Reference Genomes and Gene Models Impact on Variant Interpretation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Expert Curation of the Human and Mouse Olfactory Receptor Gene Repertoires Identifies Conserved Coding Regions Split Across Two Exons
bioRxiv preprint doi: https://doi.org/10.1101/774612; this version posted October 30, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Expert Curation of the Human and Mouse Olfactory Receptor Gene Repertoires Identifies Conserved Coding Regions Split Across Two Exons 5 If H. A. Barnes1†#, Ximena Ibarra-Soria2,3†#, Stephen Fitzgerald3, Jose M. Gonzalez1, Claire Davidson1, Matthew P. Hardy1, Deepa Manthravadi4, Laura Van Gerven5, Mark Jorissen5, Zhen Zeng6, Mona Khan6, Peter Mombaerts6, Jennifer Harrow7, Darren W. Logan3,8,9 and Adam Frankish1#. 10 1. European Molecular Biology Laboratory, European Bioinformatics Institute, Wellcome Genome Campus, Hinxton, Cambridge, CB10 1SD, UK. 2. Cancer Research UK Cambridge Institute, University of Cambridge, Li Ka Shing Centre, Robinson Way, Cambridge, CB2 0RE, UK. 3. Wellcome Sanger Institute, Wellcome Genome Campus, Hinxton, Cambridge, CB10 1SA, UK. 15 4. Brandeis University, 415 South Street, Waltham, MA 02453, USA. 5. Department of ENT-HNS, UZ Leuven, Herestraat 49, 3000 Leuven, Belgium. 6. Max Planck Research Unit for Neurogenetics, Max von-Laue-Strasse 4, 60438 Frankfurt, Germany. 7. ELIXIR, Wellcome Genome Campus, Hinxton, Cambridge, CB10 1SD, UK. 8. Monell Chemical Senses Center, Philadelphia, PA 19104, USA. 20 9. Waltham Centre for Pet Nutrition, Leicestershire, LE14 4RT, UK. † These authors contributed equally to this work. # To whom correspondence should be addressed. Email: [email protected], [email protected] and [email protected]. -

Repetitive Elements in Humans
International Journal of Molecular Sciences Review Repetitive Elements in Humans Thomas Liehr Institute of Human Genetics, Jena University Hospital, Friedrich Schiller University, Am Klinikum 1, D-07747 Jena, Germany; [email protected] Abstract: Repetitive DNA in humans is still widely considered to be meaningless, and variations within this part of the genome are generally considered to be harmless to the carrier. In contrast, for euchromatic variation, one becomes more careful in classifying inter-individual differences as meaningless and rather tends to see them as possible influencers of the so-called ‘genetic background’, being able to at least potentially influence disease susceptibilities. Here, the known ‘bad boys’ among repetitive DNAs are reviewed. Variable numbers of tandem repeats (VNTRs = micro- and minisatellites), small-scale repetitive elements (SSREs) and even chromosomal heteromorphisms (CHs) may therefore have direct or indirect influences on human diseases and susceptibilities. Summarizing this specific aspect here for the first time should contribute to stimulating more research on human repetitive DNA. It should also become clear that these kinds of studies must be done at all available levels of resolution, i.e., from the base pair to chromosomal level and, importantly, the epigenetic level, as well. Keywords: variable numbers of tandem repeats (VNTRs); microsatellites; minisatellites; small-scale repetitive elements (SSREs); chromosomal heteromorphisms (CHs); higher-order repeat (HOR); retroviral DNA 1. Introduction Citation: Liehr, T. Repetitive In humans, like in other higher species, the genome of one individual never looks 100% Elements in Humans. Int. J. Mol. Sci. alike to another one [1], even among those of the same gender or between monozygotic 2021, 22, 2072. -

GENCODE: the Reference Human Genome Annotation for the ENCODE Project
Downloaded from genome.cshlp.org on September 26, 2012 - Published by Cold Spring Harbor Laboratory Press Resource GENCODE: The reference human genome annotation for The ENCODE Project Jennifer Harrow,1,9 Adam Frankish,1 Jose M. Gonzalez,1 Electra Tapanari,1 Mark Diekhans,2 Felix Kokocinski,1 Bronwen L. Aken,1 Daniel Barrell,1 Amonida Zadissa,1 Stephen Searle,1 If Barnes,1 Alexandra Bignell,1 Veronika Boychenko,1 Toby Hunt,1 Mike Kay,1 Gaurab Mukherjee,1 Jeena Rajan,1 Gloria Despacio-Reyes,1 Gary Saunders,1 Charles Steward,1 Rachel Harte,2 Michael Lin,3 Ce´dric Howald,4 Andrea Tanzer,5 Thomas Derrien,4 Jacqueline Chrast,4 Nathalie Walters,4 Suganthi Balasubramanian,6 Baikang Pei,6 Michael Tress,7 Jose Manuel Rodriguez,7 Iakes Ezkurdia,7 Jeltje van Baren,8 Michael Brent,8 David Haussler,2 Manolis Kellis,3 Alfonso Valencia,7 Alexandre Reymond,4 Mark Gerstein,6 Roderic Guigo´,5 and Tim J. Hubbard1,9 1Wellcome Trust Sanger Institute, Wellcome Trust Campus, Hinxton, Cambridge CB10 1SA, United Kingdom; 2University of California, Santa Cruz, California 95064, USA; 3Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA; 4Center for Integrative Genomics, University of Lausanne, 1015 Lausanne, Switzerland; 5Centre for Genomic Regulation (CRG) and UPF, 08003 Barcelona, Catalonia, Spain; 6Yale University, New Haven, Connecticut 06520-8047, USA; 7Spanish National Cancer Research Centre (CNIO), E-28029 Madrid, Spain; 8Center for Genome Sciences & Systems Biology, St. Louis, Missouri 63130, USA The GENCODE Consortium aims to identify all gene features in the human genome using a combination of computa- tional analysis, manual annotation, and experimental validation. -

GENCODE Reference Annotation for the Human and Mouse Genomes
D766–D773 Nucleic Acids Research, 2019, Vol. 47, Database issue Published online 24 October 2018 doi: 10.1093/nar/gky955 GENCODE reference annotation for the human and mouse genomes Adam Frankish1, Mark Diekhans2, Anne-Maud Ferreira3, Rory Johnson4,5, Irwin Jungreis 6,7, Jane Loveland 1, Jonathan M. Mudge1, Cristina Sisu8,9, Downloaded from https://academic.oup.com/nar/article-abstract/47/D1/D766/5144133 by Universite and EPFL Lausanne user on 11 April 2019 James Wright10, Joel Armstrong2, If Barnes1, Andrew Berry1, Alexandra Bignell1, Silvia Carbonell Sala11, Jacqueline Chrast3, Fiona Cunningham 1,Tomas´ Di Domenico 12, Sarah Donaldson1, Ian T. Fiddes2, Carlos Garc´ıa Giron´ 1, Jose Manuel Gonzalez1, Tiago Grego1, Matthew Hardy1, Thibaut Hourlier 1, Toby Hunt1, Osagie G. Izuogu1, Julien Lagarde11, Fergal J. Martin 1, Laura Mart´ınez12, Shamika Mohanan1, Paul Muir13,14, Fabio C.P. Navarro8, Anne Parker1, Baikang Pei8, Fernando Pozo12, Magali Ruffier 1, Bianca M. Schmitt1, Eloise Stapleton1, Marie-Marthe Suner 1, Irina Sycheva1, Barbara Uszczynska-Ratajczak15,JinuriXu8, Andrew Yates1, Daniel Zerbino 1, Yan Zhang8,16, Bronwen Aken1, Jyoti S. Choudhary10, Mark Gerstein8,17,18, Roderic Guigo´ 11,19, Tim J.P. Hubbard20, Manolis Kellis6,7, Benedict Paten2, Alexandre Reymond3, Michael L. Tress12 and Paul Flicek 1,* 1European Molecular Biology Laboratory, European Bioinformatics Institute, Wellcome Genome Campus, Hinxton, Cambridge CB10 1SD, UK, 2UC Santa Cruz Genomics Institute, University of California, Santa Cruz, Santa Cruz, CA 95064, USA, -

Downloaded from the Tranche Distributed File System (Tranche.Proteomecommons.Org) and Ftp://Ftp.Thegpm.Org/Data/Msms
Research Article Title: The shrinking human protein coding complement: are there now fewer than 20,000 genes? Authors: Iakes Ezkurdia1*, David Juan2*, Jose Manuel Rodriguez3, Adam Frankish4, Mark Diekhans5, Jennifer Harrow4, Jesus Vazquez 6, Alfonso Valencia2,3, Michael L. Tress2,*. Affiliations: 1. Unidad de Proteómica, Centro Nacional de Investigaciones Cardiovasculares, CNIC, Melchor Fernández Almagro, 3, rid, 28029, MadSpain 2. Structural Biology and Bioinformatics Programme, Spanish National Cancer Research Centre (CNIO), Melchor Fernández Almagro, 3, 28029, Madrid, Spain 3. National Bioinformatics Institute (INB), Spanish National Cancer Research Centre (CNIO), Melchor Fernández Almagro, 3, 28029, Madrid, Spain 4. Wellcome Trust Sanger Institute, Wellcome Trust Campus, Hinxton, Cambridge CB10 1SA, UK 5. Center for Biomolecular Science and Engineering, School of Engineering, University of California Santa Cruz (UCSC), 1156 High Street, Santa Cruz, CA 95064, USA 6. Laboratorio de Proteómica Cardiovascular, Centro Nacional de Investigaciones Cardiovasculares, CNIC, Melchor Fernández Almagro, 3, 28029, Madrid, Spain *: these two authors wish to be considered as joint first authors of the paper. Corresponding author: Michael Tress, [email protected], Tel: +34 91 732 80 00 Fax: +34 91 224 69 76 Running title: Are there fewer than 20,000 protein-coding genes? Keywords: Protein coding genes, proteomics, evolution, genome annotation Abstract Determining the full complement of protein-coding genes is a key goal of genome annotation. The most powerful approach for confirming protein coding potential is the detection of cellular protein expression through peptide mass spectrometry experiments. Here we map the peptides detected in 7 large-scale proteomics studies to almost 60% of the protein coding genes in the GENCODE annotation the human genome. -
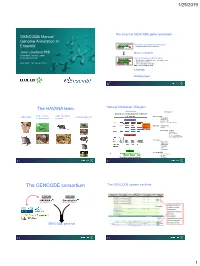
The GENCODE Consortium the GENCODE Update Trackhub
1/25/2019 The status of GENCODE gene annotation GENCODE Manual Genome Annotation in Reference genebuild ‘First pass’ systematic chr annotation Ensembl • Analysis of cDNA, ESTs, build genes Jane Loveland PhD Mouse complete Annotation Project Leader Ensembl-HAVANA Maturing genebuild Targeted improvement of models • Identification of additional gene, transcripts, exons • ‘Completion’ of models th PAG XXVII, 13 January 2019 • Correct functional annotation Major focus of human work TAGENE MANE project The HAVANA team Manual Annotation: Biotypes Annotation: Biotypes based on transcriptional evidence Whole Genome Targeted regions GENCODE Community projects Protein Coding or chromosome or genes Known_CDS Novel_CDS Putative_CDS Nonsense_mediated_decay Sequences from Transcript retained intron databases putative Non-coding lincRNA Antisense Sense_intronic Sense_overlapping 3’_overlapping_ncRNA Pseudogene Processed Unprocessed Transcribed Translated Unitary Polymorphic Immunoglobulin IG_pseudogene IG_Gene Structural and functional TR_Gene The GENCODE consortium The GENCODE update trackhub HAVANA Genebuild Manual annotation Computational annotation GENCODE gene set 1 1/25/2019 Ensembl gene view: ENO1 updated annotation (more transcripts) Walking across the mouse genome Updated annotation Walking across the mouse genome Walking across the mouse genome Walking across the mouse genome Walking across the mouse genome 2 1/25/2019 Walking across the mouse genome The GENCODE consortium current gene counts Human Mouse Total No of Transcripts 206694 141283 -

High-Throughput Annotation of Full-Length Long Noncoding
bioRxiv preprint doi: https://doi.org/10.1101/105064; this version posted September 4, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 2 High-throughput annotation of full-length 3 long noncoding RNAs with Capture Long- 4 Read Sequencing (CLS) 5 6 Authors 7 Julien Lagarde*1,2, Barbara Uszczynska-Ratajczak*1,2,6, Silvia Carbonell3, Sílvia Pérez-Lluch1,2, 8 Amaya Abad1,2, Carrie Davis4, Thomas R. Gingeras4, Adam Frankish5, Jennifer Harrow5,7, 9 Roderic Guigo#1,2, Rory Johnson#1,2,8 10 * Equal contribution 11 # Corresponding authors: [email protected], [email protected] 12 Author affiliations 13 1 Centre for Genomic Regulation (CRG), Barcelona Institute of Science and Technology (BIST), 14 Dr. Aiguader 88, 08003 Barcelona, Spain. 15 2 Universitat Pompeu Fabra (UPF), Barcelona, Spain. 16 3 R&D Department, Quantitative Genomic Medicine Laboratories (qGenomics), Barcelona, 17 Spain. 18 4 Functional Genomics Group, Cold Spring Harbor Laboratory, 1 Bungtown Road, Cold Spring 19 Harbor, New York 11724, USA. 20 5 Wellcome Trust Sanger Institute, Hinxton, Cambridgeshire, UK CB10 1HH. 21 6 Present address: International Institute of Molecular and Cell Biology, Ks. Trojdena 4, 02-109 22 Warsaw, Poland 23 7 Present address: Illumina, Cambridge, UK. 24 8 Present address: Department of Clinical Research, University of Bern, Murtenstrasse 35, 3010 25 Bern, Switzerland. 1 bioRxiv preprint doi: https://doi.org/10.1101/105064; this version posted September 4, 2017. -

3 Characterization of Intergenic Regions and Gene Definition
ENCODE 3 Characterization of intergenic regions and gene definition The prevalence and analysis of ENCODE data are changing the definition and characterization of intergenic and genic regions The cumulative coverage of transcribed regions in the 15 cell lines across the human genome is 62.1% and 74.7% for processed and primary transcripts, respectively (Supplementary Table 10 and Supplementary Fig. 22). On average, for each cell line, 39% of the genome is covered by primary transcripts and 22% by processed RNAs. No cell line showed transcription of more than 56.7% of the union of the expressed transcriptomes across all cell lines. When mapping the current RNA-seq data to the ENCODE pilot regions (Supplementary Table 10), we observed a similar, albeit higher, extent of transcriptional coverage of 73.3% for processed RNAs and 84.5% for primary transcripts. Previously reported estimates in these regions for processed and primary transcripts were 24% and 93%, respectively (Supplementary Table 2.4.3 and ref. 3). The increased genome coverage by processed RNAs stems largely from the inclusion of non-polyadenylated RNAs in the current study. Other than that, given the differences in the samples studied, the selection of pilot regions with high genic content, the increase of annotated genomic regions over time, and the different technologies used to interrogate transcription, both estimates are in reasonable agreement. As a consequence of both the expansion of genic regions by the discovery of new isoforms and the identification of novel intergenic transcripts, there has been a marked increase in the number of intergenic regions (from 32,481 to 60,250) due to their fragmentation and a decrease in their lengths (from 14,170 bp to 3,949 bp median length; Fig. -

Universal Alternative Splicing of Noncoding Exons
ManuscriptbioRxiv REV preprint doi: https://doi.org/10.1101/136275; this version posted December 18, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Universal alternative splicing of noncoding exons Ira W. Deveson1,2*, Marion E. Brunck3,4*, James Blackburn1,5, Elizabeth Tseng6, Ting Hon6, Tyson A. Clark6, Michael B. Clark1,7, Joanna Crawford8, Marcel E. Dinger1,5, Lars K. Nielsen4,9, John S. Mattick1,2,5è & Tim R. Mercer1,5,10è⌘ 5 1 Garvan Institute of Medical Research, NSW, Australia 2 School of Biotechnology and Biomolecular Sciences, Faculty of Science, University of New South Wales, Sydney, Australia 3 Centro de Biotecnologia FEMSA, Tecnologico de Monterrey, Campus Monterrey, Ave. Eugenio Garza Sada, Monterrey, NL, Mexico. 10 4 Australian Institute for Bioengineering and Nanotechnology, University of Queensland, QLD, Australia 5 St Vincent’s Clinical School, University of New South Wales, Sydney, Australia 6 Pacific Biosciences, Menlo Park, CA, USA 7 Department of Psychiatry, Warneford Hospital, University of Oxford, Oxford, UK 8 Institute for Molecular Bioscience, University of Queensland, QLD, Australia 15 9 Novo Nordisk Foundation Center for Biosustainability, Technical University of Denmark, Denmark 10 Altius Institute for Biomedical Sciences, Seattle, WA, USA * These authors contributed equally è Joint corresponding authors: [email protected], [email protected] ⌘ Lead Contact 20 The human transcriptome is so large, diverse and dynamic that, even after a decade of investigation by RNA sequencing (RNA-Seq), we are yet to resolve its true dimensions. -

Nearly All New Protein-Coding Predictions in the CHESS Database Are Not Protein-Coding Irwin Jungreis*,✝,1,2, Michael L
bioRxiv preprint doi: https://doi.org/10.1101/360602; this version posted July 2, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Nearly all new protein-coding predictions in the CHESS database are not protein-coding Irwin Jungreis*,✝,1,2, Michael L. Tress*,3, Jonathan Mudge*,4, Cristina Sisu5,6, Toby Hunt4, Rory Johnson7,8, Barbara Uszczynska-Ratajczak9, Julien Lagarde10,11,12, James Wright13, Paul Muir14,15, Mark Gerstein5,16,17, Roderic Guigo10,11,12, Manolis Kellis1,2, Adam Frankish✝,4, Paul Flicek4, The GENCODE Consortium 1MIT Computer Science and Artificial Intelligence Laboratory, Cambridge, MA; 2Broad Institute of MIT and Harvard, Cambridge, MA; 3Bioinformatics Unit, Spanish National Cancer Research Centre, Madrid, Spain; 4European Molecular Biology Laboratory, European Bioinformatics Institute, Wellcome Genome Campus, Hinxton, Cambridge CB10 1SD, UK; 5Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT 06520, USA; 6Department of Bioscience, Brunel University London, Uxbridge, UB8 3PH, UK; 7Department of Medical Oncology, Inselspital, University Hospital, University of Bern, Bern, Switzerland; 8Department of Biomedical Research (DBMR), University of Bern, Bern, Switzerland; 9Centre of New Technologies, University of Warsaw, Warsaw, Poland; 10Centre for Genomic Regulation (CRG), The Barcelona Institute for -

Accurate Mutation Annotation and Functional Prediction Enhance the Applicability of -Omics Data in Precision Medicine
The Texas Medical Center Library DigitalCommons@TMC The University of Texas MD Anderson Cancer Center UTHealth Graduate School of The University of Texas MD Anderson Cancer Biomedical Sciences Dissertations and Theses Center UTHealth Graduate School of (Open Access) Biomedical Sciences 5-2016 Accurate mutation annotation and functional prediction enhance the applicability of -omics data in precision medicine Tenghui Chen Follow this and additional works at: https://digitalcommons.library.tmc.edu/utgsbs_dissertations Part of the Bioinformatics Commons, Computational Biology Commons, Genomics Commons, and the Medicine and Health Sciences Commons Recommended Citation Chen, Tenghui, "Accurate mutation annotation and functional prediction enhance the applicability of -omics data in precision medicine" (2016). The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access). 666. https://digitalcommons.library.tmc.edu/utgsbs_dissertations/666 This Dissertation (PhD) is brought to you for free and open access by the The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences at DigitalCommons@TMC. It has been accepted for inclusion in The University of Texas MD Anderson Cancer Center UTHealth Graduate School of Biomedical Sciences Dissertations and Theses (Open Access) by an authorized administrator of DigitalCommons@TMC. For more information, please contact [email protected]. ACCURATE MUTATION ANNOTATION AND FUNCTIONAL PREDICTION -

RNA-Seq Analysis Reveals Localization-Associated Alternative Splicing Across 13 Cell Lines
G C A T T A C G G C A T genes Article RNA-Seq Analysis Reveals Localization-Associated Alternative Splicing across 13 Cell Lines Chao Zeng 1,2,* and Michiaki Hamada 1,2,3,4,* 1 AIST-Waseda University Computational Bio Big-Data Open Innovation Laboratory (CBBD-OIL), Tokyo 169-8555, Japan 2 Faculty of Science and Engineering, Waseda University, Tokyo 169-8555, Japan 3 Institute for Medical-oriented Structural Biology, Waseda University, Tokyo 162-8480, Japan 4 Graduate School of Medicine, Nippon Medical School, Tokyo 113-8602, Japan * Correspondence: [email protected] (C.Z.); [email protected] (M.H.) Received: 19 June 2020; Accepted: 17 July 2020; Published: 18 July 2020 Abstract: Alternative splicing, a ubiquitous phenomenon in eukaryotes, is a regulatory mechanism for the biological diversity of individual genes. Most studies have focused on the effects of alternative splicing for protein synthesis. However, the transcriptome-wide influence of alternative splicing on RNA subcellular localization has rarely been studied. By analyzing RNA-seq data obtained from subcellular fractions across 13 human cell lines, we identified 8720 switching genes between the cytoplasm and the nucleus. Consistent with previous reports, intron retention was observed to be enriched in the nuclear transcript variants. Interestingly, we found that short and structurally stable introns were positively correlated with nuclear localization. Motif analysis reveals that fourteen RNA-binding protein (RBPs) are prone to be preferentially bound with such introns. To our knowledge, this is the first transcriptome-wide study to analyze and evaluate the effect of alternative splicing on RNA subcellular localization.