Peroxisome Proliferator-Activated Receptor Alpha: Insight Into the Structure, Function and Energy Homeostasis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

An Animal Model with a Cardiomyocyte-Specific Deletion of Estrogen Receptor Alpha: Functional, Metabolic, and Differential Netwo
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2014 An animal model with a cardiomyocyte-specific deletion of estrogen receptor alpha: Functional, metabolic, and differential network analysis Sriram Devanathan Washington University School of Medicine in St. Louis Timothy Whitehead Washington University School of Medicine in St. Louis George G. Schweitzer Washington University School of Medicine in St. Louis Nicole Fettig Washington University School of Medicine in St. Louis Attila Kovacs Washington University School of Medicine in St. Louis See next page for additional authors Follow this and additional works at: https://digitalcommons.wustl.edu/open_access_pubs Recommended Citation Devanathan, Sriram; Whitehead, Timothy; Schweitzer, George G.; Fettig, Nicole; Kovacs, Attila; Korach, Kenneth S.; Finck, Brian N.; and Shoghi, Kooresh I., ,"An animal model with a cardiomyocyte-specific deletion of estrogen receptor alpha: Functional, metabolic, and differential network analysis." PLoS One.9,7. e101900. (2014). https://digitalcommons.wustl.edu/open_access_pubs/3326 This Open Access Publication is brought to you for free and open access by Digital Commons@Becker. It has been accepted for inclusion in Open Access Publications by an authorized administrator of Digital Commons@Becker. For more information, please contact [email protected]. Authors Sriram Devanathan, Timothy Whitehead, George G. Schweitzer, Nicole Fettig, Attila Kovacs, Kenneth S. Korach, Brian N. Finck, and Kooresh I. Shoghi This open access publication is available at Digital Commons@Becker: https://digitalcommons.wustl.edu/open_access_pubs/3326 An Animal Model with a Cardiomyocyte-Specific Deletion of Estrogen Receptor Alpha: Functional, Metabolic, and Differential Network Analysis Sriram Devanathan1, Timothy Whitehead1, George G. Schweitzer2, Nicole Fettig1, Attila Kovacs3, Kenneth S. -

Dualism of Peroxisome Proliferator-Activated Receptor Α/Γ: a Potent Clincher in Insulin Resistance
AEGAEUM JOURNAL ISSN NO: 0776-3808 Dualism of Peroxisome Proliferator-Activated Receptor α/γ: A Potent Clincher in Insulin Resistance Mr. Ravikumar R. Thakar1 and Dr. Nilesh J. Patel1* 1Faculty of Pharmacy, Shree S. K. Patel College of Pharmaceutical Education & Research, Ganpat University, Gujarat, India. [email protected] Abstract: Diabetes mellitus is clinical syndrome which is signalised by augmenting level of sugar in blood stream, which produced through lacking of insulin level and defective insulin activity or both. As per worldwide epidemiology data suggested that the numbers of people with T2DM living in developing countries is increasing with 80% of people with T2DM. Peroxisome proliferator-activated receptors are a family of ligand-activated transcription factors; modulate the expression of many genes. PPARs have three isoforms namely PPARα, PPARβ/δ and PPARγ that play a central role in regulating glucose, lipid and cholesterol metabolism where imbalance can lead to obesity, T2DM and CV ailments. It have pathogenic role in diabetes. PPARα is regulates the metabolism of lipids, carbohydrates, and amino acids, activated by ligands such as polyunsaturated fatty acids, and drugs used as Lipid lowering agents. PPAR β/δ could envision as a therapeutic option for the correction of diabetes and a variety of inflammatory conditions. PPARγ is well categorized, an element of the PPARs, also pharmacological effective as an insulin resistance lowering agents, are used as a remedy for insulin resistance integrated with type- 2 diabetes mellitus. There are mechanistic role of PPARα, PPARβ/δ and PPARγ in diabetes mellitus and insulin resistance. From mechanistic way, it revealed that dual PPAR-α/γ agonist play important role in regulating both lipids as well as glycemic levels with essential safety issues. -

Dimerization Specificity of Myogenic Helix-Loop-Helix DNA-Binding Factors Directed by Nonconserved Hydrophilic Residues
Downloaded from genesdev.cshlp.org on October 3, 2021 - Published by Cold Spring Harbor Laboratory Press Dimerization specificity of myogenic helix-loop-helix DNA-binding factors directed by nonconserved hydrophilic residues Masaki Shirakata, Fred K. Friedman, 1 Qin Wei, and Bruce M. Paterson Laboratory of Biochemistry, ~Laboratory of Molecular Carcinogenesis, National Cancer Institute, National Institutes of Health, Bethesda, Maryland 20892 USA The myogenic regulatory factor MyoD dimerizes with other positive and negative regulatory factors through a conserved region called the helix-loop-helix (HLH) domain. Using a non-DNA-binding MyoD mutant with a normal HLH domain as a dimerization competitor in gel mobility shift assays in conjunction with various MyoD HLH mutants, nonhydrophobic amino acids were identified in the HLH domain that contribute to dimerization specificity with El2. The assay detected subtle differences in dimerization activity among the mutant MyoD proteins that correlated with their ability to activate transcription in vivo, but this correlation was not apparent in the absence of competitor. The identification of such nonhydrophobic residues enabled us to predict the differences in dimerization affinity among the four vertebrate myogenic factors with El2. The experiments confirmed the prediction. Furthermore, a high-affinity homodimerizing analog of MyoD was designed by a single substitution at one of these residue positions. These experimental results were strengthened when they were analyzed in terms of the crystal structure for the Max bHLHZip domain homodimer. This analysis has allowed us to identify those residues that form charged residue pairs between the two HLH domains of MyoD and El2 and determine the dimerization specificity of the bHLH proteins. -

Leucine Zippers
Leucine Zippers Leucine Zippers Advanced article Toshio Hakoshima, Nara Institute of Science and Technology, Nara, Japan Article contents Introduction The leucine zipper (ZIP) motif consists of a periodic repetition of a leucine residue at every Structural Basis of ZIP seventh position and forms an a-helical conformation, which facilitates dimerization and in Occurrence of ZIP and Coiled-coil Motifs some cases higher oligomerization of proteins. In many eukaryotic gene regulatory proteins, Dimerization Specificity of ZIP the ZIP motif is flanked at its N-terminus by a basic region containing characteristic residues DNA-binding Specificity of bZIP that facilitate DNA binding. doi: 10.1038/npg.els.0005049 Introduction protein modules for protein–protein interactions. Knowing the structure and function of these motifs A structure referred to as the leucine zipper or enables us to understand the molecular recognition simply as ZIP has been proposed to explain how a system in several biological processes. class of eukaryotic gene regulatory proteins works (Landschulz et al., 1988). A segment of the mammalian CCAAT/enhancer binding protein (C/EBP) of 30 Structural Basis of ZIP amino acids shares notable sequence similarity with a segment of the cellular Myc transforming protein. The The a helix is a secondary structure element that segments have been found to contain a periodic occurs frequently in proteins. Alpha helices are repetition of a leucine residue at every seventh stabilized in proteins by being packed into the position. A periodic array of at least four leucines hydrophobic core of a protein through hydrophobic has also been noted in the sequences of the Fos and side chains. -

Synthesis of a Coumarin-Based PPAR Fluorescence Probe for Competitive Binding Assay
International Journal of Molecular Sciences Article Synthesis of a Coumarin-Based PPARγ Fluorescence Probe for Competitive Binding Assay Chisato Yoshikawa, Hiroaki Ishida , Nami Ohashi and Toshimasa Itoh * Laboratory of Drug Design and Medicinal Chemistry, Showa Pharmaceutical University, 3-3165 Higashi-Tamagawagakuen, Machida, Tokyo 194-8543, Japan; [email protected] (C.Y.); [email protected] (H.I.); [email protected] (N.O.) * Correspondence: [email protected] Abstract: Peroxisome proliferator-activated receptor γ (PPARγ) is a molecular target of metabolic syndrome and inflammatory disease. PPARγ is an important nuclear receptor and numerous PPARγ ligands were developed to date; thus, efficient assay methods are important. Here, we investigated the incorporation of 7-diethylamino coumarin into the PPARγ agonist rosiglitazone and used the com- pound in a binding assay for PPARγ. PPARγ-ligand-incorporated 7-methoxycoumarin, 1, showed weak fluorescence intensity in a previous report. We synthesized PPARγ-ligand-incorporating coumarin, 2, in this report, and it enhanced the fluorescence intensity. The PPARγ ligand 2 main- tained the rosiglitazone activity. The obtained partial agonist 6 appeared to act through a novel mechanism. The fluorescence intensity of 2 and 6 increased by binding to the ligand binding domain (LBD) of PPARγ and the affinity of reported PPARγ ligands were evaluated using the probe. Keywords: PPARγ ligand; coumarin; fluorescent ligand; screening; crystal structure Citation: Yoshikawa, C.; Ishida, H.; Ohashi, N.; Itoh, T. Synthesis of a Coumarin-Based PPARγ 1. Introduction Fluorescence Probe for Competitive Binding Assay. Int. J. Mol. Sci. 2021, Peroxisome proliferator-activated receptors (PPARs) belong to the nuclear receptor su- 22, 4034. -

NATURAL KILLER CELLS, HYPOXIA, and EPIGENETIC REGULATION of HEMOCHORIAL PLACENTATION by Damayanti Chakraborty Submitted to the G
NATURAL KILLER CELLS, HYPOXIA, AND EPIGENETIC REGULATION OF HEMOCHORIAL PLACENTATION BY Damayanti Chakraborty Submitted to the graduate degree program in Pathology and Laboratory Medicine and the Graduate Faculty of the University of Kansas in partial fulfillment ofthe requirements for the degree of Doctor of Philosophy. ________________________________ Chair: Michael J. Soares, Ph.D. ________________________________ Jay Vivian, Ph.D. ________________________________ Patrick Fields, Ph.D. ________________________________ Soumen Paul, Ph.D. ________________________________ Michael Wolfe, Ph.D. ________________________________ Adam J. Krieg, Ph.D. Date Defended: 04/01/2013 The Dissertation Committee for Damayanti Chakraborty certifies that this is the approved version of the following dissertation: NATURAL KILLER CELLS, HYPOXIA, AND EPIGENETIC REGULATION OF HEMOCHORIAL PLACENTATION ________________________________ Chair: Michael J. Soares, Ph.D. Date approved: 04/01/2013 ii ABSTRACT During the establishment of pregnancy, uterine stromal cells differentiate into decidual cells and recruit natural killer (NK) cells. These NK cells are characterized by low cytotoxicity and distinct cytokine production. In rodent as well as in human pregnancy, the uterine NK cells peak in number around mid-gestation after which they decline. NK cells associate with uterine spiral arteries and are implicated in pregnancy associated vascular remodeling processes and potentially in modulating trophoblast invasion. Failure of trophoblast invasion and vascular remodeling has been shown to be associated with pathological conditions like preeclampsia syndrome, hypertension in mother and/or fetal growth restriction. We hypothesize that NK cells fundamentally contribute to the organization of the placentation site. In order to study the in vivo role of NK cells during pregnancy, gestation stage- specific NK cell depletion was performed in rats using anti asialo GM1 antibodies. -

140503 IPF Signatures Supplement Withfigs Thorax
Supplementary material for Heterogeneous gene expression signatures correspond to distinct lung pathologies and biomarkers of disease severity in idiopathic pulmonary fibrosis Daryle J. DePianto1*, Sanjay Chandriani1⌘*, Alexander R. Abbas1, Guiquan Jia1, Elsa N. N’Diaye1, Patrick Caplazi1, Steven E. Kauder1, Sabyasachi Biswas1, Satyajit K. Karnik1#, Connie Ha1, Zora Modrusan1, Michael A. Matthay2, Jasleen Kukreja3, Harold R. Collard2, Jackson G. Egen1, Paul J. Wolters2§, and Joseph R. Arron1§ 1Genentech Research and Early Development, South San Francisco, CA 2Department of Medicine, University of California, San Francisco, CA 3Department of Surgery, University of California, San Francisco, CA ⌘Current address: Novartis Institutes for Biomedical Research, Emeryville, CA. #Current address: Gilead Sciences, Foster City, CA. *DJD and SC contributed equally to this manuscript §PJW and JRA co-directed this project Address correspondence to Paul J. Wolters, MD University of California, San Francisco Department of Medicine Box 0111 San Francisco, CA 94143-0111 [email protected] or Joseph R. Arron, MD, PhD Genentech, Inc. MS 231C 1 DNA Way South San Francisco, CA 94080 [email protected] 1 METHODS Human lung tissue samples Tissues were obtained at UCSF from clinical samples from IPF patients at the time of biopsy or lung transplantation. All patients were seen at UCSF and the diagnosis of IPF was established through multidisciplinary review of clinical, radiological, and pathological data according to criteria established by the consensus classification of the American Thoracic Society (ATS) and European Respiratory Society (ERS), Japanese Respiratory Society (JRS), and the Latin American Thoracic Association (ALAT) (ref. 5 in main text). Non-diseased normal lung tissues were procured from lungs not used by the Northern California Transplant Donor Network. -

The Function and Evolution of C2H2 Zinc Finger Proteins and Transposons
The function and evolution of C2H2 zinc finger proteins and transposons by Laura Francesca Campitelli A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy Department of Molecular Genetics University of Toronto © Copyright by Laura Francesca Campitelli 2020 The function and evolution of C2H2 zinc finger proteins and transposons Laura Francesca Campitelli Doctor of Philosophy Department of Molecular Genetics University of Toronto 2020 Abstract Transcription factors (TFs) confer specificity to transcriptional regulation by binding specific DNA sequences and ultimately affecting the ability of RNA polymerase to transcribe a locus. The C2H2 zinc finger proteins (C2H2 ZFPs) are a TF class with the unique ability to diversify their DNA-binding specificities in a short evolutionary time. C2H2 ZFPs comprise the largest class of TFs in Mammalian genomes, including nearly half of all Human TFs (747/1,639). Positive selection on the DNA-binding specificities of C2H2 ZFPs is explained by an evolutionary arms race with endogenous retroelements (EREs; copy-and-paste transposable elements), where the C2H2 ZFPs containing a KRAB repressor domain (KZFPs; 344/747 Human C2H2 ZFPs) are thought to diversify to bind new EREs and repress deleterious transposition events. However, evidence of the gain and loss of KZFP binding sites on the ERE sequence is sparse due to poor resolution of ERE sequence evolution, despite the recent publication of binding preferences for 242/344 Human KZFPs. The goal of my doctoral work has been to characterize the Human C2H2 ZFPs, with specific interest in their evolutionary history, functional diversity, and coevolution with LINE EREs. -

Tgfβ-Regulated Gene Expression by Smads and Sp1/KLF-Like Transcription Factors in Cancer VOLKER ELLENRIEDER
ANTICANCER RESEARCH 28 : 1531-1540 (2008) Review TGFβ-regulated Gene Expression by Smads and Sp1/KLF-like Transcription Factors in Cancer VOLKER ELLENRIEDER Signal Transduction Laboratory, Internal Medicine, Department of Gastroenterology and Endocrinology, University of Marburg, Marburg, Germany Abstract. Transforming growth factor beta (TGF β) controls complex induces the canonical Smad signaling molecules which vital cellular functions through its ability to regulate gene then translocate into the nucleus to regulate transcription (2). The expression. TGFβ binding to its transmembrane receptor cellular response to TGF β can be extremely variable depending kinases initiates distinct intracellular signalling cascades on the cell type and the activation status of a cell at a given time. including the Smad signalling and transcription factors and also For instance, TGF β induces growth arrest and apoptosis in Smad-independent pathways. In normal epithelial cells, TGF β healthy epithelial cells, whereas it can also promote tumor stimulation induces a cytostatic program which includes the progression through stimulation of cell proliferation and the transcriptional repression of the c-Myc oncogene and the later induction of an epithelial-to-mesenchymal transition of tumor induction of the cell cycle inhibitors p15 INK4b and p21 Cip1 . cells (1, 3). In the last decade it has become clear that both the During carcinogenesis, however, many tumor cells lose their tumor suppressing and the tumor promoting functions of TGF β ability to respond to TGF β with growth inhibition, and instead, are primarily regulated on the level of gene expression through activate genes involved in cell proliferation, invasion and Smad-dependent and -independent mechanisms (1, 2, 4). -

The Opportunities and Challenges of Peroxisome Proliferator-Activated Receptors Ligands in Clinical Drug Discovery and Development
International Journal of Molecular Sciences Review The Opportunities and Challenges of Peroxisome Proliferator-Activated Receptors Ligands in Clinical Drug Discovery and Development Fan Hong 1,2, Pengfei Xu 1,*,† and Yonggong Zhai 1,2,* 1 Beijing Key Laboratory of Gene Resource and Molecular Development, College of Life Sciences, Beijing Normal University, Beijing 100875, China; [email protected] 2 Key Laboratory for Cell Proliferation and Regulation Biology of State Education Ministry, College of Life Sciences, Beijing Normal University, Beijing 100875, China * Correspondence: [email protected] (P.X.); [email protected] (Y.Z.); Tel.: +86-156-005-60991 (P.X.); +86-10-5880-6656 (Y.Z.) † Current address: Center for Pharmacogenetics and Department of Pharmaceutical Sciences, University of Pittsburgh, Pittsburgh, PA 15213, USA. Received: 22 June 2018; Accepted: 24 July 2018; Published: 27 July 2018 Abstract: Peroxisome proliferator-activated receptors (PPARs) are a well-known pharmacological target for the treatment of multiple diseases, including diabetes mellitus, dyslipidemia, cardiovascular diseases and even primary biliary cholangitis, gout, cancer, Alzheimer’s disease and ulcerative colitis. The three PPAR isoforms (α, β/δ and γ) have emerged as integrators of glucose and lipid metabolic signaling networks. Typically, PPARα is activated by fibrates, which are commonly used therapeutic agents in the treatment of dyslipidemia. The pharmacological activators of PPARγ include thiazolidinediones (TZDs), which are insulin sensitizers used in the treatment of type 2 diabetes mellitus (T2DM), despite some drawbacks. In this review, we summarize 84 types of PPAR synthetic ligands introduced to date for the treatment of metabolic and other diseases and provide a comprehensive analysis of the current applications and problems of these ligands in clinical drug discovery and development. -
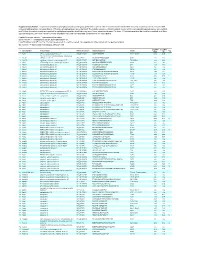
Supplementary Table 1
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding -

Regulation of Enac-Mediated Sodium Reabsorption by Peroxisome Proliferator-Activated Receptors
Hindawi Publishing Corporation PPAR Research Volume 2010, Article ID 703735, 9 pages doi:10.1155/2010/703735 Review Article Regulation of ENaC-Mediated Sodium Reabsorption by Peroxisome Proliferator-Activated Receptors Tengis S. Pavlov,1 John D. Imig,2, 3 and Alexander Staruschenko1, 4 1 Department of Physiology, Medical College of Wisconsin, Milwaukee, WI 53226, USA 2 Department of Pharmacology and Toxicology, Medical College of Wisconsin, Milwaukee, WI 53226, USA 3 Cardiovascular Research Center, Medical College of Wisconsin, Milwaukee, WI 53226, USA 4 Kidney Disease Center, Medical College of Wisconsin, Milwaukee, WI 53226, USA Correspondence should be addressed to Alexander Staruschenko, [email protected] Received 6 January 2010; Revised 16 March 2010; Accepted 14 April 2010 Academic Editor: Tianxin Yang Copyright © 2010 Tengis S. Pavlov et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Peroxisome proliferator-activated receptors (PPARs) are members of a steroid hormone receptor superfamily that responds to changes in lipid and glucose homeostasis. Peroxisomal proliferator-activated receptor subtype γ (PPARγ)hasreceivedmuch attention as the target for antidiabetic drugs, as well as its role in responding to endogenous compounds such as prostaglandin J2. However, thiazolidinediones (TZDs), the synthetic agonists of the PPARγ are tightly associated with fluid retention and edema, as potentially serious side effects. The epithelial sodium channel (ENaC) represents the rate limiting step for sodium absorption in the renal collecting duct. Consequently, ENaC is a central effector impacting systemic blood volume and pressure. The role of PPARγ agonists on ENaC activity remains controversial.