Computational Integration of Genome-Wide Observational And
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

9, 2015 Glasgow, Scotland, United Kingdom Abstracts
Volume 23 Supplement 1 June 2015 www.nature.com/ejhg European Human Genetics Conference 2015 June 6 - 9, 2015 Glasgow, Scotland, United Kingdom Abstracts EJHG_OFC.indd 1 4/1/2006 10:58:05 AM ABSTRACTS European Human Genetics Conference joint with the British Society of Genetics Medicine June 6 - 9, 2015 Glasgow, Scotland, United Kingdom Abstracts ESHG 2015 | GLASGOW, SCOTLAND, UK | WWW.ESHG.ORG 1 ABSTRACTS Committees – Board - Organisation European Society of Human Genetics ESHG Office Executive Board 2014-2015 Scientific Programme Committee European Society President Chair of Human Genetics Helena Kääriäinen, FI Brunhilde Wirth, DE Andrea Robinson Vice-President Members Karin Knob Han Brunner, NL Tara Clancy, UK c/o Vienna Medical Academy Martina Cornel, NL Alser Strasse 4 President-Elect Yanick Crow, FR 1090 Vienna Feliciano Ramos, ES Paul de Bakker, NL Austria Secretary-General Helene Dollfus, FR T: 0043 1 405 13 83 20 or 35 Gunnar Houge, NO David FitzPatrick, UK F: 0043 1 407 82 74 Maurizio Genuardi, IT E: [email protected] Deputy-Secretary-General Daniel Grinberg, ES www.eshg.org Karin Writzl, SI Gunnar Houge, NO Treasurer Erik Iwarsson, SE Andrew Read, UK Xavier Jeunemaitre, FR Mark Longmuir, UK Executive Officer Jose C. Machado, PT Jerome del Picchia, AT Dominic McMullan, UK Giovanni Neri, IT William Newman, UK Minna Nyström, FI Pia Ostergaard, UK Francesc Palau, ES Anita Rauch, CH Samuli Ripatti, FI Peter N. Robinson, DE Kristel van Steen, BE Joris Veltman, NL Joris Vermeesch, BE Emma Woodward, UK Karin Writzl, SI Board Members Liaison Members Yasemin Alanay, TR Stan Lyonnet, FR Martina Cornel, NL Martijn Breuning, NL Julie McGaughran, AU Ulf Kristoffersson, SE Pascal Borry, BE Bela Melegh, HU Thomas Liehr, DE Nina Canki-Klain, HR Will Newman, UK Milan Macek Jr., CZ Ana Carrió, ES Markus Nöthen, DE Tayfun Ozcelik, TR Isabella Ceccherini, IT Markus Perola, FI Milena Paneque, PT Angus John Clarke, UK Dijana Plaseska-Karanfilska, MK Hans Scheffer, NL Koen Devriendt, BE Trine E. -

Computational and Experimental Approaches for Evaluating the Genetic Basis of Mitochondrial Disorders
Computational and Experimental Approaches For Evaluating the Genetic Basis of Mitochondrial Disorders The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Lieber, Daniel Solomon. 2013. Computational and Experimental Approaches For Evaluating the Genetic Basis of Mitochondrial Disorders. Doctoral dissertation, Harvard University. Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:11158264 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA Computational and Experimental Approaches For Evaluating the Genetic Basis of Mitochondrial Disorders A dissertation presented by Daniel Solomon Lieber to The Committee on Higher Degrees in Systems Biology in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the subject of Systems Biology Harvard University Cambridge, Massachusetts April 2013 © 2013 - Daniel Solomon Lieber All rights reserved. Dissertation Adviser: Professor Vamsi K. Mootha Daniel Solomon Lieber Computational and Experimental Approaches For Evaluating the Genetic Basis of Mitochondrial Disorders Abstract Mitochondria are responsible for some of the cell’s most fundamental biological pathways and metabolic processes, including aerobic ATP production by the mitochondrial respiratory chain. In humans, mitochondrial dysfunction can lead to severe disorders of energy metabolism, which are collectively referred to as mitochondrial disorders and affect approximately 1:5,000 individuals. These disorders are clinically heterogeneous and can affect multiple organ systems, often within a single individual. Symptoms can include myopathy, exercise intolerance, hearing loss, blindness, stroke, seizures, diabetes, and GI dysmotility. -

Literature Mining Sustains and Enhances Knowledge Discovery from Omic Studies
LITERATURE MINING SUSTAINS AND ENHANCES KNOWLEDGE DISCOVERY FROM OMIC STUDIES by Rick Matthew Jordan B.S. Biology, University of Pittsburgh, 1996 M.S. Molecular Biology/Biotechnology, East Carolina University, 2001 M.S. Biomedical Informatics, University of Pittsburgh, 2005 Submitted to the Graduate Faculty of School of Medicine in partial fulfillment of the requirements for the degree of Doctor of Philosophy University of Pittsburgh 2016 UNIVERSITY OF PITTSBURGH SCHOOL OF MEDICINE This dissertation was presented by Rick Matthew Jordan It was defended on December 2, 2015 and approved by Shyam Visweswaran, M.D., Ph.D., Associate Professor Rebecca Jacobson, M.D., M.S., Professor Songjian Lu, Ph.D., Assistant Professor Dissertation Advisor: Vanathi Gopalakrishnan, Ph.D., Associate Professor ii Copyright © by Rick Matthew Jordan 2016 iii LITERATURE MINING SUSTAINS AND ENHANCES KNOWLEDGE DISCOVERY FROM OMIC STUDIES Rick Matthew Jordan, M.S. University of Pittsburgh, 2016 Genomic, proteomic and other experimentally generated data from studies of biological systems aiming to discover disease biomarkers are currently analyzed without sufficient supporting evidence from the literature due to complexities associated with automated processing. Extracting prior knowledge about markers associated with biological sample types and disease states from the literature is tedious, and little research has been performed to understand how to use this knowledge to inform the generation of classification models from ‘omic’ data. Using pathway analysis methods to better understand the underlying biology of complex diseases such as breast and lung cancers is state-of-the-art. However, the problem of how to combine literature- mining evidence with pathway analysis evidence is an open problem in biomedical informatics research. -

Complete Transcriptome Profiling of Normal and Age-Related Macular
www.nature.com/scientificreports OPEN Complete Transcriptome Profling of Normal and Age-Related Macular Degeneration Eye Tissues Reveals Received: 16 June 2017 Accepted: 30 January 2018 Dysregulation of Anti-Sense Published: xx xx xxxx Transcription Eun Ji Kim1, Gregory R. Grant1,2, Anita S. Bowman3,4, Naqi Haider3,4, Harini V. Gudiseva3 & Venkata Ramana Murthy Chavali3,4 Age-related macular degeneration (AMD) predominantly afects the retina and retinal pigment epithelium in the posterior eye. While there are numerous studies investigating the non-coding transcriptome of retina and RPE, few signifcant diferences between AMD and normal tissues have been reported. Strand specifc RNA sequencing of both peripheral retina (PR) and RPE-Choroid-Sclera (PRCS), in both AMD and matched normal controls were generated. The transcriptome analysis reveals a highly signifcant and consistent impact on anti-sense transcription as well as moderate changes in the regulation of non-coding (sense) RNA. Hundreds of genes that do not express anti-sense transcripts in normal PR and PRCS demonstrate signifcant anti-sense expression in AMD in all patient samples. Several pathways are highly enriched in the upregulated anti-sense transcripts—in particular the EIF2 signaling pathway. These results call for a deeper exploration into anti-sense and noncoding RNA regulation in AMD and their potential as therapeutic targets. Age-related macular degeneration (AMD) is the third largest cause of vision loss worldwide1. It is a progressive retinal disorder that involves loss of central vision, hypo- and hyper-pigmentation of the RPE, deposition of drusen in the Bruch’s membrane, and loss of photoreceptors, especially in the 8th or 9th decade2–4. -

Insights Into MYC Biology Through Investigation of Synthetic Lethal Interactions with MYC Deregulation
Insights into MYC biology through investigation of synthetic lethal interactions with MYC deregulation Mai Sato Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy under the Executive Committee of the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2014 © 2014 Mai Sato All Rights Reserved ABSTRACT Insights into MYC biology through investigation of synthetic lethal interactions with MYC deregulation Mai Sato MYC (or c-myc) is a bona fide “cancer driver” oncogene that is deregulated in up to 70% of human tumors. In addition to its well-characterized role as a transcription factor that can directly promote tumorigenic growth and proliferation, MYC has transcription-independent functions in vital cellular processes including DNA replication and protein synthesis, contributing to its complex biology. MYC expression, activity, and stability are highly regulated through multiple mechanisms. MYC deregulation triggers genome instability and oncogene-induced DNA replication stress, which are thought to be critical in promoting cancer via mechanisms that are still unclear. Because regulated MYC activity is essential for normal cell viability and MYC is a difficult protein to target pharmacologically, targeting genes or pathways that are essential to survive MYC deregulation offer an attractive alternative as a means to combat tumor cells with MYC deregulation. To this end, we conducted a genome-wide synthetic lethal shRNA screen in MCF10A breast epithelial cells stably expressing an inducible MYCER transgene. We identified and validated FBXW7 as a high-confidence synthetic lethal (MYC-SL) candidate gene. FBXW7 is a component of an E3 ubiquitin ligase complex that degrades MYC. FBXW7 knockdown in MCF10A cells selectively induced cell death in MYC-deregulated cells compared to control. -

Identification of Genomic Targets of Krüppel-Like Factor 9 in Mouse Hippocampal
Identification of Genomic Targets of Krüppel-like Factor 9 in Mouse Hippocampal Neurons: Evidence for a role in modulating peripheral circadian clocks by Joseph R. Knoedler A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Neuroscience) in the University of Michigan 2016 Doctoral Committee: Professor Robert J. Denver, Chair Professor Daniel Goldman Professor Diane Robins Professor Audrey Seasholtz Associate Professor Bing Ye ©Joseph R. Knoedler All Rights Reserved 2016 To my parents, who never once questioned my decision to become the other kind of doctor, And to Lucy, who has pushed me to be a better person from day one. ii Acknowledgements I have a huge number of people to thank for having made it to this point, so in no particular order: -I would like to thank my adviser, Dr. Robert J. Denver, for his guidance, encouragement, and patience over the last seven years; his mentorship has been indispensable for my growth as a scientist -I would also like to thank my committee members, Drs. Audrey Seasholtz, Dan Goldman, Diane Robins and Bing Ye, for their constructive feedback and their willingness to meet in a frequently cold, windowless room across campus from where they work -I am hugely indebted to Pia Bagamasbad and Yasuhiro Kyono for teaching me almost everything I know about molecular biology and bioinformatics, and to Arasakumar Subramani for his tireless work during the home stretch to my dissertation -I am grateful for the Neuroscience Program leadership and staff, in particular -

Agricultural University of Athens
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΤΩΝ ΖΩΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΗΣ ΖΩΙΚΗΣ ΠΑΡΑΓΩΓΗΣ ΕΡΓΑΣΤΗΡΙΟ ΓΕΝΙΚΗΣ ΚΑΙ ΕΙΔΙΚΗΣ ΖΩΟΤΕΧΝΙΑΣ ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ ΑΘΗΝΑ 2020 ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Genome-wide association analysis and gene network analysis for (re)production traits in commercial broilers ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ Τριμελής Επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Επταμελής εξεταστική επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Πηνελόπη Μπεμπέλη (Καθ. ΓΠΑ) Δημήτριος Βλαχάκης (Επ. Καθ. ΓΠΑ) Ευάγγελος Ζωίδης (Επ.Καθ. ΓΠΑ) Γεώργιος Θεοδώρου (Επ.Καθ. ΓΠΑ) 2 Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Περίληψη Σκοπός της παρούσας διδακτορικής διατριβής ήταν ο εντοπισμός γενετικών δεικτών και υποψηφίων γονιδίων που εμπλέκονται στο γενετικό έλεγχο δύο τυπικών πολυγονιδιακών ιδιοτήτων σε κρεοπαραγωγικά ορνίθια. Μία ιδιότητα σχετίζεται με την ανάπτυξη (σωματικό βάρος στις 35 ημέρες, ΣΒ) και η άλλη με την αναπαραγωγική -

WO 2013/095793 Al 27 June 2013 (27.06.2013) W P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2013/095793 Al 27 June 2013 (27.06.2013) W P O P C T (51) International Patent Classification: (81) Designated States (unless otherwise indicated, for every C12Q 1/68 (2006.01) kind of national protection available): AE, AG, AL, AM, AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, (21) International Application Number: BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, PCT/US2012/063579 DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, (22) International Filing Date: HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KM, KN, KP, 5 November 20 12 (05 .11.20 12) KR, KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, (25) Filing Language: English NO, NZ, OM, PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, (26) Publication Language: English RW, SC, SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, (30) Priority Data: ZM, ZW. 61/579,530 22 December 201 1 (22. 12.201 1) US (84) Designated States (unless otherwise indicated, for every (71) Applicant: AVEO PHARMACEUTICALS, INC. kind of regional protection available): ARIPO (BW, GH, [US/US]; 75 Sidney Street, Fourth Floor, Cambridge, MA GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, SZ, TZ, 02139 (US). -
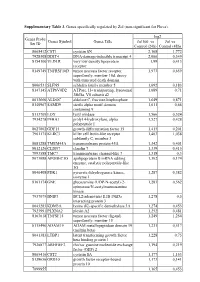
Supplementary Table 3. Genes Specifically Regulated by Zol (Non-Significant for Fluva)
Supplementary Table 3. Genes specifically regulated by Zol (non-significant for Fluva). log2 Genes Probe Genes Symbol Genes Title Zol100 vs Zol vs Set ID Control (24h) Control (48h) 8065412 CST1 cystatin SN 2,168 1,772 7928308 DDIT4 DNA-damage-inducible transcript 4 2,066 0,349 8154100 VLDLR very low density lipoprotein 1,99 0,413 receptor 8149749 TNFRSF10D tumor necrosis factor receptor 1,973 0,659 superfamily, member 10d, decoy with truncated death domain 8006531 SLFN5 schlafen family member 5 1,692 0,183 8147145 ATP6V0D2 ATPase, H+ transporting, lysosomal 1,689 0,71 38kDa, V0 subunit d2 8013660 ALDOC aldolase C, fructose-bisphosphate 1,649 0,871 8140967 SAMD9 sterile alpha motif domain 1,611 0,66 containing 9 8113709 LOX lysyl oxidase 1,566 0,524 7934278 P4HA1 prolyl 4-hydroxylase, alpha 1,527 0,428 polypeptide I 8027002 GDF15 growth differentiation factor 15 1,415 0,201 7961175 KLRC3 killer cell lectin-like receptor 1,403 1,038 subfamily C, member 3 8081288 TMEM45A transmembrane protein 45A 1,342 0,401 8012126 CLDN7 claudin 7 1,339 0,415 7993588 TMC7 transmembrane channel-like 7 1,318 0,3 8073088 APOBEC3G apolipoprotein B mRNA editing 1,302 0,174 enzyme, catalytic polypeptide-like 3G 8046408 PDK1 pyruvate dehydrogenase kinase, 1,287 0,382 isozyme 1 8161174 GNE glucosamine (UDP-N-acetyl)-2- 1,283 0,562 epimerase/N-acetylmannosamine kinase 7937079 BNIP3 BCL2/adenovirus E1B 19kDa 1,278 0,5 interacting protein 3 8043283 KDM3A lysine (K)-specific demethylase 3A 1,274 0,453 7923991 PLXNA2 plexin A2 1,252 0,481 8163618 TNFSF15 tumor necrosis -

Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations Fall 2010 Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress Renuka Nayak University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Computational Biology Commons, and the Genomics Commons Recommended Citation Nayak, Renuka, "Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress" (2010). Publicly Accessible Penn Dissertations. 1559. https://repository.upenn.edu/edissertations/1559 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/1559 For more information, please contact [email protected]. Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress Abstract Genes interact in networks to orchestrate cellular processes. Here, we used coexpression networks based on natural variation in gene expression to study the functions and interactions of human genes. We asked how these networks change in response to stress. First, we studied human coexpression networks at baseline. We constructed networks by identifying correlations in expression levels of 8.9 million gene pairs in immortalized B cells from 295 individuals comprising three independent samples. The resulting networks allowed us to infer interactions between biological processes. We used the network to predict the functions of poorly-characterized human genes, and provided some experimental support. Examining genes implicated in disease, we found that IFIH1, a diabetes susceptibility gene, interacts with YES1, which affects glucose transport. Genes predisposing to the same diseases are clustered non-randomly in the network, suggesting that the network may be used to identify candidate genes that influence disease susceptibility. -

Ryan D. Hernandez
Natural Selection Ryan D. Hernandez [email protected] 1 The Effect of Positive Selection Adaptive Neutral Nearly Neutral Mildly Deleterious Fairly Deleterious Strongly Deleterious "2 The Effect of Positive Selection Adaptive! Neutral Nearly Neutral Mildly Deleterious Fairly Deleterious Strongly Deleterious "3 Coding regions tend to have the lowest levels of diversity in the genome 4 What are the predominant evolutionary forces driving human genomes?! Eyre-Walker & Keightley ~40% of amino acid substitutions (2009) were advantageous 10-20% of amino acid substitutions Boyko et al (2008) were advantageous 10% of the genome affected by Williamson et al (2007) selective sweeps 5 Diversity levels around a selective sweep Thornton et al (2007): Simulation of patterns of neutral diversity around a selective sweep 6 The footprint of adaptive amino acid substitutions Neutral diversity levels … Amino acid substitution Reflects the typical strength of selection Reflects the fraction of • Goal: compare the pattern amino acid substitutions around amino acid that are adaptive substitutions to the pattern … n substitutions around synonymous substitutions. 7 Other organisms... Drosophila NS SYN Sattath et al (2011) estimate ~13% of amino acid substitutions were adaptive. 8 Observed Patterns of Diversity Around Human Substitutions 9 Hernandez, et al. Science (2011) Other organisms... Drosophila Chimpanzee NS SYN NATURE PLANTS DOI: 10.1038/NPLANTS.2016.84 ARTICLES Maize Teosinte a 1.1 b 1.1 0.008 0.013 1.0 1.0 0.007 Pairwise diversity Pairwise 0.011 0.9 0.9 diversity Pairwise 0.8 0.006 0.8 0.009 0.7 0.7 0.005 1.2 0.009 1.2 0.6 1.0 0.6 0.013 0.8 0.006 0.007 Diversity/neutral diversity Diversity/neutral Diversity/neutral diversity Diversity/neutral 0.004 0.8 0.009 0.6 0.5 0.5 0.4 0.003 Synonymous 0.4 0.005 Synonymous −0.15 −0.05 0.05 0.15 0.4 Non-synonymous 0.003 0.4 −0.15 −0.05 0.05 0.15 Non-synonymous 0.005 Beissinger , −0.002 −0.001 0.000 0.001 0.002 −0.002 −0.001 0.000 0.001 0.002 10 Distance to nearest substitution (cM) Distance to nearest substitution (cM) et al.