Sctenifoldnet: a Machine Learning Workflow for Constructing and Comparing Transcriptome-Wide Gene Regulatory Networks from Single-Cell Data
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Mineralocorticoid Receptor Leads to Increased Expression of EGFR
www.nature.com/scientificreports OPEN The mineralocorticoid receptor leads to increased expression of EGFR and T‑type calcium channels that support HL‑1 cell hypertrophy Katharina Stroedecke1,2, Sandra Meinel1,2, Fritz Markwardt1, Udo Kloeckner1, Nicole Straetz1, Katja Quarch1, Barbara Schreier1, Michael Kopf1, Michael Gekle1 & Claudia Grossmann1* The EGF receptor (EGFR) has been extensively studied in tumor biology and recently a role in cardiovascular pathophysiology was suggested. The mineralocorticoid receptor (MR) is an important efector of the renin–angiotensin–aldosterone‑system and elicits pathophysiological efects in the cardiovascular system; however, the underlying molecular mechanisms are unclear. Our aim was to investigate the importance of EGFR for MR‑mediated cardiovascular pathophysiology because MR is known to induce EGFR expression. We identifed a SNP within the EGFR promoter that modulates MR‑induced EGFR expression. In RNA‑sequencing and qPCR experiments in heart tissue of EGFR KO and WT mice, changes in EGFR abundance led to diferential expression of cardiac ion channels, especially of the T‑type calcium channel CACNA1H. Accordingly, CACNA1H expression was increased in WT mice after in vivo MR activation by aldosterone but not in respective EGFR KO mice. Aldosterone‑ and EGF‑responsiveness of CACNA1H expression was confrmed in HL‑1 cells by Western blot and by measuring peak current density of T‑type calcium channels. Aldosterone‑induced CACNA1H protein expression could be abrogated by the EGFR inhibitor AG1478. Furthermore, inhibition of T‑type calcium channels with mibefradil or ML218 reduced diameter, volume and BNP levels in HL‑1 cells. In conclusion the MR regulates EGFR and CACNA1H expression, which has an efect on HL‑1 cell diameter, and the extent of this regulation seems to depend on the SNP‑216 (G/T) genotype. -

Monogenic Causation in Chronic Kidney Disease
University of Dublin, Trinity College School of Medicine, Department of Medicine Investigation of the monogenic causes of chronic kidney disease PhD Thesis April 2020 Dervla Connaughton Supervisor: Professor Mark Little Co-Supervisors: Professor Friedhelm Hildebrandt and Professor Peter Conlon 1 DECLARATION I declare that this thesis has not been submitted as an exercise for a degree at this or any other university and it is entirely my own work. This work was funding by the Health Research Board, Ireland (HPF-206-674), the International Pediatric Research Foundation Early Investigators’ Exchange Program and the Amgen® Irish Nephrology Society Specialist Registrar Bursary. I agree to deposit this thesis in the University’s open access institutional repository or allow the Library to do so on my behalf, subject to Irish Copyright Legislation and Trinity College Library conditions of use and acknowledgement. I consent to the examiner retaining a copy of the thesis beyond the examining period, should they so wish (EU GDPR May 2018). _____________________ Dervla Connaughton 2 SUMMARY Chapter 1 provides an introduction to the topic while Chapter 2 provides details of the methods employed in this work. In Chapter 3 I provide an overview of the currently known monogenic causes for human chronic kidney disease (CKD). I also describe how next- generation sequencing can facilitate molecular genetic diagnostics in individuals with suspected genetic kidney disease. Chapter 4 details the findings of a multi-centre, cross-sectional study of patients with CKD in the Republic of Ireland. The primary aim of this study (the Irish Kidney Gene Project) was to describe the prevalence of reporting a positive family history of CKD among a representation sample of the CKD population. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -
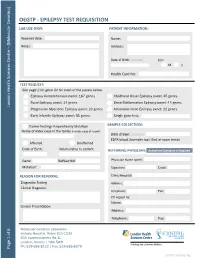
Oegtp - Epilepsy Test Requisition Lab Use Only: Patient Information
OEGTP - EPILEPSY TEST REQUISITION LAB USE ONLY: PATIENT INFORMATION: Received date: Name: Notes: Address: Date of Birth: YY/MM/DD Sex: M F Health Card No: TEST REQUEST: See page 2 for gene list for each of the panels below Epilepsy Comprehensive panel: 167 genes Childhood Onset Epilepsy panel: 45 genes Focal Epilepsy panel: 14 genes Brain Malformation Epilepsy panel: 44 genes London Health Sciences Centre – (Molecular Genetics) London Health Sciences Centre Progressive Myoclonic Epilepsy panel: 20 genes Actionable Gene Epilepsy panel: 22 genes Early Infantile Epilepsy panel: 51 genes Single gene test: Carrier Testing/ KnownFamily Mutation SAMPLE COLLECTION: Name of index case in the family (include copy of report) Date drawn: YY/MM/DD EDTA blood (lavender top) (5ml at room temp) Affected Unaffected Date of Birth: Relationship to patient: REFERRING PHYSICIAN: Authorized Signature is Required Gene: RefSeq:NM Physician Name (print): Mutation: Signature: Email: REASON FOR REFERRAL: Clinic/Hospital: Diagnostic Testing Address: Clinical Diagnosis: Telephone: Fax: CC report to: Name: Clinical Presentation: Address: Telephone: Fax: Molecular Genetics Laboratory Victoria Hospital, Room B10-123A 800 Commissioners Rd. E. London, Ontario | N6A 5W9 Pathology and Laboratory Medicine Ph: 519-685-8122 | Fax: 519-685-8279 Page 1 of 6 Page OEGTP (2021/05/28) OEGTP - EPILEPSY TEST PANELS Patient Identifier: COMPREHENSIVE EPILEPSY PANEL: 167 Genes ACTB, ACTG1, ADSL, AKT3, ALDH7A1, AMT, AP3B2, ARFGEF2, ARHGEF9, ARV1, ARX, ASAH1, ASNS, ATP1A3, ATP6V0A2, ATP7A, -

Anticancer Drug Oxaliplatin Induces Acute Cooling-Aggravated
Anticancer drug oxaliplatin induces acute cooling-aggravated neuropathy via sodium channel subtype NaV1.6-resurgent and persistent current Ruth Sittla,1, Angelika Lampertb,1, Tobias Huthb, E. Theresa Schuyb, Andrea S. Linkb, Johannes Fleckensteinc, Christian Alzheimerb, Peter Grafea, and Richard W. Carra,d,2 aInstitute of Physiology and cDepartment of Anesthesiology, Ludwig-Maximilians University, 80336 Munich, Germany; bInstitute of Physiology and Pathophysiology, Friedrich-Alexander Universität Erlangen-Nürnberg, 91054 Erlangen, Germany; and dDepartment of Anesthesia and Intensive Care Medicine, Medical Faculty Mannheim, Heidelberg University, 68167 Mannheim, Germany Edited by Richard W. Aldrich, University of Texas, Austin, TX, and approved March 8, 2012 (received for review November 2, 2011) Infusion of the chemotherapeutic agent oxaliplatin leads to an arrhythmia (NaV1.5) (14), paramyotonia congenita (Nav1.4) (15, acute and a chronic form of peripheral neuropathy. Acute oxaliplatin 16), and pain (Nav1.7) (15, 17). INaR was first described in cerebellar neuropathy is characterized by sensory paresthesias and muscle Purkinje neurons and refers to a transient surge of inward sodium cramps that are notably exacerbated by cooling. Painful dysesthesias current occurring upon repolarization from a preceding period of are rarely reported for acute oxaliplatin neuropathy, whereas a strong depolarization (18). Because of its unorthodox activation common symptom of chronic oxaliplatin neuropathy is pain. Here profile, INaR is thought to promote burst discharge (11, 12). we examine the role of the sodium channel isoform NaV1.6 in medi- Pain associated with paroxysmal extreme pain disorder (17) and ating the symptoms of acute oxaliplatin neuropathy. Compound and muscle cramps experienced by paramyotonia patients (16) are of- single-action potential recordings from human and mouse peripheral ten exacerbated or triggered by cooling, similar to the symptoms of axons showed that cooling in the presence of oxaliplatin (30–100 μM; acute oxaliplatin neuropathy. -

Dominant-Negative Calcium Channel Suppression by Truncated Constructs Involves a Kinase Implicated in the Unfolded Protein Response
5400 • The Journal of Neuroscience, June 9, 2004 • 24(23):5400–5409 Cellular/Molecular Dominant-Negative Calcium Channel Suppression by Truncated Constructs Involves a Kinase Implicated in the Unfolded Protein Response Karen M. Page,* Fay Heblich,* Anthony Davies,* Adrian J. Butcher,* Jeroˆme Leroy, Federica Bertaso, Wendy S. Pratt, and Annette C. Dolphin Department of Pharmacology, University College London, London WC1E 6BT, United Kingdom Expression of the calcium channel CaV2.2 is markedly suppressed by coexpression with truncated constructs of CaV2.2. Furthermore, a two-domain construct of CaV2.1 mimicking an episodic ataxia-2 mutation strongly inhibited CaV2.1 currents. We have now determined the specificity of this effect, identified a potential mechanism, and have shown that such constructs also inhibit endogenous calcium currents when transfected into neuronal cell lines. Suppression of calcium channel expression requires interaction between truncated and full-length channels, because there is inter-subfamily specificity. Although there is marked cross-suppression within the CaV2 calcium channel family, there is no cross-suppression between CaV2 and CaV3 channels. The mechanism involves activation of a compo- nent of the unfolded protein response, the endoplasmic reticulum resident RNA-dependent kinase (PERK), because it is inhibited by expression of dominant-negative constructs of this kinase. Activation of PERK has been shown previously to cause translational arrest, which has the potential to result in a generalized effect on protein synthesis. In agreement with this, coexpression of the truncated domain ␣ ␦ IofCaV2.2, together with full-length CaV2.2, reduced the level not only of CaV2.2 protein but also the coexpressed 2 -2. -

Inhibition of Radiation and Temozolomide-Induced Glioblastoma Invadopodia Activity Using Ion Channel Drugs
cancers Article Inhibition of Radiation and Temozolomide-Induced Glioblastoma Invadopodia Activity Using Ion Channel Drugs Marija Dinevska 1 , Natalia Gazibegovic 2 , Andrew P. Morokoff 1,3, Andrew H. Kaye 1,4, Katharine J. Drummond 1,3, Theo Mantamadiotis 1,5 and Stanley S. Stylli 1,3,* 1 Department of Surgery, The University of Melbourne, The Royal Melbourne Hospital, Parkville 3050, Victoria, Australia; [email protected] (M.D.); morokoff@unimelb.edu.au (A.P.M.); [email protected] (A.H.K.); [email protected] (K.J.D.); [email protected] (T.M.) 2 Victoria University, St. Albans 3021, Victoria, Australia; [email protected] 3 Department of Neurosurgery, The Royal Melbourne Hospital, Parkville 3050, Victoria, Australia 4 Hadassah University Medical Centre, Jerusalem 91120, Israel 5 Department of Microbiology & Immunology, School of Biomedical Sciences, The University of Melbourne, Parkville 3010, Victoria, Australia * Correspondence: [email protected] or [email protected] Received: 8 September 2020; Accepted: 30 September 2020; Published: 8 October 2020 Simple Summary: Glioblastoma accounts for approximately 40–50% of all primary brain cancers and is a highly aggressive cancer that rapidly disseminates within the surrounding normal brain. Dynamic actin-rich protrusions known as invadopodia facilitate this invasive process. Ion channels have also been linked to a pro-invasive phenotype and may contribute to facilitating invadopodia activity in cancer cells. The aim of our study was to screen ion channel-targeting drugs for their cytotoxic efficacy and potential anti-invadopodia properties in glioblastoma cells. We demonstrated that the targeting of ion channels in glioblastoma cells can lead to a reduction in invadopodia activity and protease secretion. -

Ion Channels 3 1
r r r Cell Signalling Biology Michael J. Berridge Module 3 Ion Channels 3 1 Module 3 Ion Channels Synopsis Ion channels have two main signalling functions: either they can generate second messengers or they can function as effectors by responding to such messengers. Their role in signal generation is mainly centred on the Ca2 + signalling pathway, which has a large number of Ca2+ entry channels and internal Ca2+ release channels, both of which contribute to the generation of Ca2 + signals. Ion channels are also important effectors in that they mediate the action of different intracellular signalling pathways. There are a large number of K+ channels and many of these function in different + aspects of cell signalling. The voltage-dependent K (KV) channels regulate membrane potential and + excitability. The inward rectifier K (Kir) channel family has a number of important groups of channels + + such as the G protein-gated inward rectifier K (GIRK) channels and the ATP-sensitive K (KATP) + + channels. The two-pore domain K (K2P) channels are responsible for the large background K current. Some of the actions of Ca2 + are carried out by Ca2+-sensitive K+ channels and Ca2+-sensitive Cl − channels. The latter are members of a large group of chloride channels and transporters with multiple functions. There is a large family of ATP-binding cassette (ABC) transporters some of which have a signalling role in that they extrude signalling components from the cell. One of the ABC transporters is the cystic − − fibrosis transmembrane conductance regulator (CFTR) that conducts anions (Cl and HCO3 )and contributes to the osmotic gradient for the parallel flow of water in various transporting epithelia. -

Spatial Distribution of Leading Pacemaker Sites in the Normal, Intact Rat Sinoa
Supplementary Material Supplementary Figure 1: Spatial distribution of leading pacemaker sites in the normal, intact rat sinoatrial 5 nodes (SAN) plotted along a normalized y-axis between the superior vena cava (SVC) and inferior vena 6 cava (IVC) and a scaled x-axis in millimeters (n = 8). Colors correspond to treatment condition (black: 7 baseline, blue: 100 µM Acetylcholine (ACh), red: 500 nM Isoproterenol (ISO)). 1 Supplementary Figure 2: Spatial distribution of leading pacemaker sites before and after surgical 3 separation of the rat SAN (n = 5). Top: Intact SAN preparations with leading pacemaker sites plotted during 4 baseline conditions. Bottom: Surgically cut SAN preparations with leading pacemaker sites plotted during 5 baseline conditions (black) and exposure to pharmacological stimulation (blue: 100 µM ACh, red: 500 nM 6 ISO). 2 a &DUGLDFIoQChDQQHOV .FQM FOXVWHU &DFQDG &DFQDK *MD &DFQJ .FQLS .FQG .FQK .FQM &DFQDF &DFQE .FQM í $WSD .FQD .FQM í .FQN &DVT 5\U .FQM &DFQJ &DFQDG ,WSU 6FQD &DFQDG .FQQ &DFQDJ &DFQDG .FQD .FQT 6FQD 3OQ 6FQD +FQ *MD ,WSU 6FQE +FQ *MG .FQN .FQQ .FQN .FQD .FQE .FQQ +FQ &DFQDD &DFQE &DOP .FQM .FQD .FQN .FQG .FQN &DOP 6FQD .FQD 6FQE 6FQD 6FQD ,WSU +FQ 6FQD 5\U 6FQD 6FQE 6FQD .FQQ .FQH 6FQD &DFQE 6FQE .FQM FOXVWHU V6$1 L6$1 5$ /$ 3 b &DUGLDFReFHSWRUV $GUDF FOXVWHU $GUDD &DY &KUQE &KUP &KJD 0\O 3GHG &KUQD $GUE $GUDG &KUQE 5JV í 9LS $GUDE 7SP í 5JV 7QQF 3GHE 0\K $GUE *QDL $QN $GUDD $QN $QN &KUP $GUDE $NDS $WSE 5DPS &KUP 0\O &KUQD 6UF &KUQH $GUE &KUQD FOXVWHU V6$1 L6$1 5$ /$ 4 c 1HXURQDOPURWHLQV -

Conditional Knockout of Nav1.6 in Adult Mice Ameliorates Neuropathic Pain
www.nature.com/scientificreports OPEN Conditional knockout of NaV1.6 in adult mice ameliorates neuropathic pain Received: 22 November 2017 Lubin Chen 1,2,3, Jianying Huang1,2,3, Peng Zhao1,2,3, Anna-Karin Persson1,2,3, Accepted: 19 February 2018 Fadia B. Dib-Hajj1,2,3, Xiaoyang Cheng1,2,3, Andrew Tan1,2,3, Stephen G. Waxman1,2,3 & Published: xx xx xxxx Sulayman D. Dib-Hajj 1,2,3 Voltage-gated sodium channels NaV1.7, NaV1.8 and NaV1.9 have been the focus for pain studies because their mutations are associated with human pain disorders, but the role of NaV1.6 in pain is less understood. In this study, we selectively knocked out NaV1.6 in dorsal root ganglion (DRG) neurons, using NaV1.8-Cre directed or adeno-associated virus (AAV)-Cre mediated approaches, and examined the specifc contribution of NaV1.6 to the tetrodotoxin-sensitive (TTX-S) current in these neurons and its role in neuropathic pain. We report here that NaV1.6 contributes up to 60% of the TTX-S current in large, and 34% in small DRG neurons. We also show NaV1.6 accumulates at nodes of Ranvier within the neuroma following spared nerve injury (SNI). Although NaV1.8-Cre driven NaV1.6 knockout does not alter acute, infammatory or neuropathic pain behaviors, AAV-Cre mediated NaV1.6 knockout in adult mice partially attenuates SNI-induced mechanical allodynia. Additionally, AAV-Cre mediated NaV1.6 knockout, mostly in large DRG neurons, signifcantly attenuates excitability of these neurons after SNI and reduces NaV1.6 accumulation at nodes of Ranvier at the neuroma. -

The L-Type Voltage-Gated Calcium Channel Cav1.2 Mediates Fear Extinction and Modulates Synaptic Tone in the Lateral Amygdala
Downloaded from learnmem.cshlp.org on September 26, 2021 - Published by Cold Spring Harbor Laboratory Press Research The L-type voltage-gated calcium channel CaV1.2 mediates fear extinction and modulates synaptic tone in the lateral amygdala Stephanie J. Temme1 and Geoffrey G. Murphy1,2,3 1Neuroscience Graduate Program; 2Molecular and Behavioral Neuroscience Institute; 3Department of Molecular and Integrative Physiology, University of Michigan, Ann Arbor, Michigan 48109-2200, USA L-type voltage-gated calcium channels (LVGCCs) have been implicated in both the formation and the reduction of fear through Pavlovian fear conditioning and extinction. Despite the implication of LVGCCs in fear learning and extinction, studies of the individual LVGCC subtypes, CaV1.2 and CaV1.3, using transgenic mice have failed to find a role of either subtype in fear extinction. This discontinuity between the pharmacological studies of LVGCCs and the studies investigating individual subtype contributions could be due to the limited neuronal deletion pattern of the CaV1.2 conditional knockout mice previously studied to excitatory neurons in the forebrain. To investigate the effects of deletion of CaV1.2 in all neu- ronal populations, we generated CaV1.2 conditional knockout mice using the synapsin1 promoter to drive Cre recombinase expression. Pan-neuronal deletion of CaV1.2 did not alter basal anxiety or fear learning. However, pan-neuronal deletion of CaV1.2 resulted in a significant deficit in extinction of contextual fear, implicating LVGCCs, specifically CaV1.2, in extinction learning. Further exploration on the effects of deletion of CaV1.2 on inhibitory and excitatory input onto the principle neurons of the lateral amygdala revealed a significant shift in inhibitory/excitatory balance. -

Application of Microrna Database Mining in Biomarker Discovery and Identification of Therapeutic Targets for Complex Disease
Article Application of microRNA Database Mining in Biomarker Discovery and Identification of Therapeutic Targets for Complex Disease Jennifer L. Major, Rushita A. Bagchi * and Julie Pires da Silva * Department of Medicine, Division of Cardiology, University of Colorado Anschutz Medical Campus, Aurora, CO 80045, USA; [email protected] * Correspondence: [email protected] (R.A.B.); [email protected] (J.P.d.S.) Supplementary Tables Methods Protoc. 2021, 4, 5. https://doi.org/10.3390/mps4010005 www.mdpi.com/journal/mps Methods Protoc. 2021, 4, 5. https://doi.org/10.3390/mps4010005 2 of 25 Table 1. List of all hsa-miRs identified by Human microRNA Disease Database (HMDD; v3.2) analysis. hsa-miRs were identified using the term “genetics” and “circulating” as input in HMDD. Targets CAD hsa-miR-1 Targets IR injury hsa-miR-423 Targets Obesity hsa-miR-499 hsa-miR-146a Circulating Obesity Genetics CAD hsa-miR-423 hsa-miR-146a Circulating CAD hsa-miR-149 hsa-miR-499 Circulating IR Injury hsa-miR-146a Circulating Obesity hsa-miR-122 Genetics Stroke Circulating CAD hsa-miR-122 Circulating Stroke hsa-miR-122 Genetics Obesity Circulating Stroke hsa-miR-26b hsa-miR-17 hsa-miR-223 Targets CAD hsa-miR-340 hsa-miR-34a hsa-miR-92a hsa-miR-126 Circulating Obesity Targets IR injury hsa-miR-21 hsa-miR-423 hsa-miR-126 hsa-miR-143 Targets Obesity hsa-miR-21 hsa-miR-223 hsa-miR-34a hsa-miR-17 Targets CAD hsa-miR-223 hsa-miR-92a hsa-miR-126 Targets IR injury hsa-miR-155 hsa-miR-21 Circulating CAD hsa-miR-126 hsa-miR-145 hsa-miR-21 Targets Obesity hsa-mir-223 hsa-mir-499 hsa-mir-574 Targets IR injury hsa-mir-21 Circulating IR injury Targets Obesity hsa-mir-21 Targets CAD hsa-mir-22 hsa-mir-133a Targets IR injury hsa-mir-155 hsa-mir-21 Circulating Stroke hsa-mir-145 hsa-mir-146b Targets Obesity hsa-mir-21 hsa-mir-29b Methods Protoc.