University of Florida Thesis Or Dissertation Formatting
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Symbiosis Regulation in a Facultatively Symbiotic Temperate Coral: Zooxanthellae Division and Expulsion
Coral Reefs (2008) 27:601–604 DOI 10.1007/s00338-008-0363-x NOTE Symbiosis regulation in a facultatively symbiotic temperate coral: zooxanthellae division and expulsion J. Dimond Æ E. Carrington Received: 18 October 2007 / Accepted: 10 February 2008 / Published online: 29 February 2008 Springer-Verlag 2008 Abstract Zooxanthellae mitotic index (MI) and expul- Keywords Temperate coral Astrangia Zooxanthellae sion rates were measured in the facultatively symbiotic Expulsion Facultative symbiosis scleractinian Astrangia poculata during winter and summer off the southern New England coast, USA. While MI was significantly higher in summer than in winter, mean Introduction expulsion rates were comparable between seasons. Corals therefore appear to allow increases in symbiont density Many anthozoans and some other invertebrates are well when symbiosis is advantageous during the warm season, known for their endosymbiotic associations with zooxan- followed by a net reduction during the cold season when thellae (Symbiodinium sp. dinoflagellates). Living within zooxanthellae may draw resources from the coral. Given gastrodermal cells, zooxanthellae utilize host wastes and previous reports that photosynthesis in A. poculata sym- translocate photosynthetic products to the animal, in some bionts does not occur below approximately 6 C, cases fulfilling nearly all of the host’s energy demands considerable zooxanthellae division at 3 C and in darkness (Muscatine 1990). Host cells have a flexible, but finite suggests that zooxanthellae are heterotrophic at low sea- capacity for zooxanthellae, and must therefore either grow sonal temperatures. Finally, examination of expulsion as a additional cells to accommodate dividing symbionts or function of zooxanthellae density revealed that corals with regulate their numbers (Muscatine et al. -

Checklist of Fish and Invertebrates Listed in the CITES Appendices
JOINTS NATURE \=^ CONSERVATION COMMITTEE Checklist of fish and mvertebrates Usted in the CITES appendices JNCC REPORT (SSN0963-«OStl JOINT NATURE CONSERVATION COMMITTEE Report distribution Report Number: No. 238 Contract Number/JNCC project number: F7 1-12-332 Date received: 9 June 1995 Report tide: Checklist of fish and invertebrates listed in the CITES appendices Contract tide: Revised Checklists of CITES species database Contractor: World Conservation Monitoring Centre 219 Huntingdon Road, Cambridge, CB3 ODL Comments: A further fish and invertebrate edition in the Checklist series begun by NCC in 1979, revised and brought up to date with current CITES listings Restrictions: Distribution: JNCC report collection 2 copies Nature Conservancy Council for England, HQ, Library 1 copy Scottish Natural Heritage, HQ, Library 1 copy Countryside Council for Wales, HQ, Library 1 copy A T Smail, Copyright Libraries Agent, 100 Euston Road, London, NWl 2HQ 5 copies British Library, Legal Deposit Office, Boston Spa, Wetherby, West Yorkshire, LS23 7BQ 1 copy Chadwick-Healey Ltd, Cambridge Place, Cambridge, CB2 INR 1 copy BIOSIS UK, Garforth House, 54 Michlegate, York, YOl ILF 1 copy CITES Management and Scientific Authorities of EC Member States total 30 copies CITES Authorities, UK Dependencies total 13 copies CITES Secretariat 5 copies CITES Animals Committee chairman 1 copy European Commission DG Xl/D/2 1 copy World Conservation Monitoring Centre 20 copies TRAFFIC International 5 copies Animal Quarantine Station, Heathrow 1 copy Department of the Environment (GWD) 5 copies Foreign & Commonwealth Office (ESED) 1 copy HM Customs & Excise 3 copies M Bradley Taylor (ACPO) 1 copy ^\(\\ Joint Nature Conservation Committee Report No. -

Deep‐Sea Coral Taxa in the U.S. Gulf of Mexico: Depth and Geographical Distribution
Deep‐Sea Coral Taxa in the U.S. Gulf of Mexico: Depth and Geographical Distribution by Peter J. Etnoyer1 and Stephen D. Cairns2 1. NOAA Center for Coastal Monitoring and Assessment, National Centers for Coastal Ocean Science, Charleston, SC 2. National Museum of Natural History, Smithsonian Institution, Washington, DC This annex to the U.S. Gulf of Mexico chapter in “The State of Deep‐Sea Coral Ecosystems of the United States” provides a list of deep‐sea coral taxa in the Phylum Cnidaria, Classes Anthozoa and Hydrozoa, known to occur in the waters of the Gulf of Mexico (Figure 1). Deep‐sea corals are defined as azooxanthellate, heterotrophic coral species occurring in waters 50 m deep or more. Details are provided on the vertical and geographic extent of each species (Table 1). This list is adapted from species lists presented in ʺBiodiversity of the Gulf of Mexicoʺ (Felder & Camp 2009), which inventoried species found throughout the entire Gulf of Mexico including areas outside U.S. waters. Taxonomic names are generally those currently accepted in the World Register of Marine Species (WoRMS), and are arranged by order, and alphabetically within order by suborder (if applicable), family, genus, and species. Data sources (references) listed are those principally used to establish geographic and depth distribution. Only those species found within the U.S. Gulf of Mexico Exclusive Economic Zone are presented here. Information from recent studies that have expanded the known range of species into the U.S. Gulf of Mexico have been included. The total number of species of deep‐sea corals documented for the U.S. -

Adaptive Divergence, Neutral Panmixia, and Algal Symbiont Population Structure in the Temperate Coral Astrangia Poculata Along the Mid-Atlantic United States
Adaptive divergence, neutral panmixia, and algal symbiont population structure in the temperate coral Astrangia poculata along the Mid-Atlantic United States Hannah E. Aichelman1,2 and Daniel J. Barshis2 1 Department of Biology, Boston University, Boston, MA, USA 2 Department of Biological Sciences, Old Dominion University, Norfolk, VA, USA ABSTRACT Astrangia poculata is a temperate scleractinian coral that exists in facultative symbiosis with the dinoflagellate alga Breviolum psygmophilum across a range spanning the Gulf of Mexico to Cape Cod, Massachusetts. Our previous work on metabolic thermal performance of Virginia (VA) and Rhode Island (RI) populations of A. poculata revealed physiological signatures of cold (RI) and warm (VA) adaptation of these populations to their respective local thermal environments. Here, we used whole-transcriptome sequencing (mRNA-Seq) to evaluate genetic differences and identify potential loci involved in the adaptive signature of VA and RI populations. Sequencing data from 40 A. poculata individuals, including 10 colonies from each population and symbiotic state (VA-white, VA-brown, RI-white, and RI-brown), yielded a total of 1,808 host-associated and 59 algal symbiont-associated single nucleotide polymorphisms (SNPs) post filtration. Fst outlier analysis identified 66 putative high outlier SNPs in the coral host and 4 in the algal symbiont. Differentiation of VA and RI populations in the coral host was driven by putatively adaptive loci, not neutral divergence (Fst = 0.16, p = 0.001 and Fst = 0.002, p = 0.269 for outlier and neutral SNPs respectively). In contrast, we found evidence of neutral population differentiation in B. psygmophilum (Fst = 0.093, p = 0.001). -

Voestalpine Essential Fish Habitat Assessment for PSD Greenhouse Gas Permit
Essential Fish Habitat Assessment: Texas Project Site voestalpine Stahl GmbH San Patricio County, Texas January 31, 2013 www.erm.com voestalpine Stahl GmbH Essential Fish Habitat Assessment: Texas Project Site January 31, 2013 Project No. 0172451 San Patricio County, Texas Alicia Smith Partner-in-Charge Graham Donaldson Project Manager Travis Wycoff Project Consultant Environmental Resources Management 15810 Park Ten Place, Suite 300 Houston, Texas 77084-5140 T: 281-600-1000 F: 281-600-1001 Texas Registered Engineering Firm F-2393 TABLE OF CONTENTS LIST OF ACRONYMS IV EXECUTIVE SUMMARY VI 1.0 INTRODUCTION 1 1.1 PROPOSED ACTION 1 1.2 AGENCY REGULATIONS 1 1.2.1 Magnuson-Stevens Fishery Conservation and Management Act 1 1.2.1 Essential Fish Habitat Defined 2 2.0 PROJECT DESCRIPTION 4 2.1 PROJECT SCHEDULE 4 2.2 PROJECT LOCATION 4 2.3 SITE DESCRIPTION 5 2.4 SITE HISTORY 7 2.5 EMISSIONS CONTROLS 8 2.6 NOISE 9 2.7 DUST 10 2.8 WATER AND WASTEWATER 10 2.8.1 Water Sourcing and Water Rights 11 2.8.2 Wastewater Discharge 13 3.0 IDENTIFICATION OF THE ACTION AREA 15 3.1 ACTION AREA DEFINED 15 3.2 ACTION AREA DELINEATION METHODOLOGY AND RESULTS 16 3.2.1 Significant Impact Level Dispersion Modeling 16 3.2.2 Other Contaminants 17 4.0 ESSENTIAL FISH HABITAT IN THE VICINITY OF THE PROJECT 19 4.1 SPECIES OF PARTICULAR CONCERN 19 4.1.1 Brown Shrimp 19 4.1.2 Gray Snapper 20 4.1.3 Pink Shrimp 20 4.1.4 Red Drum 20 4.1.5 Spanish Mackerel 21 4.1.6 White Shrimp 21 4.2 HABITAT AREAS OF PARTICULAR CONCERN 22 5.0 ENVIRONMENTAL BASELINE CONDITIONS AND EFFECTS ANALYSIS -

Season, but Not Symbiont State, Drives Microbiome Structure in the Temperate Coral Astrangia Poculata
Roger Williams University DOCS@RWU Arts & Sciences Faculty Publications Arts and Sciences 2017 Season, but not symbiont state, drives microbiome structure in the temperate coral Astrangia poculata. Koty H. Sharp Roger Williams University, [email protected] Zoe A. Pratte Georgia Institute of Technology Allison H. Kerwin University of Connecticut, Storrs Follow this and additional works at: https://docs.rwu.edu/fcas_fp Part of the Biology Commons, Marine Biology Commons, and the Microbiology Commons Recommended Citation Sharp, K.H., Pratte, Z.A., Kerwin, A.H., Rotjan, R.D., & Stewart, F. J. (2017). Season, but not symbiont state, drives microbiome structure in the temperate coral Astrangia poculata. Microbiome 5(1), 120. This Article is brought to you for free and open access by the Arts and Sciences at DOCS@RWU. It has been accepted for inclusion in Arts & Sciences Faculty Publications by an authorized administrator of DOCS@RWU. For more information, please contact [email protected]. Sharp et al. Microbiome (2017) 5:120 DOI 10.1186/s40168-017-0329-8 RESEARCH Open Access Season, but not symbiont state, drives microbiome structure in the temperate coral Astrangia poculata Koty H. Sharp1*, Zoe A. Pratte2, Allison H. Kerwin3, Randi D. Rotjan 4,5 and Frank J. Stewart2 Abstract Background: Understanding the associations among corals, their photosynthetic zooxanthella symbionts (Symbiodinium), and coral-associated prokaryotic microbiomes is critical for predicting the fidelity and strength of coral symbioses in the face of growing environmental threats. Most coral-microbiome associations are beneficial, yet the mechanisms that determine the composition of the coral microbiome remain largely unknown. Here, we characterized microbiome diversity in the temperate, facultatively symbiotic coral Astrangia poculata at four seasonal time points near the northernmost limit of the species range. -
Summary Output
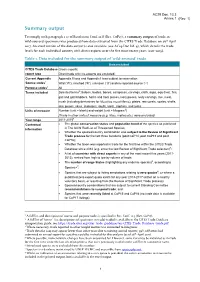
AC29 Doc. 13.3 Annex 1 Summary output To comply with paragraph 1 a) of Resolution Conf. 12.8 (Rev. CoP17), a summary output of trade in wild-sourced specimens was produced from data extracted from the CITES Trade Database on 26th April 2017. An excel version of the data output is also available (see AC29 Doc Inf. 4), which details the trade levels for each individual country with direct exports over the five most recent years (2011-2015). Table 1. Data included for the summary output of ‘wild-sourced’ trade Data included CITES Trade Database Gross exports; report type Direct trade only (re-exports are excluded) Current Appendix Appendix II taxa and Appendix I taxa subject to reservation Source codes1 Wild (‘W’), ranched (‘R’), unknown (‘U’) and no reported source (‘-’) Purpose codes1 All Terms included Selected terms2: baleen, bodies, bones, carapaces, carvings, cloth, eggs, egg (live), fins, gall and gall bladders, horns and horn pieces, ivory pieces, ivory carvings, live, meat, musk (including derivatives for Moschus moschiferus), plates, raw corals, scales, shells, skin pieces, skins, skeletons, skulls, teeth, trophies, and tusks. Units of measure Number (unit = blank) and weight (unit = kilogram3) [Trade in other units of measure (e.g. litres, metres etc.) were excluded] Year range 2011-20154 Contextual The global conservation status and population trend of the species as published information in The IUCN Red List of Threatened Species; Whether the species/country combination was subject to the Review of Significant Trade process for the last three iterations (post CoP14, post CoP15 and post CoP16); Whether the taxon was reported in trade for the first time within the CITES Trade Database since 2012 (e.g. -

Astrangia Poculata Koty H
Sharp et al. Microbiome (2017) 5:120 DOI 10.1186/s40168-017-0329-8 RESEARCH Open Access Season, but not symbiont state, drives microbiome structure in the temperate coral Astrangia poculata Koty H. Sharp1*, Zoe A. Pratte2, Allison H. Kerwin3, Randi D. Rotjan 4,5 and Frank J. Stewart2 Abstract Background: Understanding the associations among corals, their photosynthetic zooxanthella symbionts (Symbiodinium), and coral-associated prokaryotic microbiomes is critical for predicting the fidelity and strength of coral symbioses in the face of growing environmental threats. Most coral-microbiome associations are beneficial, yet the mechanisms that determine the composition of the coral microbiome remain largely unknown. Here, we characterized microbiome diversity in the temperate, facultatively symbiotic coral Astrangia poculata at four seasonal time points near the northernmost limit of the species range. The facultative nature of this system allowed us to test seasonal influence and symbiotic state (Symbiodinium density in the coral) on microbiome community composition. Results: Change in season had a strong effect on A. poculata microbiome composition. The seasonal shift was greatest upon the winter to spring transition, during which time A. poculata microbiome composition became more similar among host individuals. Within each of the four seasons, microbiome composition differed significantly from that of surrounding seawater but was surprisingly uniform between symbiotic and aposymbiotic corals, even in summer, when differences in Symbiodinium density between brown and white colonies are the highest, indicating that the observed seasonal shifts are not likely due to fluctuations in Symbiodinium density. Conclusions: Our results suggest that symbiotic state may not be a primary driver of coral microbial community organization in A. -

Inventory and Atlas of Corals and Coral Reefs, with Emphasis on Deep-Water Coral Reefs from the U
Inventory and Atlas of Corals and Coral Reefs, with Emphasis on Deep-Water Coral Reefs from the U. S. Caribbean EEZ Jorge R. García Sais SEDAR26-RD-02 FINAL REPORT Inventory and Atlas of Corals and Coral Reefs, with Emphasis on Deep-Water Coral Reefs from the U. S. Caribbean EEZ Submitted to the: Caribbean Fishery Management Council San Juan, Puerto Rico By: Dr. Jorge R. García Sais dba Reef Surveys P. O. Box 3015;Lajas, P. R. 00667 [email protected] December, 2005 i Table of Contents Page I. Executive Summary 1 II. Introduction 4 III. Study Objectives 7 IV. Methods 8 A. Recuperation of Historical Data 8 B. Atlas map of deep reefs of PR and the USVI 11 C. Field Study at Isla Desecheo, PR 12 1. Sessile-Benthic Communities 12 2. Fishes and Motile Megabenthic Invertebrates 13 3. Statistical Analyses 15 V. Results and Discussion 15 A. Literature Review 15 1. Historical Overview 15 2. Recent Investigations 22 B. Geographical Distribution and Physical Characteristics 36 of Deep Reef Systems of Puerto Rico and the U. S. Virgin Islands C. Taxonomic Characterization of Sessile-Benthic 49 Communities Associated With Deep Sea Habitats of Puerto Rico and the U. S. Virgin Islands 1. Benthic Algae 49 2. Sponges (Phylum Porifera) 53 3. Corals (Phylum Cnidaria: Scleractinia 57 and Antipatharia) 4. Gorgonians (Sub-Class Octocorallia 65 D. Taxonomic Characterization of Sessile-Benthic Communities 68 Associated with Deep Sea Habitats of Puerto Rico and the U. S. Virgin Islands 1. Echinoderms 68 2. Decapod Crustaceans 72 3. Mollusks 78 E. -

Hermit Crabs - Paguridae and Diogenidae
Identification Guide to Marine Invertebrates of Texas by Brenda Bowling Texas Parks and Wildlife Department April 12, 2019 Version 4 Page 1 Marine Crabs of Texas Mole crab Yellow box crab Giant hermit Surf hermit Lepidopa benedicti Calappa sulcata Petrochirus diogenes Isocheles wurdemanni Family Albuneidae Family Calappidae Family Diogenidae Family Diogenidae Blue-spot hermit Thinstripe hermit Blue land crab Flecked box crab Paguristes hummi Clibanarius vittatus Cardisoma guanhumi Hepatus pudibundus Family Diogenidae Family Diogenidae Family Gecarcinidae Family Hepatidae Calico box crab Puerto Rican sand crab False arrow crab Pink purse crab Hepatus epheliticus Emerita portoricensis Metoporhaphis calcarata Persephona crinita Family Hepatidae Family Hippidae Family Inachidae Family Leucosiidae Mottled purse crab Stone crab Red-jointed fiddler crab Atlantic ghost crab Persephona mediterranea Menippe adina Uca minax Ocypode quadrata Family Leucosiidae Family Menippidae Family Ocypodidae Family Ocypodidae Mudflat fiddler crab Spined fiddler crab Longwrist hermit Flatclaw hermit Uca rapax Uca spinicarpa Pagurus longicarpus Pagurus pollicaris Family Ocypodidae Family Ocypodidae Family Paguridae Family Paguridae Dimpled hermit Brown banded hermit Flatback mud crab Estuarine mud crab Pagurus impressus Pagurus annulipes Eurypanopeus depressus Rithropanopeus harrisii Family Paguridae Family Paguridae Family Panopeidae Family Panopeidae Page 2 Smooth mud crab Gulf grassflat crab Oystershell mud crab Saltmarsh mud crab Hexapanopeus angustifrons Dyspanopeus -

Assessing the Effects of Cold Thermal Stress on Astrangia Poculata
Assessing the effects of cold thermal stress on Astrangia poculata quiescence: analyzing photosynthetic efficiency, calcification rate, polyp behavior and Symbiodinium Sara Brennan, Mary Liesegang, Adeline Almanzar, Daniel Chavez Abstract: Astrangia poculata is a model organism due to its tolerance to both cold and warm water and its facultative relationship with its Symbiodinium. Understanding how this temperate coral tolerates such a wide range of temperatures, with and without symbionts, can help us understand what will happen in the future as the climate becomes more variable and the world experiences both higher high temperatures and colder cold temperatures. We conducted a common garden laboratory experiment in order to test the impacts of cold thermal stress on A. poculata. Data from photosynthetic efficiency, calcification rate, polyp behavior, and Symbiodinium were used to analyze the decreasing health of the coral as it approached and ultimately entered quiescence. We found no significant difference in calcification rates between the controls at 22°C and the experimentals at decreasing temperatures. We also found a decreasing trend in photosynthetic efficiency in experimental corals and a decrease in polyp behavior as a result of cold stress. Our results show that A. poculata is very resilient, quickly recovers from thermal stress and enters dormancy at about 7°C. Introduction: From spikes in global temperatures to Most coral species exhibit obligate symbiosis, adapted migration patterns of organisms, the meaning that they will eventually die if they Earth is changing rapidly due to global climate experience an extended amount of time change. This is particularly problematic for without their Symbiodinium, a single celled coral reefs, comprising sensitive, slow algae living in the coral (Muscatine & Porter, growing organism, with 30% of corals already 1977). -

2019 Astrangia Research Workshop Program Schedule at a Glance Wednesday, May 29 8:00 – 9:00 Breakfast/Check-In 9:00 – 9:30
2019 Astrangia Research Workshop Program Schedule At A Glance Wednesday, May 29 8:00 – 9:00 Breakfast/Check-In 9:00 – 9:30 Welcome/Introductions 9:30 – 10:15 Introduction and Inspiration: History of Astrangia Science 10:15 – 11:15 Research Session I – Community Ecology 11:15 - 12:15 Poster Session 12:15 – 1:30 Lunch (Upper Dining Commons, RWU) 1:30 – 3:30 Research Session II – Genetic and Genomic Insights 3:30 - 3:45 Coffee Break 3:45 - 5:15 Research Session III - Microbiome 5:30 – 6:00 RWU Oyster Sampling (RWU Docks) 6:00 – 7:00 Dinner (Upper Dining Commons, RWU) 7:30 – 10:00 Sunset Porch Party (Portsmouth, RI) Thursday, May 30 8:00 – 9:00 Breakfast 9:00 – 10:45 Research Session IV – Resilience Ecology 10:45 - 11:30 Research Session V: Astrangia Spawning & Early Life Cycle 11:30 – 1:00 Working Lunch & Small Breakout Groups: Education and Outreach 1:00 - 2:00 Education/Outreach Consensus Discussion 2:00 – 4:00 Looking to the Future: Questions Remaining 4:00 Workshop Wrap-Up 2019 Astrangia Research Workshop Program Wednesday, May 29 8:00 – 9:00 Breakfast/Check-In 9:00 – 9:30 Welcome/Introductions 9:30 – 10:15 Introduction and Inspiration: Sean Grace - “History of Astrangia Science” 10:15 – 11:15 Research Session I: Community Ecology Moderator: Koty Sharp Gabriella DiPreta - “Increasing prey availability with a cost-effective light rack inhibits quiescence and dormancy in the temperate scleractinian coral Astrangia poculata” (lightning talk) Sean Grace - “Shallow subtidal community changes in Southern New England; future challenges for temperate corals” Sean Grace - “Current Projects in the Grace Lab” (lightning talk) Caroline Fleming - “Building a dynamic energy budget for A.