Systematic Analyses of Drugs and Disease Indications In
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Therapeutic Potential of Nicotinamide Adenine Dinucleotide (NAD) T ∗ Marta Arenas-Jala,B, , J.M
European Journal of Pharmacology 879 (2020) 173158 Contents lists available at ScienceDirect European Journal of Pharmacology journal homepage: www.elsevier.com/locate/ejphar Therapeutic potential of nicotinamide adenine dinucleotide (NAD) T ∗ Marta Arenas-Jala,b, , J.M. Suñé-Negrea, Encarna García-Montoyaa a Pharmacy and Pharmaceutical Technology Department (Faculty of Pharmacy and Food Sciences), University of Barcelona, Barcelona, Spain b ICN2 – Catalan Institute of Nanoscience and Nanotechnology (Autonomous University of Barcelona), Bellaterra (Barcelona), Spain ARTICLE INFO ABSTRACT Keywords: Nicotinamide adenine nucleotide (NAD) is a small ubiquitous hydrophilic cofactor that participates in several NAD aspects of cellular metabolism. As a coenzyme it has an essential role in the regulation of energetic metabolism, Metabolism but it is also a cosubstrate for enzymes that regulate fundamental biological processes such as transcriptional Therapeutic potential regulation, signaling and DNA repairing among others. The fluctuation and oxidative state of NAD levels reg- Drug discovery ulate the activity of these enzymes, which is translated into marked effects on cellular function. While alterations Supplementation in NAD homeostasis are a common feature of different conditions and age-associated diseases, in general, in- creased NAD levels have been associated with beneficial health effects. Due to its therapeutic potential, the interest in this molecule has been renewed, and the regulation of NAD metabolism has become an attractive target for drug discovery. In fact, different approaches to replenish or increase NAD levels have been tested, including enhancement of biosynthesis and inhibition of NAD breakdown. Despite further research is needed, this review provides an overview and update on NAD metabolism, including the therapeutic potential of its regulation, as well as pharmacokinetics, safety, precautions and formulation challenges of NAD supplementa- tion. -

Metabolic-Hydroxy and Carboxy Functionalization of Alkyl Moieties in Drug Molecules: Prediction of Structure Influence and Pharmacologic Activity
molecules Review Metabolic-Hydroxy and Carboxy Functionalization of Alkyl Moieties in Drug Molecules: Prediction of Structure Influence and Pharmacologic Activity Babiker M. El-Haj 1,* and Samrein B.M. Ahmed 2 1 Department of Pharmaceutical Sciences, College of Pharmacy and Health Sciences, University of Science and Technology of Fujairah, Fufairah 00971, UAE 2 College of Medicine, Sharjah Institute for Medical Research, University of Sharjah, Sharjah 00971, UAE; [email protected] * Correspondence: [email protected] Received: 6 February 2020; Accepted: 7 April 2020; Published: 22 April 2020 Abstract: Alkyl moieties—open chain or cyclic, linear, or branched—are common in drug molecules. The hydrophobicity of alkyl moieties in drug molecules is modified by metabolic hydroxy functionalization via free-radical intermediates to give primary, secondary, or tertiary alcohols depending on the class of the substrate carbon. The hydroxymethyl groups resulting from the functionalization of methyl groups are mostly oxidized further to carboxyl groups to give carboxy metabolites. As observed from the surveyed cases in this review, hydroxy functionalization leads to loss, attenuation, or retention of pharmacologic activity with respect to the parent drug. On the other hand, carboxy functionalization leads to a loss of activity with the exception of only a few cases in which activity is retained. The exceptions are those groups in which the carboxy functionalization occurs at a position distant from a well-defined primary pharmacophore. Some hydroxy metabolites, which are equiactive with their parent drugs, have been developed into ester prodrugs while carboxy metabolites, which are equiactive to their parent drugs, have been developed into drugs as per se. -

Global LC/MS Metabolomics Profiling of Calcium Stressed and Immunosuppressant Drug Treated Saccharomyces Cerevisiae
Metabolites 2013, 3, 1102-1117; doi:10.3390/metabo3041102 OPEN ACCESS metabolites ISSN 2218-1989 www.mdpi.com/journal/metabolites/ Article Global LC/MS Metabolomics Profiling of Calcium Stressed and Immunosuppressant Drug Treated Saccharomyces cerevisiae Stefan Jenkins 1,2,3,*, Steven M. Fischer 2, Lily Chen 3 and Theodore R. Sana 2 1 Life Sciences Division, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA 2 Agilent Technologies, Life Sciences, Diagnostics and Applied Markets, Santa Clara, CA 95051, USA; E-Mails: [email protected] (S.M.F.); [email protected] (T.R.S.) 3 Biology Department, San Francisco State University, San Francisco, CA 94132, USA; E-Mail: [email protected] * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: +1-650-922-5046; Fax: +1-510-486-4545. Received: 15 October 2013; in revised form: 20 November 2013 / Accepted: 25 November 2013 / Published: 6 December 2013 Abstract: Previous studies have shown that calcium stressed Saccharomyces cerevisiae, challenged with immunosuppressant drugs FK506 and Cyclosporin A, responds with comprehensive gene expression changes and attenuation of the generalized calcium stress response. Here, we describe a global metabolomics workflow for investigating the utility of tracking corresponding phenotypic changes. This was achieved by efficiently analyzing relative abundance differences between intracellular metabolite pools from wild-type and calcium stressed cultures, with and without prior immunosuppressant drugs exposure. We used pathway database content from WikiPathways and YeastCyc to facilitate the projection of our metabolomics profiling results onto biological pathways. A key challenge was to increase the coverage of the detected metabolites. This was achieved by applying both reverse phase (RP) and aqueous normal phase (ANP) chromatographic separations, as well as electrospray ionization (ESI) and atmospheric pressure chemical ionization (APCI) sources for detection in both ion polarities. -

STRING V10: Protein–Protein Interaction Networks, Integrated
Published online 28 October 2014 Nucleic Acids Research, 2015, Vol. 43, Database issue D447–D452 doi: 10.1093/nar/gku1003 STRING v10: protein–protein interaction networks, integrated over the tree of life Damian Szklarczyk1, Andrea Franceschini1, Stefan Wyder1, Kristoffer Forslund2, Davide Heller1, Jaime Huerta-Cepas2, Milan Simonovic1, Alexander Roth1, Alberto Santos3, Kalliopi P. Tsafou3, Michael Kuhn4,5, Peer Bork2,*, Lars J. Jensen3,* and Christian von Mering1,* 1Institute of Molecular Life Sciences and Swiss Institute of Bioinformatics, University of Zurich, 8057 Zurich, Switzerland, 2European Molecular Biology Laboratory, 69117 Heidelberg, Germany, 3Novo Nordisk Foundation Center for Protein Research, University of Copenhagen, 2200 Copenhagen N, Denmark, 4Biotechnology Center, Technische Universitat¨ Dresden, 01062 Dresden, Germany and 5Max Planck Institute of Molecular Cell Biology and Downloaded from Genetics, 01062 Dresden, Germany Received September 15, 2014; Accepted October 07, 2014 ABSTRACT INTRODUCTION http://nar.oxfordjournals.org/ The many functional partnerships and interactions For a full description of a protein’s function, knowledge that occur between proteins are at the core of cel- about its specific interaction partners is an important pre- lular processing and their systematic characteriza- requisite. The concept of protein ‘function’ is somewhat hi- tion helps to provide context in molecular systems erarchical (1–4), and at all levels in this hierarchy, interac- biology. However, known and predicted interactions tions between proteins help to describe and narrow down a protein’s function: its three-dimensional structure may are scattered over multiple resources, and the avail- become meaningful only in the context of a larger pro- able data exhibit notable differences in terms of qual- tein assembly, its molecular actions may be regulated by at University of Aarhus on January 22, 2015 ity and completeness. -

And Macro-Level Network Metrics Unveils Top Communicative Gene Modules in Psoriasis
G C A T T A C G G C A T genes Article Using Micro- and Macro-Level Network Metrics Unveils Top Communicative Gene Modules in Psoriasis 1, 2 3 Reyhaneh Naderi y, Homa Saadati Mollaei , Arne Elofsson 3, , and Saman Hosseini Ashtiani * y 1 Department of Artificial Intelligence and Robotics, Faculty of Computer Engineering, Iran University of Science and Technology, Tehran 1684613114, Iran; [email protected] 2 Department of Advanced Sciences and Technology, Islamic Azad University Tehran Medical Sciences, Tehran 1916893813, Iran; [email protected] 3 Department of Biochemistry and Biophysics and Science for Life Laboratory, Stockholm University, 106 91 Stockholm, Sweden; [email protected] * Correspondence: [email protected]; Tel.: +46-762623644 These authors contributed equally to this work and should be considered joint first authors. y Received: 9 July 2020; Accepted: 6 August 2020; Published: 10 August 2020 Abstract: (1) Background: Psoriasis is a multifactorial chronic inflammatory disorder of the skin, with significant morbidity, characterized by hyperproliferation of the epidermis. Even though psoriasis’ etiology is not fully understood, it is believed to be multifactorial, with numerous key components. (2) Methods: In order to cast light on the complex molecular interactions in psoriasis vulgaris at both protein–protein interactions and transcriptomics levels, we studied a set of microarray gene expression analyses consisting of 170 paired lesional and non-lesional samples. Afterwards, a network analysis was conducted on the protein–protein interaction network of differentially expressed genes based on micro- and macro-level network metrics at a systemic level standpoint. (3) Results: We found 17 top communicative genes, all of which were experimentally proven to be pivotal in psoriasis, which were identified in two modules, namely the cell cycle and immune system. -

Biological Pathways
Biological pathways Bing Zhang Department of Biomedical Informatics Vanderbilt University [email protected] Biological pathway A biological pathway is a series of actions among molecules in a cell that leads to a certain product or a change in a cell. Different types of biological pathways Signal transduction pathways Gene regulation pathways Metabolic pathways 2 Applied Bioinformatics, Spring 2011 Outline Pathway databases Pathway assembly and editing Pathway mapping and enrichment analysis 3 Applied Bioinformatics, Spring 2011 Selected pathway databases KEGG Kyoto Encyclopedia of Genes and Genomes http://www.genome.jp/kegg/pathway.html Reactome Ontario Institute for Cancer Research, Cold Spring Harbor Laboratory, New York University School of Medicine and The European Bioinformatics Institute http://www.reactome.org/ WikiPathways University of Maastricht and UCSF http://www.wikipathways.org/ 4 Applied Bioinformatics, Spring 2011 KEGG: data model Molecular building blocks KEGG GENES: genes and proteins generated by genome sequencing projects KEGG ORTHOLOGY: orthology (KO) groups KEGG COMPOUND: small molecules KEGG REACTIONS: biochemical reactions KEGG PATHWAY: pathway maps Created in a general way to be applicable to all organisms, in terms of the orthologs defined by KO groups Organism-specific pathways can be generated by converting KO groups to gene identifiers in a given organism 5 Applied Bioinformatics, Spring 2011 KEGG: TCA cycle 6 Applied Bioinformatics, Spring 2011 Reactome: data model Entities -

Drugbank 3.0: a Comprehensive Resource for 'Omics' Research On
Published online 8 November 2010 Nucleic Acids Research, 2011, Vol. 39, Database issue D1035–D1041 doi:10.1093/nar/gkq1126 DrugBank 3.0: a comprehensive resource for ‘Omics’ research on drugs Craig Knox1, Vivian Law2, Timothy Jewison1, Philip Liu3, Son Ly2, Alex Frolkis1, Allison Pon1, Kelly Banco2, Christine Mak2, Vanessa Neveu1, Yannick Djoumbou3, Roman Eisner1, An Chi Guo1 and David S. Wishart1,2,3,4,* 1Department of Computing Science, University of Alberta, Edmonton, AB, Canada T6G 2E8, 2Faculty of Pharmacy and Pharmaceutical Sciences, University of Alberta, Edmonton, AB, Canada T6G 2N8, 3Department of Biological Sciences, University of Alberta, Edmonton, AB, Canada T6G 2E8 and 4National Institute for Nanotechnology, 11421 Saskatchewan Drive, Edmonton, AB, Canada T6G 2M9 Received September 15, 2010; Revised October 20, 2010; Accepted October 21, 2010 ABSTRACT drug target, drug description and drug action data. DrugBank (http://www.drugbank.ca) is a richly DrugBank 3.0 represents the result of 2 years annotated database of drug and drug target infor- of manual annotation work aimed at making mation. It contains extensive data on the nomencla- the database much more useful for a wide ture, ontology, chemistry, structure, function, range of ‘omics’ (i.e. pharmacogenomic, action, pharmacology, pharmacokinetics, metabol- pharmacoproteomic, pharmacometabolomic and ism and pharmaceutical properties of both small even pharmacoeconomic) applications. molecule and large molecule (biotech) drugs. It also contains comprehensive information on the INTRODUCTION target diseases, proteins, genes and organisms on which these drugs act. First released in 2006, Historically most of the known information on drugs, DrugBank has become widely used by pharmacists, drug targets and drug action has resided in books, medicinal chemists, pharmaceutical researchers, journals and expensive commercial databases. -

Smart Drugs: a Review
International Journal for Innovation Education and Research www.ijier.net Vol:-8 No-11, 2020 Smart Drugs: A Review Sahjesh Soni, Dr Rashmi Srivastava, Ayush Bhandari Mumbai Educational Trust, India Abstracts Smart drugs can change the way our mind functions. Smart drugs are also known as nootropics, which literally means the ability to bend or shape our mind. Smart drugs are classified into two main categories. They are classified based on their pharmacological action and their availability. The stimulant category of drugs is highly used and misused. There has been a rampant increase in the sale of smart drugs, which could be attributed to the rise in competition all over the world. Two major criteria for selecting a good drug are its mechanism of action and bioavailability. Owing to the short-term benefits of smart drugs, many countries have openly accepted this concept. There is still no concrete scientific evidence backing the safety and efficacy of these drugs. Some believe that this is just a fad that will soon pass, while others believe that this is something that will revolutionize our future. Key Words: Smart drugs, Nootropics, Cognitive enhancers, Stimulants, Uses and Side effects. What are Smart Drugs? "Smart drugs" are a group of compounds that can promote brain performance. They have got a lot of attention due to our stressful lifestyle, and these drugs help to boost our memory, focus, creativity, intelligence, and motivation. The origin of the word comes from the Greek language meaning “to bend or shape the mind”.1 These chemicals have many mechanisms of action. -

Confirmation of Pathogenic Mechanisms by SARS-Cov-2–Host
Messina et al. Cell Death and Disease (2021) 12:788 https://doi.org/10.1038/s41419-021-03881-8 Cell Death & Disease ARTICLE Open Access Looking for pathways related to COVID-19: confirmation of pathogenic mechanisms by SARS-CoV-2–host interactome Francesco Messina 1, Emanuela Giombini1, Chiara Montaldo1, Ashish Arunkumar Sharma2, Antonio Zoccoli3, Rafick-Pierre Sekaly2, Franco Locatelli4, Alimuddin Zumla5, Markus Maeurer6,7, Maria R. Capobianchi1, Francesco Nicola Lauria1 and Giuseppe Ippolito 1 Abstract In the last months, many studies have clearly described several mechanisms of SARS-CoV-2 infection at cell and tissue level, but the mechanisms of interaction between host and SARS-CoV-2, determining the grade of COVID-19 severity, are still unknown. We provide a network analysis on protein–protein interactions (PPI) between viral and host proteins to better identify host biological responses, induced by both whole proteome of SARS-CoV-2 and specific viral proteins. A host-virus interactome was inferred, applying an explorative algorithm (Random Walk with Restart, RWR) triggered by 28 proteins of SARS-CoV-2. The analysis of PPI allowed to estimate the distribution of SARS-CoV-2 proteins in the host cell. Interactome built around one single viral protein allowed to define a different response, underlining as ORF8 and ORF3a modulated cardiovascular diseases and pro-inflammatory pathways, respectively. Finally, the network-based approach highlighted a possible direct action of ORF3a and NS7b to enhancing Bradykinin Storm. This network-based representation of SARS-CoV-2 infection could be a framework for pathogenic evaluation of specific 1234567890():,; 1234567890():,; 1234567890():,; 1234567890():,; clinical outcomes. -

Drug Knowledge Bases and Their Applications in Biomedical Informatics Research Yongjun Zhu, Olivier Elemento, Jyotishman Pathak and Fei Wang
Briefings in Bioinformatics, 2018, 1–14 doi: 10.1093/bib/bbx169 Paper Drug knowledge bases and their applications in biomedical informatics research Yongjun Zhu, Olivier Elemento, Jyotishman Pathak and Fei Wang Corresponding author: Fei Wang, Division of Health Informatics, Department of Healthcare Policy and Research at Weill Cornell Medicine at Cornell University, 425 East 61st Street, Suite 301, DV-308, New York, NY 10065, USA. E-mail: [email protected] Abstract Recent advances in biomedical research have generated a large volume of drug-related data. To effectively handle this flood of data, many initiatives have been taken to help researchers make good use of them. As the results of these initiatives, many drug knowledge bases have been constructed. They range from simple ones with specific focuses to comprehensive ones that contain information on almost every aspect of a drug. These curated drug knowledge bases have made significant contributions to the development of efficient and effective health information technologies for better health-care service delivery. Understanding and comparing existing drug knowledge bases and how they are applied in various biomedical studies will help us recognize the state of the art and design better knowledge bases in the future. In addition, researchers can get insights on novel applications of the drug knowledge bases through a review of successful use cases. In this study, we provide a review of existing popular drug knowledge bases and their applications in drug-related studies. We discuss challenges in constructing and using drug knowledge bases as well as future research directions toward a better ecosystem of drug knowledge bases. -
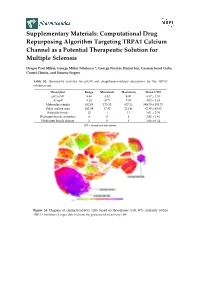
Computational Drug Repurposing Algorithm Targeting TRPA1 Calcium Channel As a Potential Therapeutic Solution for Multiple Sclerosis
Supplementary Materials: Computational Drug Repurposing Algorithm Targeting TRPA1 Calcium Channel as a Potential Therapeutic Solution for Multiple Sclerosis Dragos Paul Mihai, George Mihai Nitulescu *, George Nicolae Daniel Ion, Cosmin Ionut Ciotu, Cornel Chirita, and Simona Negres Table S1. Descriptive statistics for pIC50 and druglikeness-related descriptors for the TRPA1 inhibitors set. Descriptor Range Minimum Maximum Mean ± SD pIC50 (M) 4.48 4.52 9.00 6.57 ± 1.01 ALogP 8.28 −0.71 7.57 4.02 ± 1.34 Molecular weight 482.03 175.10 657.13 389.70 ± 101.73 Polar surface area 193.59 17.82 211.41 82.80 ± 40.83 Rotatable bonds 12 1 13 5.01 ± 2.08 Hydrogen bonds acceptors 8 0 8 2.92 ± 1.46 Hydrogen bonds donors 3 0 3 1.06 ± 0.54 SD – standard deviation. Figure S1. Diagram of similarity/activity cliffs based on flexophores with 80% similarity within TRPA1 inhibitors. Larger dots indicate the presence of an activity cliff. Figure S2. Representative structures for similarity/activity cliffs analysis of TRPA1 inhibitors. Table S2. Highest similarity pairs between TRPA1 inhibitors and screened drugs based on flexophore descriptors data mining procedure. TRPA1 inhibitors Repurposing dataset Entry Similarity (ChEMBL ID) (DrugBank ID) 1 CHEMBL3298238 DB08135 0.9832 2 CHEMBL3220230 DB08561 0.9696 3 CHEMBL3220228 DB08561 0.9614 4 CHEMBL593902 DB07311 0.9553 5 CHEMBL3297780 DB01065 0.9533 6 CHEMBL3220448 DB08561 0.9509 Figure S3. Diagram of similarity/activity cliffs based on flexophores with 80% similarity threshold for merged TRPA1 inhibitors dataset (colored dots) and similar DrugBank entries (grey dots). Table S3. -

Package 'Dbparser'
Package ‘dbparser’ August 26, 2020 Title 'DrugBank' Database XML Parser Version 1.2.0 Description This tool is for parsing the 'DrugBank' XML database <https://www.drugbank.ca/>. The parsed data are then returned in a proper 'R' dataframe with the ability to save them in a given database. License MIT + file LICENSE Encoding UTF-8 LazyData true Imports DBI, dplyr, odbc, progress, purrr, readr, RMariaDB, RSQLite, tibble, tools, XML RoxygenNote 7.1.0 Suggests knitr, rmarkdown, testthat VignetteBuilder knitr URL https://docs.ropensci.org/dbparser/, https://github.com/ropensci/dbparser/ BugReports https://github.com/ropensci/dbparser/issues Depends R (>= 2.10) NeedsCompilation no Author Mohammed Ali [aut, cre], Ali Ezzat [aut], Hao Zhu [rev], Emma Mendelsohn [rev] Maintainer Mohammed Ali <[email protected]> Repository CRAN Date/Publication 2020-08-26 12:10:03 UTC 1 2 R topics documented: R topics documented: articles . .3 attachments . .5 books . .8 cett.............................................. 10 cett_actions_doc . 12 cett_doc . 14 cett_ex_identity_doc . 17 cett_go_doc . 19 cett_poly_doc . 21 cett_poly_pfms_doc . 24 cett_poly_syn_doc . 26 dbparser . 28 drugs . 29 drug_affected_organisms . 31 drug_ahfs_codes . 33 drug_atc_codes . 35 drug_calc_prop . 36 drug_categories . 38 drug_classification . 40 drug_dosages . 42 drug_element . 44 drug_element_options . 46 drug_exp_prop . 47 drug_external_links . 49 drug_ex_identity . 51 drug_food_interactions . 53 drug_general_information . 54 drug_groups . 57 drug_interactions . 58 drug_intern_brand