Unipro UGENE Online User Manual
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

T-Coffee Documentation Release Version 13.45.47.Aba98c5
T-Coffee Documentation Release Version_13.45.47.aba98c5 Cedric Notredame Aug 31, 2021 Contents 1 T-Coffee Installation 3 1.1 Installation................................................3 1.1.1 Unix/Linux Binaries......................................4 1.1.2 MacOS Binaries - Updated...................................4 1.1.3 Installation From Source/Binaries downloader (Mac OSX/Linux)...............4 1.2 Template based modes: PSI/TM-Coffee and Expresso.........................5 1.2.1 Why do I need BLAST with T-Coffee?.............................6 1.2.2 Using a BLAST local version on Unix.............................6 1.2.3 Using the EBI BLAST client..................................6 1.2.4 Using the NCBI BLAST client.................................7 1.2.5 Using another client.......................................7 1.3 Troubleshooting.............................................7 1.3.1 Third party packages......................................7 1.3.2 M-Coffee parameters......................................9 1.3.3 Structural modes (using PDB)................................. 10 1.3.4 R-Coffee associated packages................................. 10 2 Quick Start Regressive Algorithm 11 2.1 Introduction............................................... 11 2.2 Installation from source......................................... 12 2.3 Examples................................................. 12 2.3.1 Fast and accurate........................................ 12 2.3.2 Slower and more accurate.................................... 12 2.3.3 Very Fast........................................... -

Comparative Analysis of Multiple Sequence Alignment Tools
I.J. Information Technology and Computer Science, 2018, 8, 24-30 Published Online August 2018 in MECS (http://www.mecs-press.org/) DOI: 10.5815/ijitcs.2018.08.04 Comparative Analysis of Multiple Sequence Alignment Tools Eman M. Mohamed Faculty of Computers and Information, Menoufia University, Egypt E-mail: [email protected]. Hamdy M. Mousa, Arabi E. keshk Faculty of Computers and Information, Menoufia University, Egypt E-mail: [email protected], [email protected]. Received: 24 April 2018; Accepted: 07 July 2018; Published: 08 August 2018 Abstract—The perfect alignment between three or more global alignment algorithm built-in dynamic sequences of Protein, RNA or DNA is a very difficult programming technique [1]. This algorithm maximizes task in bioinformatics. There are many techniques for the number of amino acid matches and minimizes the alignment multiple sequences. Many techniques number of required gaps to finds globally optimal maximize speed and do not concern with the accuracy of alignment. Local alignments are more useful for aligning the resulting alignment. Likewise, many techniques sub-regions of the sequences, whereas local alignment maximize accuracy and do not concern with the speed. maximizes sub-regions similarity alignment. One of the Reducing memory and execution time requirements and most known of Local alignment is Smith-Waterman increasing the accuracy of multiple sequence alignment algorithm [2]. on large-scale datasets are the vital goal of any technique. The paper introduces the comparative analysis of the Table 1. Pairwise vs. multiple sequence alignment most well-known programs (CLUSTAL-OMEGA, PSA MSA MAFFT, BROBCONS, KALIGN, RETALIGN, and Compare two biological Compare more than two MUSCLE). -

Ple Sequence Alignment Methods: Evidence from Data
Mul$ple sequence alignment methods: evidence from data Tandy Warnow Alignment Error/Accuracy • SPFN: percentage of homologies in the true alignment that are not recovered (false negave homologies) • SPFP: percentage of homologies in the es$mated alignment that are false (false posi$ve homologies) • TC: total number of columns correctly recovered • SP-score: percentage of homologies in the true alignment that are recovered • Pairs score: 1-(avg of SP-FN and SP-FP) Benchmarks • Simulaons: can control everything, and true alignment is not disputed – Different simulators • Biological: can’t control anything, and reference alignment might not be true alignment – BAliBASE, HomFam, Prefab – CRW (Comparave Ribosomal Website) Alignment Methods (Sample) • Clustal-Omega • MAFFT • Muscle • Opal • Prank/Pagan • Probcons Co-es$maon of trees and alignments • Bali-Phy and Alifritz (stas$cal co-es$maon) • SATe-1, SATe-2, and PASTA (divide-and-conquer co- es$maon) • POY and Beetle (treelength op$mizaon) Other Criteria • Tree topology error • Tree branch length error • Gap length distribu$on • Inser$on/dele$on rao • Alignment length • Number of indels How does the guide tree impact accuracy? • Does improving the accuracy of the guide tree help? • Do all alignment methods respond iden$cally? (Is the same guide tree good for all methods?) • Do the default sengs for the guide tree work well? Alignment criteria • Does the relave performance of methods depend on the alignment criterion? • Which alignment criteria are predic$ve of tree accuracy? • How should we design MSA methods to produce best accuracy? Choice of best MSA method • Does it depend on type of data (DNA or amino acids?) • Does it depend on rate of evolu$on? • Does it depend on gap length distribu$on? • Does it depend on existence of fragments? Katoh and Standley . -
An Open-Sourced Bioinformatic Pipeline for the Processing of Next-Generation Sequencing Derived Nucleotide Reads
bioRxiv preprint doi: https://doi.org/10.1101/2020.04.20.050369; this version posted May 28, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. An open-sourced bioinformatic pipeline for the processing of Next-Generation Sequencing derived nucleotide reads: Identification and authentication of ancient metagenomic DNA Thomas C. Collin1, *, Konstantina Drosou2, 3, Jeremiah Daniel O’Riordan4, Tengiz Meshveliani5, Ron Pinhasi6, and Robin N. M. Feeney1 1School of Medicine, University College Dublin, Ireland 2Division of Cell Matrix Biology Regenerative Medicine, University of Manchester, United Kingdom 3Manchester Institute of Biotechnology, School of Earth and Environmental Sciences, University of Manchester, United Kingdom [email protected] 5Institute of Paleobiology and Paleoanthropology, National Museum of Georgia, Tbilisi, Georgia 6Department of Evolutionary Anthropology, University of Vienna, Austria *Corresponding Author Abstract The emerging field of ancient metagenomics adds to these Bioinformatic pipelines optimised for the processing and as- processing complexities with the need for additional steps sessment of metagenomic ancient DNA (aDNA) are needed in the separation and authentication of ancient sequences from modern sequences. Currently, there are few pipelines for studies that do not make use of high yielding DNA cap- available for the analysis of ancient metagenomic DNA ture techniques. These bioinformatic pipelines are tradition- 1 4 ally optimised for broad aDNA purposes, are contingent on (aDNA) ≠ The limited number of bioinformatic pipelines selection biases and are associated with high costs. -

Determination of Optimal Parameters of MAFFT Program Based on Balibase3.0 Database
Long et al. SpringerPlus (2016) 5:736 DOI 10.1186/s40064-016-2526-5 RESEARCH Open Access Determination of optimal parameters of MAFFT program based on BAliBASE3.0 database HaiXia Long1, ManZhi Li2* and HaiYan Fu1 *Correspondence: myresearch_hainnu@163. Abstract com Background: Multiple sequence alignment (MSA) is one of the most important 2 School of Mathematics and Statistics, Hainan Normal research contents in bioinformatics. A number of MSA programs have emerged. The University, Haikou 571158, accuracy of MSA programs highly depends on the parameters setting, mainly including HaiNan, China gap open penalties (GOP), gap extension penalties (GEP) and substitution matrix (SM). Full list of author information is available at the end of the This research tries to obtain the optimal GOP, GEP and SM rather than MAFFT default article parameters. Results: The paper discusses the MAFFT program benchmarked on BAliBASE3.0 data- base, and the optimal parameters of MAFFT program are obtained, which are better than the default parameters of CLUSTALW and MAFFT program. Conclusions: The optimal parameters can improve the results of multiple sequence alignment, which is feasible and efficient. Keywords: Multiple sequence alignment, Gap open penalties, Gap extension penalties, Substitution matrix, MAFFT program, Default parameters Background Multiple sequence alignment (MSA), one of the most basic bioinformatics tool, has wide applications in sequence analysis, gene recognition, protein structure prediction, and phylogenetic tree reconstruction, etc. MSA computation is a NP-complete problem (Lathrop 1995), whose time and space complexity have sharp increase while the length and the number of sequences are increasing. At present, many scholars have developed open source online alignment tools, such as CLUSTALW, T-COFFEE, MAFFT, (Thompson et al. -

The Art of Multiple Sequence Alignment in R
The Art of Multiple Sequence Alignment in R Erik S. Wright May 19, 2021 Contents 1 Introduction 1 2 Alignment Speed 2 3 Alignment Accuracy 4 4 Recommendations for optimal performance 7 5 Single Gene Alignment 8 5.1 Example: Protein coding sequences . 8 5.2 Example: Non-coding RNA sequences . 9 5.3 Example: Aligning two aligned sequence sets . 9 6 Advanced Options & Features 10 6.1 Example: Building a Guide Tree . 10 6.2 Example: Post-processing an existing multiple alignment . 12 7 Aligning Homologous Regions of Multiple Genomes 12 8 Session Information 15 1 Introduction This document is intended to illustrate the art of multiple sequence alignment in R using DECIPHER. Even though its beauty is often con- cealed, multiple sequence alignment is a form of art in more ways than one. Take a look at Figure 1 for an illustration of what is happening behind the scenes during multiple sequence alignment. The practice of sequence alignment is one that requires a degree of skill, and it is that art which this vignette intends to convey. It is simply not enough to \plug" sequences into a multiple sequence aligner and blindly trust the result. An appreciation for the art as well a careful consideration of the results are required. What really is multiple sequence alignment, and is there a single cor- rect alignment? Generally speaking, alignment seeks to perform the act of taking multiple divergent biological sequences of the same \type" and Figure 1: The art of multiple se- fitting them to a form that reflects some shared quality. -

HMMER User's Guide
HMMER User’s Guide Biological sequence analysis using profile hidden Markov models http://hmmer.org/ Version 3.0rc1; February 2010 Sean R. Eddy for the HMMER Development Team Janelia Farm Research Campus 19700 Helix Drive Ashburn VA 20147 USA http://eddylab.org/ Copyright (C) 2010 Howard Hughes Medical Institute. Permission is granted to make and distribute verbatim copies of this manual provided the copyright notice and this permission notice are retained on all copies. HMMER is licensed and freely distributed under the GNU General Public License version 3 (GPLv3). For a copy of the License, see http://www.gnu.org/licenses/. HMMER is a trademark of the Howard Hughes Medical Institute. 1 Contents 1 Introduction 5 How to avoid reading this manual . 5 How to avoid using this software (links to similar software) . 5 What profile HMMs are . 5 Applications of profile HMMs . 6 Design goals of HMMER3 . 7 What’s still missing in HMMER3 . 8 How to learn more about profile HMMs . 9 2 Installation 10 Quick installation instructions . 10 System requirements . 10 Multithreaded parallelization for multicores is the default . 11 MPI parallelization for clusters is optional . 11 Using build directories . 12 Makefile targets . 12 3 Tutorial 13 The programs in HMMER . 13 Files used in the tutorial . 13 Searching a sequence database with a single profile HMM . 14 Step 1: build a profile HMM with hmmbuild . 14 Step 2: search the sequence database with hmmsearch . 16 Searching a profile HMM database with a query sequence . 22 Step 1: create an HMM database flatfile . 22 Step 2: compress and index the flatfile with hmmpress . -

Improvement in the Accuracy of Multiple Sequence Alignment Program MAFFT
22 Genome Informatics 16(1): 22-33 (2005) Improvement in the Accuracy of Multiple Sequence Alignment Program MAFFT Kazutaka Katohl Kei-ichi Kuma1 [email protected] [email protected] Takashi Miyata2" Hiroyuki Toh5 [email protected] [email protected] 1 Bioinformatics Center , Institute for Chemical Research, Kyoto University, Uji, Kyoto 611-0011, Japan 2 Biohistory Research Hall , Takatsuki, Osaka 569-1125, Japan 3 Department of Electrical Engineering and Bioscience , Science and Engineering, Waseda University, Tokyo 169-8555, Japan 4 Department of Biophysics , Graduate School of Science, Kyoto University, Kyoto 606- 8502, Japan 5 Division of Bioinformatics , Research Center for Prevention of Infectious Diseases, Medical Institute of Bioregulation, Kyushu University, Fukuoka 812-8582,Japan Abstract In 2002, we developed and released a rapid multiple sequence alignment program MAFFT that was designed to handle a huge (up to•`5,000 sequences) and long data (,-2,000 as or •`5,000 nt) in a reasonable time on a standard desktop PC. As for the accuracy, however, the previous versions (v.4 and lower) of MAFFT were outperformed by ProbCons and TCoffee v.2, both of which were released in 2004, in several benchmark tests. Here we report a recent extension of MAFFT that aims to improve the accuracy with as little cost of calculation time as possible. The extended version of MAFFT (v.5) has new iterative refinement options, G-INS-i and L-INS-i (collectively denoted as [GL]-INS-iin this report). These options use a new objective function combining the weighted sum-of-pairs (WSP) score and a score similar to COFFEE derived from all pairwise alignments. -

Multiple Sequence Alignment
Bioinformatics Algorithms Multiple Sequence Alignment David Hoksza http://siret.ms.mff.cuni.cz/hoksza Outline • Motivation • Algorithms • Scoring functions • exhaustive • multidimensional dynamic programming • heuristics • progressive alignment • iterative alignment/refinement • block(local)-based alignment Multiple sequence alignment (MSA) • Goal of MSA is to find “optimal” mapping of a set of sequences • Homologous residues (originating in the same position in a common ancestor) among a set of sequences are aligned together in columns • Usually employs multiple pairwise alignment (PA) computations to reveal the evolutionarily equivalent positions across all sequences Motivation • Distant homologues • faint similarity can become apparent when present in many sequences • motifs might not be apparent from pairwise alignment only • Detection of key functional residues • amino acids critical for function tend to be conserved during the evolution and therefore can be revealed by inspecting sequences within given family • Prediction of secondary/tertiary structure • Inferring evolutionary history 4 Representation of MSA • Column-based representation • Profile representation (position specific scoring matrix) • Sequence logo Manual MSA • High quality MSA can be carried out automatic MSA algorithms by hand using expert knowledge • specific columns • BAliBASE • highly conserved residues • https://lbgi.fr/balibase/ • buried hydrophobic residues • PROSITE • secondary structure (especially in RNA • http://prosite.expasy.org/ alignment) • Pfam • expected -
Align with Clustalw Element
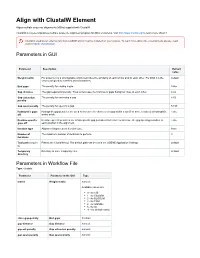
Align with ClustalW Element Aligns multiple sequence alignments (MSAs) supplied with ClustalW. ClustalW is a general purpose multiple sequence alignment program for DNA or proteins. Visit http://www.clustal.org/ to learn more about it. Clustal is used as an external tool from UGENE and it must be installed on your system. To learn more about the external tools, please, read main UGENE User Manual. Parameters in GUI Parameter Description Default value Weight matrix For proteins it is a scoring table which describes the similarity of each amino acid to each other. For DNA it is the default scores assigned to matches and mismatches. End gaps The penalty for closing a gap. False Gap distance The gap separation penalty. Tries to decrease the chances of gaps being too close to each other. 4.42 Gap extension The penalty for extending a gap. 8.52 penalty Gap open penalty The penalty for opening a gap. 53.90 Hydrophilic gaps Hydrophilic gap penalties are used to increase the chances of a gap within a run (5 or more residues) of hydrophilic False off amino acids. Residue-specific Residue-specific penalties are amino specific gap penalties that reduce or increase the gap opening penalties at False gaps off each position in the alignment. Iteration type Alignment improvement iteration type. None Number of The maximum number of iterations to perform. 3 iterations Tool path (require Path to the ClustalW tool. The default path can be set in the UGENE Application Settings. default d) Temporary Directory to store temporary files. default directory Parameters -

Human Genome Graph Construction from Multiple Long-Read Assemblies
F1000Research 2018, 7:1391 Last updated: 14 OCT 2019 SOFTWARE TOOL ARTICLE NovoGraph: Human genome graph construction from multiple long-read de novo assemblies [version 2; peer review: 2 approved] A genome graph representation of seven ethnically diverse whole human genomes Evan Biederstedt1,2, Jeffrey C. Oliver3, Nancy F. Hansen4, Aarti Jajoo5, Nathan Dunn 6, Andrew Olson7, Ben Busby8, Alexander T. Dilthey 4,9 1Weill Cornell Medicine, New York, NY, 10065, USA 2New York Genome Center, New York, NY, 10013, USA 3Office of Digital Innovation and Stewardship, University Libraries, University of Arizona, Tucson, AZ, 85721, USA 4National Human Genome Research Institute, National Institutes of Health, Bethesda, MD, 20817, USA 5Baylor College of Medicine, Houston, TX, 77030, USA 6Lawrence Berkeley National Laboratory, Berkeley, CA, 94720, USA 7Cold Spring Harbor Laboratory, Cold Spring Harbor, NY, 11724, USA 8National Center for Biotechnology Information, National Institutes of Health, Bethesda, MD, 20817, USA 9Institute of Medical Microbiology and Hospital Hygiene, Heinrich Heine University Düsseldorf, Düsseldorf, 40225, Germany First published: 03 Sep 2018, 7:1391 ( Open Peer Review v2 https://doi.org/10.12688/f1000research.15895.1) Latest published: 10 Dec 2018, 7:1391 ( https://doi.org/10.12688/f1000research.15895.2) Reviewer Status Abstract Invited Reviewers Genome graphs are emerging as an important novel approach to the 1 2 analysis of high-throughput human sequencing data. By explicitly representing genetic variants and alternative haplotypes in a mappable data structure, they can enable the improved analysis of structurally version 2 report variable and hyperpolymorphic regions of the genome. In most existing published approaches, graphs are constructed from variant call sets derived from 10 Dec 2018 short-read sequencing. -

Real Time Collaboration on Multiple Sequence Alignments Jeffrey Yunes Submitted in Partial Satisfaction of the Require
ShareMSA: real time collaboration on multiple sequence alignments Jeffrey Yunes Submitted in partial satisfaction of the requirements for the degree of m in the GRADUATE DIVISION UNIVERSITY OF CALIFORNIA, SAN FRANCISCO ii Acknowledgements I am very thankful for the guidance and support of my thesis advisor and chair, Patricia C. Babbitt. I thank my thesis committee members Hao Li and Ian Holmes. I am grateful to Elaine Meng for feedback on user experience and Eric Pettersen for help with Chimera plugins. This work was funded by NIH GM60595 to Patricia C. Babbitt. iii Table of Contents Chapter1.................................................. 1 Abstract................................................. 1 Introduction............................................... 1 Features................................................. 1 ExamplesofUse ............................................ 2 Conclusion ............................................... 4 References . 5 iv List of Figures Figure 1 Screenshot from the mobile version of the site . 3 v Abstract A Multiple Sequence Alignment (MSA) is a widely used data structure in bioinformatics for applications ranging from evolutionary analyses to protein engineering. ShareMSA is a website that allows users to visualize, share, and collaborate on MSAs. ShareMSA makes MSAs more accessible by allowing collaborators one-click access to the MSA and annotations without the need to install software. In addition, when those with requisite permissions change the annotations or legend, all viewers of the MSA see the changes in real time. A Chimera plugin is available via the ShareMSA website, allowing Chimera users to publish MSAs from within this sequence-structure visualization software. Availability and Implementation ShareMSA is freely available at https://www.sharemsa.com, and does not require registration. The front end is implemented in HTML and JavaScript and runs on all modern browsers that support WebSockets.