A Reusable Javascript Component Biojs
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
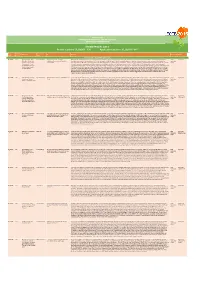
DATA Poster Numbers: P Da001 - 130 Application Posters: P Da001 - 041
POSTER LIST ORDERED ALPHABETICALLY BY POSTER TITLE GROUPED BY THEME/TRACK THEME/TRACK: DATA Poster numbers: P_Da001 - 130 Application posters: P_Da001 - 041 Poster EasyChair Presenting Author list Title Abstract Theme/track Topics number number author APPLICATION POSTERS WITHIN DATA THEME P_Da001 773 Benoît Carrères, Anne Benoît Carrères A systems approach to explore Microalgae are promising platforms for sustainable biofuel production. They produce triacyl-glycerides (TAG) which are easily converted into biofuel. When exposed to nitrogen limitation, Data/ Application Klok, Maria Suarez Diez, triacylglycerol production in Neochloris Neochloris oleoabundans accumulates up to 40% of its dry weight in TAG. However, a feasible production requires a decrease of production costs, which can be partially reached by Application Lenny de Jaeger, Mark oleoabundans increasing TAG yield.We built a constraint-based model describing primary metabolism of N. oleoabundans. It was grown in combinations of light absorption and nitrate supply rates and the poster Sturme, Packo Lamers, parameters needed for modeling of metabolism were measured. Fluxes were then calculated by flux balance analysis. cDNA samples of 16 experimental conditions were sequenced, Rene' Wijffels, Vitor Dos assembled and functionally annotated. Relative expression changes and relative flux changes for all reactions in the model were compared.The model predicts a maximum TAG yield on light Santos, Peter Schaap of 1.07g (mol photons)-1, more than 3 times current yield under optimal conditions. Furthermore, from optimization scenarios we concluded that increasing light efficiency has much higher and Dirk Martens potential to increase TAG yield than blocking entire pathways.Certain reaction expression patterns suggested an interdependence of the response to nitrogen and light supply. -

A Molecular Phylogenetic Toolkit Using Node.Js 1 2 3 Damien M
bioRxiv preprint doi: https://doi.org/10.1101/075101; this version posted October 9, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Phylo-Node: a molecular phylogenetic toolkit using Node.js 2 3 4 Damien M. O’Halloran1,2* 5 6 1. Department of Biological Sciences, The George Washington University, Science 7 and Engineering Hall, Rm 6000, 800 22nd Street N.W., Washington DC 20052, USA. 8 2. Institute for Neuroscience, The George Washington University, 636 Ross Hall, 9 2300 I St. N.W. Washington DC 20052, USA. 10 11 12 13 *Corresponding author address: 14 Damien O’Halloran, The George Washington University, 636 Ross Hall, 2300 I St. 15 N.W. Washington DC 20052, USA. 16 Tel: 202-994-8955 17 Fax: 202-994-6100 18 EMAIL: [email protected] 19 20 21 22 23 24 25 1 bioRxiv preprint doi: https://doi.org/10.1101/075101; this version posted October 9, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 26 ABSTRACT 27 Background: Node.js is an open-source and cross-platform environment that 28 provides a JavaScript codebase for back-end server-side applications. JavaScript 29 has been used to develop very fast, and user-friendly front-end tools for bioinformatic 30 and phylogenetic analyses. However, no such toolkits are available using Node.js to 31 conduct comprehensive molecular phylogenetic analysis. -

Biojs-HGV Viewer: Genetic Variation Visualizer
bioRxiv preprint doi: https://doi.org/10.1101/032573; this version posted November 23, 2015. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. BioJS-HGV Viewer: Genetic Variation Visualizer Saket Choudhary 1, Leyla Garcia 2, Andrew Nightingale 2 and Maria Martin 2 1Molecular and Computational Biology, University of Southern California, Los Angeles, California, 90089, USA 2European Molecular Biology laboratory - European Bioinformatics Institute (EMBL-EBI), Hinxton, Cambridge, CB10 1SD, UK ABSTRACT BioJS-HGV Viewer is a BioJS (Gomez´ et al. (2013)) component Motivation: Studying the pattern of genetic variants is a primary developed to visualize genetic variants in a comprehensive manner. step in deciphering the basis of biological diversity, identifying key Using it requires very limited knowledge of HTML and javascript. ‘driver variants’ that affect disease states and evolution of a species. BioJS is an open source javascript library providing various Catalogs of genetic variants contain vast numbers of variants and components to visualize biological data. The visualizations are web are growing at an exponential rate, but lack an interactive exploratory based and hence are almost platform independent. interface. Results: We present BioJS-HGV Viewer, a BioJS component to represent and visualize genetic variants pooled from different sources. The tool displays sequences and variants at different levels 2 METHODS of detail, facilitating representation of variant sites and annotations in The functionality provided by BioJS-HGV Viewer has two mode a user friendly and interactive manner. -

a Biojs Component to Visualize KEGG Pathways Keggviewer [V1
F1000Research 2014, 3:43 Last updated: 05 MAR 2015 WEB TOOL KEGGViewer, a BioJS component to visualize KEGG Pathways [v1; ref status: indexed, http://f1000r.es/2uq] Jose M. Villaveces1, Rafael C. Jimenez2, Bianca H. Habermann1 1Max Planck Institute of Biochemistry, Am Klopferspitz 18, 82152, Germany 2European Bioinformatics Institute, Wellcome Trust Genome Campus, Hinxton, Cambridge, CB10 1SD, UK v1 First published: 13 Feb 2014, 3:43 (doi: 10.12688/f1000research.3-43.v1) Open Peer Review Latest published: 13 Feb 2014, 3:43 (doi: 10.12688/f1000research.3-43.v1) Referee Status: Abstract Summary: Signaling pathways provide essential information on complex regulatory processes within the cell. They are moreover widely used to interpret Invited Referees and integrate data from large-scale studies, such as expression or functional 1 2 screens. We present KEGGViewer a BioJS component to visualize KEGG pathways and to allow their visual integration with functional data. version 1 Availability: KEGGViewer is an open-source tool freely available at the BioJS published report report Registry. Instructions on how to use the tool are available at 13 Feb 2014 http://goo.gl/dVeWpg and the source code can be found at http://github.com/biojs/biojs and DOI:10.5281/zenodo.7708. 1 Hedi Peterson, University of Geneva Switzerland, Priit Adler, University of Tartu Estonia 2 Alexander Pico, Gladstone Institutes USA This article is included in the BioJS Discuss this article Comments (0) Corresponding author: Bianca H. Habermann ([email protected]) How to cite this article: Villaveces JM, Jimenez RC and Habermann BH. KEGGViewer, a BioJS component to visualize KEGG Pathways [v1; ref status: indexed, http://f1000r.es/2uq] F1000Research 2014, 3:43 (doi: 10.12688/f1000research.3-43.v1) Copyright: © 2014 Villaveces JM et al. -

1 Phylo-Node: a Molecular Phylogenetic Toolkit Using Node.Js
bioRxiv preprint doi: https://doi.org/10.1101/075101; this version posted September 19, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Phylo-Node: a molecular phylogenetic toolkit using Node.js 2 3 4 Damien M. O’Halloran1,2* 5 6 1. Department of Biological Sciences, The George Washington University, Science and 7 Engineering Hall, Rm 6000, 800 22nd Street N.W., Washington DC 20052, USA. 8 2. Institute for Neuroscience, The George Washington University, 636 Ross Hall, 2300 I 9 St. N.W. Washington DC 20052, USA. 10 11 12 13 *Corresponding author address: 14 Damien O’Halloran, The George Washington University, 636 Ross Hall, 2300 I St. N.W. 15 Washington DC 20052, USA. 16 Tel: 202-994-8955 17 Fax: 202-994-6100 18 EMAIL: [email protected] 19 20 21 22 23 1 bioRxiv preprint doi: https://doi.org/10.1101/075101; this version posted September 19, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 24 ABSTRACT 25 Background: Node.js is an open-source and cross-platform environment that provides 26 a JavaScript codebase for back-end server-side applications. JavaScript has been used 27 to develop very fast, and user-friendly front-end tools for bioinformatic and phylogenetic 28 analyses. However, no such toolkits are available using Node.js to conduct 29 comprehensive molecular phylogenetic analysis. 30 Results: To address this problem, I have developed, Phylo-Node, which was developed 31 using Node.js and provides a fast, stable, and scalable toolkit that allows the user to go 32 from sequence retrieval to phylogeny reconstruction. -

Thesis Entire.Pdf (12.90Mb)
Faculty OF Science AND TECHNOLOGY Department OF Computer Science TOWARD ReprODUCIBLE Analysis AND ExplorATION OF High-ThrOUGHPUT Biological Datasets — Bjørn Fjukstad A DISSERTATION FOR THE DEGREE OF Philosophiae Doctor – 2018 This thesis document was typeset using the UiT Thesis LaTEX Template. © 2018 – http://github.com/egraff/uit-thesis “Ta aldri problemene på forskudd, for da får du dem to ganger, men ta gjerne seieren på forskudd, for hvis ikke er det altfor sjelden du får oppleve den.” –Ivar Tollefsen AbstrACT There is a rapid growth in the number of available biological datasets due to the advent of high-throughput data collection instruments combined with cheap compute infrastructure. Modern instruments enable the analysis of biological data at different levels, from small DNA sequences through larger cell structures, and up to the function of entire organs. These new datasets have brought the need to develop new software packages to enable novel insights into the underlying biological mechanisms in the development and progression of diseases such as cancer. The heterogeneity of biological datasets require researchers to tailor the explo- ration and analyses with a wide range of different tools and systems. However, despite the need for their integration, few of them provide standard inter- faces for analyses implemented using different programming languages and frameworks. In addition, because of the many tools, different input parame- ters, and references to databases, it is necessary to record these correctly. The lack of such details complicates reproducing the original results and the reuse of the analyses on new datasets. This increases the analysis time and leaves unrealized potential for scientific insights. -

Blastjs: a BLAST+ Wrapper for Node.Js
Page et al. BMC Res Notes (2016) 9:130 DOI 10.1186/s13104-016-1938-1 BMC Research Notes TECHNICAL NOTE Open Access blastjs: a BLAST wrapper for Node.js Martin Page1, Dan MacLean1 and Christian+ Schudoma1,2* Abstract Background: To cope with the ever-increasing amount of sequence data generated in the field of genomics, the demand for efficient and fast database searches that drive functional and structural annotation in both large- and small-scale genome projects is on the rise. The tools of the BLAST suite are the most widely employed bioinformatic method for these database searches. Recent trends in bioinformatics+ application development show an increasing number of JavaScript apps that are based on modern frameworks such as Node.js. Until now, there is no way of using database searches with the BLAST suite from a Node.js codebase. + Results: We developed blastjs, a Node.js library that wraps the search tools of the BLAST suite and thus allows to easily add significant functionality to any Node.js-based application. + Conclusion: blastjs is a library that allows the incorporation of BLAST functionality into bioinformatics applications based on JavaScript and Node.js. The library was designed to be as user-friendly+ as possible and therefore requires only a minimal amount of code in the client application. The library is freely available under the MIT license at https:// github.com/teammaclean/blastjs. Keywords: Sequence analysis, Database search, Annotation, Web services, Application development, JavaScript Findings assessment of BLAST results via output processing and/ Background or visualisation. The algorithms of the BLAST-family [1–3] are the most Rich web-application development is greatly facilitated widely employed methods for efficiently searching bio- by taking advantage of the power of modern web brows- logical databases. -

Trends in IT Innovation to Build a Next Generation Bioinformatics Solution to Manage and Analyse Biological Big Data Produced by NGS Technologies Alexandre G
Trends in IT Innovation to Build a Next Generation Bioinformatics Solution to Manage and Analyse Biological Big Data Produced by NGS Technologies Alexandre G. de Brevern, Jean-Philippe Meyniel, Cecile Fairhead, Cécile Neuvéglise, Alain Malpertuy To cite this version: Alexandre G. de Brevern, Jean-Philippe Meyniel, Cecile Fairhead, Cécile Neuvéglise, Alain Malpertuy. Trends in IT Innovation to Build a Next Generation Bioinformatics Solution to Manage and Anal- yse Biological Big Data Produced by NGS Technologies. BioMed Research International , Hindawi Publishing Corporation, 2015, pp.1-15. 10.1155/2015/904541. hal-02635313 HAL Id: hal-02635313 https://hal.inrae.fr/hal-02635313 Submitted on 27 May 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution| 4.0 International License Hindawi Publishing Corporation BioMed Research International Volume 2015, Article ID 904541, 15 pages http://dx.doi.org/10.1155/2015/904541 Review Article Trends in IT Innovation to Build a Next Generation Bioinformatics Solution -

Anatomy of Biojs, an Open Source Community for the Life Sciences
FEATURE ARTICLE elifesciences.org CUTTING EDGE Anatomy of BioJS, an open source community for the life sciences Abstract BioJS is an open source software project that develops visualization tools for different types of biological data. Here we report on the factors that influenced the growth of the BioJS user and developer community, and outline our strategy for building on this growth. The lessons we have learned on BioJS may also be relevant to other open source software projects. DOI: 10.7554/eLife.07009.001 GUY YACHDAV*, TATYANA GOLDBERG, SEBASTIAN WILZBACH, DAVID DAO, IRIS SHIH, SAKET CHOUDHARY, STEVE CROUCH, MAX FRANZ, ALEXANDER GARCIA,´ LEYLA J GARCIA,´ BJORN¨ A GRUNING,¨ DEVASENA INUPAKUTIKA, IAN SILLITOE, ANIL S THANKI, BRUNO VIEIRA, JOSE´ M VILLAVECES, MARIA V SCHNEIDER, SUZANNA LEWIS, STEVE PETTIFER, BURKHARD ROST AND MANUEL CORPAS* Introduction support needed to survive beyond the originator’s BioJavaScript (BioJS; http://biojs.net/) was set initial enthusiasm and/or funding. up to meet a need for an open source library of In the case of BioJS we were acutely aware of reusable components to visualize and analyse the need to gain buy-in from the community for biological data on the web (Figure 1; Corpas two reasons: et al., 2014). These components are discrete 1. To fulfil the vision of a suite of tools capable of modules that can be reused, extended and displaying diverse biological data requires combined to meet a particular visualization expertise and capacity that is well beyond that need. Unlike proprietary (or closed-source) of any individual group of developers working systems, which are typically distributed as in isolation. -

1 Phylo-Node: a Molecular Phylogenetic Toolkit Using Node.Js
bioRxiv preprint doi: https://doi.org/10.1101/075101; this version posted January 13, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 phylo-node: A Molecular Phylogenetic Toolkit using Node.js 2 3 4 Damien M. O’Halloran1,2* 5 6 1. Department of Biological Sciences, The George Washington University, Science 7 and Engineering Hall, Rm 6000, 800 22nd Street N.W., Washington DC 20052, USA. 8 2. Institute for Neuroscience, The George Washington University, 636 Ross Hall, 9 2300 I St. N.W. Washington DC 20052, USA. 10 11 12 13 *Corresponding author address: 14 Damien O’Halloran, The George Washington University, 636 Ross Hall, 2300 I St. 15 N.W. Washington DC 20052, USA. 16 Tel: 202-994-8955 17 Fax: 202-994-6100 18 Email: [email protected] 19 20 21 22 23 24 25 1 bioRxiv preprint doi: https://doi.org/10.1101/075101; this version posted January 13, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 26 ABSTRACT 27 28 Background: Node.js is an open-source and cross-platform environment that 29 provides a JavaScript codebase for back-end server-side applications. JavaScript 30 has been used to develop very fast and user-friendly front-end tools for bioinformatic 31 and phylogenetic analyses. However, no such toolkits are available using Node.js to 32 conduct comprehensive molecular phylogenetic analysis. -

Visually Guiding Users in Selection, Exploration, and Presentation Tasks
Submitted by DI Samuel Gratzl, Bsc. Submitted at Institute of Computer Graphics Supervisor and First Examiner Assoz.Univ.-Prof. Dipl.-Ing. Dr.techn. Marc Streit Visually Guiding Users in Second Examiner Prof. Dr. Tobias Schreck Selection, Exploration, March 2017 and Presentation Tasks Doctoral Thesis to obtain the academic degree of Doktor der technischen Wissenschaften in the Doctoral Program Technische Wissenschaften JOHANNES KEPLER UNIVERSITY LINZ Altenbergerstraße 69 4040 Linz, Osterreich¨ www.jku.at DVR 0093696 Abstract Making scientific discoveries based on large and heterogeneous datasets is challenging. The continuous improvement of data acquisition technologies makes it possible to collect more and more data. However, not only the amount of data is growing at a fast pace, but also its complexity. Visually analyzing such large, interconnected data collections requires a user to perform a combination of selection, exploration, and presentation tasks. In each of these tasks a user needs guidance in terms of (1) what data subsets are to be investigated from the data collection, (2) how to effectively and efficiently explore selected data subsets, and (3) how to effectively reproduce findings and tell the story of their discovery. On the basis of a unified model called the SPARE model, this thesis makes contributions to all three guidance tasks a user encounters during a visual analysis session: The LineUp multi-attribute ranking technique was developed to order and prioritize item collections. It is an essential building block of the proposed guidance process that has the goal of better supporting users in data selection tasks by scoring and ranking data subsets based on user-defined queries. -

Downloads Or Setups, Because They Are Built Into the Webpage Structure
informatics Review Web Apps Come of Age for Molecular Sciences Luciano A. Abriata ID Laboratory for Biomolecular Modeling, École Polytechnique Fédérale de Lausanne and Swiss Institute of Bioinformatics, CH-1015 Lausanne, Switzerland; luciano.abriata@epfl.ch; Tel.: +41-021-69-30963 Academic Editor: Igor I. Baskin Received: 11 August 2017; Accepted: 20 August 2017; Published: 24 August 2017 Abstract: Whereas server-side programs are essential to maintain databases and run data analysis pipelines and simulations, client-side web-based computing tools are also important as they allow users to access, visualize and analyze the content delivered to their devices on-the-fly and interactively. This article reviews the best-established tools for in-browser plugin-less programming, including JavaScript as used in HTML5 as well as related web technologies. Through examples based on JavaScript libraries, web applets, and even full web apps, either alone or coupled to each other, the article puts on the spotlight the potential of these technologies for carrying out numerical calculations, text processing and mining, retrieval and analysis of data through queries to online databases and web services, effective visualization of data including 3D visualization and even virtual and augmented reality; all of them in the browser at relatively low programming effort, with applications in cheminformatics, structural biology, biophysics, and genomics, among other molecular sciences. Keywords: JavaScript; HTML; programming; data analysis; internet 1. Introduction Slightly over 20 years ago, a small commentary by Prof. L. Stein in Trends in Genetics anticipated the impact that “web applets” were to have on how scientists browse, visualize, and analyze biological data available online [1].