Inactive USP14 and Inactive UCHL5 Cause Accumulation of Distinct Ubiquitinated Proteins In
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Potential Role of Necroptosis in Clinical Diseases (Review)
INTERNATIONAL JOURNAL OF MOleCular meDICine 47: 89, 2021 The potential role of necroptosis in clinical diseases (Review) WENLI DAI1*, JIN CHENG1*, XI LENG2, XIAOQING HU1 and YINGFANG AO1 1Institute of Sports Medicine, Beijing Key Laboratory of Sports Injuries, Peking University Third Hospital, Beijing 100191; 2Medical Imaging Center, The First Affiliated Hospital of Guangzhou University of Chinese Medicine, Guangzhou, Guangdong 510405, P.R. China Received November 19, 2020; Accepted March 8, 2021 DOI: 10.3892/ijmm.2021.4922 Abstract. As an important type of programmed cell death in 1. Introduction addition to apoptosis, necroptosis occurs in a variety of patho‑ physiological processes, including infections, liver diseases, Necroptosis, an emerging field closely related to apoptosis, is a kidney injury, neurodegenerative diseases, cardiovascular non‑caspase‑dependent cell death that has been implicated in diseases, and human tumors. It can be triggered by a variety the pathological processes of various diseases. It is regulated of factors, such as tumor necrosis factor receptor and Toll‑like by various genes that cause regular and ordered cell death. receptor families, intracellular DNA and RNA sensors, and Through activating specific death signaling pathways, it shares interferon, and is mainly mediated by receptor‑interacting typical characteristics of necrosis, including loss of metabolic protein kinase 1 (RIP1), RIP3, and mixed lineage kinase function and subcellular changes (1,2). Receptor‑interacting domain‑like protein. A better understanding of the mechanism protein kinase 1 (RIP1) was the first signaling molecule of necroptosis may be useful in the development of novel drugs identified in the necrosome (3). RIP1 and RIP3 interact for necroptosis‑related diseases. -

Table S1. List of Proteins in the BAHD1 Interactome
Table S1. List of proteins in the BAHD1 interactome BAHD1 nuclear partners found in this work yeast two-hybrid screen Name Description Function Reference (a) Chromatin adapters HP1α (CBX5) chromobox homolog 5 (HP1 alpha) Binds histone H3 methylated on lysine 9 and chromatin-associated proteins (20-23) HP1β (CBX1) chromobox homolog 1 (HP1 beta) Binds histone H3 methylated on lysine 9 and chromatin-associated proteins HP1γ (CBX3) chromobox homolog 3 (HP1 gamma) Binds histone H3 methylated on lysine 9 and chromatin-associated proteins MBD1 methyl-CpG binding domain protein 1 Binds methylated CpG dinucleotide and chromatin-associated proteins (22, 24-26) Chromatin modification enzymes CHD1 chromodomain helicase DNA binding protein 1 ATP-dependent chromatin remodeling activity (27-28) HDAC5 histone deacetylase 5 Histone deacetylase activity (23,29,30) SETDB1 (ESET;KMT1E) SET domain, bifurcated 1 Histone-lysine N-methyltransferase activity (31-34) Transcription factors GTF3C2 general transcription factor IIIC, polypeptide 2, beta 110kDa Required for RNA polymerase III-mediated transcription HEYL (Hey3) hairy/enhancer-of-split related with YRPW motif-like DNA-binding transcription factor with basic helix-loop-helix domain (35) KLF10 (TIEG1) Kruppel-like factor 10 DNA-binding transcription factor with C2H2 zinc finger domain (36) NR2F1 (COUP-TFI) nuclear receptor subfamily 2, group F, member 1 DNA-binding transcription factor with C4 type zinc finger domain (ligand-regulated) (36) PEG3 paternally expressed 3 DNA-binding transcription factor with -

Mouse Germ Line Mutations Due to Retrotransposon Insertions Liane Gagnier1, Victoria P
Gagnier et al. Mobile DNA (2019) 10:15 https://doi.org/10.1186/s13100-019-0157-4 REVIEW Open Access Mouse germ line mutations due to retrotransposon insertions Liane Gagnier1, Victoria P. Belancio2 and Dixie L. Mager1* Abstract Transposable element (TE) insertions are responsible for a significant fraction of spontaneous germ line mutations reported in inbred mouse strains. This major contribution of TEs to the mutational landscape in mouse contrasts with the situation in human, where their relative contribution as germ line insertional mutagens is much lower. In this focussed review, we provide comprehensive lists of TE-induced mouse mutations, discuss the different TE types involved in these insertional mutations and elaborate on particularly interesting cases. We also discuss differences and similarities between the mutational role of TEs in mice and humans. Keywords: Endogenous retroviruses, Long terminal repeats, Long interspersed elements, Short interspersed elements, Germ line mutation, Inbred mice, Insertional mutagenesis, Transcriptional interference Background promoter and polyadenylation motifs and often a splice The mouse and human genomes harbor similar types of donor site [10, 11]. Sequences of full-length ERVs can TEs that have been discussed in many reviews, to which encode gag, pol and sometimes env, although groups of we refer the reader for more in depth and general infor- LTR retrotransposons with little or no retroviral hom- mation [1–9]. In general, both human and mouse con- ology also exist [6–9]. While not the subject of this re- tain ancient families of DNA transposons, none view, ERV LTRs can often act as cellular enhancers or currently active, which comprise 1–3% of these genomes promoters, creating chimeric transcripts with genes, and as well as many families or groups of retrotransposons, have been implicated in other regulatory functions [11– which have caused all the TE insertional mutations in 13]. -

Selective Androgen Receptor Modulators (Sarms) And
(19) TZZ ¥6B_T (11) EP 2 222 636 B1 (12) EUROPEAN PATENT SPECIFICATION (45) Date of publication and mention (51) Int Cl.: of the grant of the patent: C07D 207/08 (2006.01) C07D 207/09 (2006.01) 10.04.2013 Bulletin 2013/15 C07D 498/04 (2006.01) A61K 31/402 (2006.01) A61P 5/26 (2006.01) (21) Application number: 08865188.0 (86) International application number: (22) Date of filing: 12.12.2008 PCT/US2008/013657 (87) International publication number: WO 2009/082437 (02.07.2009 Gazette 2009/27) (54) SELECTIVE ANDROGEN RECEPTOR MODULATORS (SARMS) AND USES THEREOF SELEKTIVE ANDROGENREZEPTORMODULATOREN (SARMS) UND IHRE VERWENDUNG MODULATEURS SÉLECTIFS DU RÉCEPTEUR ANDROGÈNE (SARM) ET LEURS UTILISATIONS (84) Designated Contracting States: (74) Representative: Baldock, Sharon Claire AT BE BG CH CY CZ DE DK EE ES FI FR GB GR Boult Wade Tennant HR HU IE IS IT LI LT LU LV MC MT NL NO PL PT Verulam Gardens RO SE SI SK TR 70 Gray’s Inn Road Designated Extension States: London WC1X 8BT (GB) AL BA MK RS (56) References cited: (30) Priority: 21.12.2007 US 8731 P WO-A-02/068427 WO-A-03/090672 WO-A-2005/090282 WO-A-2006/124447 (43) Date of publication of application: WO-A-2007/015567 01.09.2010 Bulletin 2010/35 • HIGUCHI ET AL: "Potent, nonsteroidal selective (60) Divisional application: androgen receptor modulators (SARMs) based 12168231.4 / 2 489 656 on 8H-[1,4]oxazino[2,3-f]quinolin-8-ones" BIOORGANIC & MEDICINAL CHEMISTRY (73) Proprietor: Ligand Pharmaceuticals Inc. -

Genomic Amplification of Chromosome 20Q13.33 Is the Early Biomarker For
Bui et al. BMC Medical Genomics 2020, 13(Suppl 10):149 https://doi.org/10.1186/s12920-020-00776-z RESEARCH Open Access Genomic amplification of chromosome 20q13.33 is the early biomarker for the development of sporadic colorectal carcinoma Vo-Minh-Hoang Bui1,2, Clément Mettling3, Jonathan Jou4 and H. Sunny Sun1,5* From The 18th Asia Pacific Bioinformatics Conference Seoul, Korea. 18-20 August 2020 Abstract Background: Colorectal carcinoma (CRC) is the third most common cancer in the world and also the third leading cause of cancer-related mortality in Taiwan. CRC tumorigenesis is a multistep process, starting from mutations causing loss of function of tumor suppressor genes, canonically demonstrated in adenomatous polyposis coli pathogenesis. Although many genes or chromosomal alterations have been shown to be involved in this process, there are still unrecognized molecular events within CRC tumorigenesis. Elucidating these mechanisms may help improve the management and treatment. Methods: In this study, we aimed to identify copy number alteration of the smallest chromosomal regions that is significantly associated with sporadic CRC tumorigenesis using high-resolution array-based Comparative Genomic Hybridization (aCGH) and quantitative Polymerase chain reaction (qPCR). In addition, microsatellite instability assay and sequencing-based mutation assay were performed to illustrate the initiation event of CRC tumorigenesis. Results: A total of 571 CRC patients were recruited and 377 paired CRC tissues from sporadic CRC cases were used to define the smallest regions with chromosome copy number changes. In addition, 198 colorectal polyps from 160 patients were also used to study the role of 20q13.33 gain in CRC tumorigenesis. -

WO 2019/079361 Al 25 April 2019 (25.04.2019) W 1P O PCT
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date WO 2019/079361 Al 25 April 2019 (25.04.2019) W 1P O PCT (51) International Patent Classification: CA, CH, CL, CN, CO, CR, CU, CZ, DE, DJ, DK, DM, DO, C12Q 1/68 (2018.01) A61P 31/18 (2006.01) DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, C12Q 1/70 (2006.01) HR, HU, ID, IL, IN, IR, IS, JO, JP, KE, KG, KH, KN, KP, KR, KW, KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, (21) International Application Number: MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, PCT/US2018/056167 OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, (22) International Filing Date: SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, 16 October 2018 (16. 10.2018) TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (25) Filing Language: English (84) Designated States (unless otherwise indicated, for every kind of regional protection available): ARIPO (BW, GH, (26) Publication Language: English GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, ST, SZ, TZ, (30) Priority Data: UG, ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, RU, TJ, 62/573,025 16 October 2017 (16. 10.2017) US TM), European (AL, AT, BE, BG, CH, CY, CZ, DE, DK, EE, ES, FI, FR, GB, GR, HR, HU, ΓΕ , IS, IT, LT, LU, LV, (71) Applicant: MASSACHUSETTS INSTITUTE OF MC, MK, MT, NL, NO, PL, PT, RO, RS, SE, SI, SK, SM, TECHNOLOGY [US/US]; 77 Massachusetts Avenue, TR), OAPI (BF, BJ, CF, CG, CI, CM, GA, GN, GQ, GW, Cambridge, Massachusetts 02139 (US). -

Produktinformation
Produktinformation Diagnostik & molekulare Diagnostik Laborgeräte & Service Zellkultur & Verbrauchsmaterial Forschungsprodukte & Biochemikalien Weitere Information auf den folgenden Seiten! See the following pages for more information! Lieferung & Zahlungsart Lieferung: frei Haus Bestellung auf Rechnung SZABO-SCANDIC Lieferung: € 10,- HandelsgmbH & Co KG Erstbestellung Vorauskassa Quellenstraße 110, A-1100 Wien T. +43(0)1 489 3961-0 Zuschläge F. +43(0)1 489 3961-7 [email protected] • Mindermengenzuschlag www.szabo-scandic.com • Trockeneiszuschlag • Gefahrgutzuschlag linkedin.com/company/szaboscandic • Expressversand facebook.com/szaboscandic PSMD10 Antibody, Biotin conjugated Product Code CSB-PA018899LD01HU Abbreviation 26S proteasome non-ATPase regulatory subunit 10 Storage Upon receipt, store at -20°C or -80°C. Avoid repeated freeze. Uniprot No. O75832 Immunogen Recombinant Human 26S proteasome non-ATPase regulatory subunit 10 protein (1-226AA) Raised In Rabbit Species Reactivity Human Tested Applications ELISA Relevance Acts as a chaperone during the assembly of the 26S proteasome, specifically of the PA700/19S regulatory complex (RC). In the initial step of the base subcomplex assembly is part of an intermediate PSMD10:PSMC4:PSMC5:PAAF1 module which probably assembles with a PSMD5:PSMC2:PSMC1:PSMD2 module. Independently of the proteasome, regulates EGF-induced AKT activation through inhibition of the RHOA/ROCK/PTEN pahway, leading to prolonged AKT activation. Plays an important role in RAS-induced tumorigenesis. Acts as an proto-oncoprotein by being involved in negative regulation of tumor suppressors RB1 and p53/TP53. Overexpression is leading to phosphorylation of RB1 and proteasomal degradation of RB1. Regulates CDK4-mediated phosphorylation of RB1 by competing with CDKN2A for binding with CDK4. Facilitates binding of MDM2 to p53/TP53 and the mono- and polyubiquitination of p53/TP53 by MDM2 suggesting a function in targeting the TP53:MDM2 complex to the 26S proteasome. -

Cancer Vulnerabilities Unveiled by Genomic Loss
Cancer Vulnerabilities Unveiled by Genomic Loss The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation Nijhawan, Deepak, Travis I. Zack, Yin Ren, Matthew R. Strickland, Rebecca Lamothe, Steven E. Schumacher, Aviad Tsherniak, et al. “Cancer Vulnerabilities Unveiled by Genomic Loss.” Cell 150, no. 4 (August 2012): 842–854. © 2012 Elsevier Inc. As Published http://dx.doi.org/10.1016/j.cell.2012.07.023 Publisher Elsevier B.V. Version Final published version Citable link http://hdl.handle.net/1721.1/91899 Terms of Use Article is made available in accordance with the publisher's policy and may be subject to US copyright law. Please refer to the publisher's site for terms of use. Cancer Vulnerabilities Unveiled by Genomic Loss Deepak Nijhawan,1,2,7,9,10 Travis I. Zack,1,2,3,9 Yin Ren,5 Matthew R. Strickland,1 Rebecca Lamothe,1 Steven E. Schumacher,1,2 Aviad Tsherniak,2 Henrike C. Besche,4 Joseph Rosenbluh,1,2,7 Shyemaa Shehata,1 Glenn S. Cowley,2 Barbara A. Weir,2 Alfred L. Goldberg,4 Jill P. Mesirov,2 David E. Root,2 Sangeeta N. Bhatia,2,5,6,7,8 Rameen Beroukhim,1,2,7,* and William C. Hahn1,2,7,* 1Departments of Cancer Biology and Medical Oncology, Dana Farber Cancer Institute, Boston, MA 02215, USA 2Broad Institute of Harvard and MIT, Cambridge, MA 02142, USA 3Biophysics Program, Harvard University, Boston, MA 02115, USA 4Department of Cell Biology, Harvard Medical School, 240 Longwood Avenue, Boston, MA 02115, USA 5Harvard-MIT Division of Health Sciences and Technology 6David H. -

PSMD10 Antibody (Center) Purified Rabbit Polyclonal Antibody (Pab) Catalog # AW5126
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 PSMD10 Antibody (Center) Purified Rabbit Polyclonal Antibody (Pab) Catalog # AW5126 Specification PSMD10 Antibody (Center) - Product Information Application IF, WB, IHC-P,E Primary Accession O75832 Other Accession Q9Z2X2 Reactivity Human Predicted Mouse Host Rabbit Clonality Polyclonal Calculated MW H=24,16;M=25;Ra t=25 KDa Isotype Rabbit Ig Antigen Source HUMAN PSMD10 Antibody (Center) - Additional Information Gene ID 5716 Fluorescent image of Hela cells stained with PSMD10 Antibody (Center)(Cat#AW5126). Antigen Region AW5126 was diluted at 1:25 dilution. An 43-76 Alexa Fluor 488-conjugated goat anti-rabbit lgG at 1:400 dilution was used as the Other Names secondary antibody (green). Cytoplasmic 26S proteasome non-ATPase regulatory subunit 10, 26S proteasome regulatory actin was counterstained with Alexa Fluor® subunit p28, Gankyrin, p28(GANK), PSMD10 555 conjugated with Phalloidin (red). Dilution IF~~1:25 WB~~ 1:1000 IHC-P~~1:25 Target/Specificity This PSMD10 antibody is generated from a rabbit immunized with a KLH conjugated synthetic peptide between 43-76 amino acids from the Central region of human PSMD10. Format Purified polyclonal antibody supplied in PBS with 0.09% (W/V) sodium azide. This antibody is purified through a protein A column, followed by peptide affinity purification. Western blot analysis of lysates from MCF-7, PC-3, K562 cell line (from left to right), using Storage PSMD10 Antibody (Center)(Cat. #AW5126). Page 1/3 10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 Maintain refrigerated at 2-8°C for up to 2 AW5126 was diluted at 1:1000 at each lane. -

Proteomic and Bioinformatic Pipeline to Screen the Ligands of S
www.nature.com/scientificreports OPEN Proteomic and bioinformatic pipeline to screen the ligands of S. pneumoniae interacting Received: 15 September 2017 Accepted: 14 March 2018 with human brain microvascular Published: xx xx xxxx endothelial cells Irene Jiménez-Munguía1, Lucia Pulzova1, Evelina Kanova1, Zuzana Tomeckova1, Petra Majerova2, Katarina Bhide1, Lubos Comor1, Ivana Sirochmanova1, Andrej Kovac2 & Mangesh Bhide1,2 The mechanisms by which Streptococcus pneumoniae penetrates the blood-brain barrier (BBB), reach the CNS and causes meningitis are not fully understood. Adhesion of bacterial cells on the brain microvascular endothelial cells (BMECs), mediated through protein-protein interactions, is one of the crucial steps in translocation of bacteria across BBB. In this work, we proposed a systematic workfow for identifcation of cell wall associated ligands of pneumococcus that might adhere to the human BMECs. The proteome of S. pneumoniae was biotinylated and incubated with BMECs. Interacting proteins were recovered by afnity purifcation and identifed by data independent acquisition (DIA). A total of 44 proteins were identifed from which 22 were found to be surface-exposed. Based on the subcellular location, ontology, protein interactive analysis and literature review, fve ligands (adhesion lipoprotein, endo-β-N-acetylglucosaminidase, PhtA and two hypothetical proteins, Spr0777 and Spr1730) were selected to validate experimentally (ELISA and immunocytochemistry) the ligand- BMECs interaction. In this study, we proposed a high-throughput approach to generate a dataset of plausible bacterial ligands followed by systematic bioinformatics pipeline to categorize the protein candidates for experimental validation. The approach proposed here could contribute in the fast and reliable screening of ligands that interact with host cells. -

Identification and the Significance of Selective Proteins in Bile And
University of Arkansas, Fayetteville ScholarWorks@UARK Theses and Dissertations 12-2014 Identification and the Significance of Selective Proteins in Bile and Plasma of Normal and Health- compromised Chickens Balamurugan Packialakshmi University of Arkansas, Fayetteville Follow this and additional works at: http://scholarworks.uark.edu/etd Part of the Analytical Chemistry Commons, Animal Diseases Commons, and the Poultry or Avian Science Commons Recommended Citation Packialakshmi, Balamurugan, "Identification and the Significance of Selective Proteins in Bile and Plasma of Normal and Health- compromised Chickens" (2014). Theses and Dissertations. 2083. http://scholarworks.uark.edu/etd/2083 This Dissertation is brought to you for free and open access by ScholarWorks@UARK. It has been accepted for inclusion in Theses and Dissertations by an authorized administrator of ScholarWorks@UARK. For more information, please contact [email protected], [email protected]. Identification and the Significance of Selective Proteins in Bile and Plasma of Normal and Health-compromised Chickens Identification and the Significance of Selective Proteins in Bile and Plasma of Normal and Health-compromised Chickens A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in Cell and Molecular Biology by Balamurugan Packialakshmi Tamilnadu Agricultural University, Bachelor of Technology in Agricultural Biotechnology, 2007 Govind Ballabh Pant University of Agriculture and Technology, Master of Science in Molecular Biology and Biotechnology, 2009 December 2014 University of Arkansas This dissertation is approved for the recommendation to the graduate council. _____________________ Dr. Narayan. C. Rath Dissertation Director _____________________ _____________________ Dr. Jackson O. Lay, Jr. Dr. Robert F. Wideman, Jr. Committee member Committee member ____________________ Dr. -
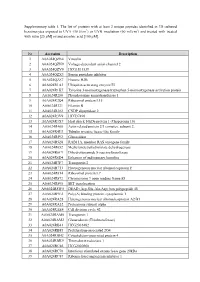
Supplementary Table 1. the List of Proteins with at Least 2 Unique
Supplementary table 1. The list of proteins with at least 2 unique peptides identified in 3D cultured keratinocytes exposed to UVA (30 J/cm2) or UVB irradiation (60 mJ/cm2) and treated with treated with rutin [25 µM] or/and ascorbic acid [100 µM]. Nr Accession Description 1 A0A024QZN4 Vinculin 2 A0A024QZN9 Voltage-dependent anion channel 2 3 A0A024QZV0 HCG1811539 4 A0A024QZX3 Serpin peptidase inhibitor 5 A0A024QZZ7 Histone H2B 6 A0A024R1A3 Ubiquitin-activating enzyme E1 7 A0A024R1K7 Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein 8 A0A024R280 Phosphoserine aminotransferase 1 9 A0A024R2Q4 Ribosomal protein L15 10 A0A024R321 Filamin B 11 A0A024R382 CNDP dipeptidase 2 12 A0A024R3V9 HCG37498 13 A0A024R3X7 Heat shock 10kDa protein 1 (Chaperonin 10) 14 A0A024R408 Actin related protein 2/3 complex, subunit 2, 15 A0A024R4U3 Tubulin tyrosine ligase-like family 16 A0A024R592 Glucosidase 17 A0A024R5Z8 RAB11A, member RAS oncogene family 18 A0A024R652 Methylenetetrahydrofolate dehydrogenase 19 A0A024R6C9 Dihydrolipoamide S-succinyltransferase 20 A0A024R6D4 Enhancer of rudimentary homolog 21 A0A024R7F7 Transportin 2 22 A0A024R7T3 Heterogeneous nuclear ribonucleoprotein F 23 A0A024R814 Ribosomal protein L7 24 A0A024R872 Chromosome 9 open reading frame 88 25 A0A024R895 SET translocation 26 A0A024R8W0 DEAD (Asp-Glu-Ala-Asp) box polypeptide 48 27 A0A024R9E2 Poly(A) binding protein, cytoplasmic 1 28 A0A024RA28 Heterogeneous nuclear ribonucleoprotein A2/B1 29 A0A024RA52 Proteasome subunit alpha 30 A0A024RAE4 Cell division cycle 42 31