A Tour of the Cell
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Nature of Genomes Viral Genomes Prokaryotic Genome
The nature of genomes • Genomics: study of structure and function of genomes • Genome size – variable, by orders of magnitude – number of genes roughly proportional to genome size • Plasmids – symbiotic DNA molecules, not essential – mostly circular in prokaryotes • Organellar DNA – chloroplast, mitochondrion – derived by endosymbiosis from bacterial ancestors Chapter 2: Genes and genomes © 2002 by W. H. Freeman and Company Chapter 2: Genes and genomes © 2002 by W. H. Freeman and Company Viral genomes • Nonliving particle In prokaryotes, viruses are – nucleic acid sometimes referred to as – protein bacteriophages. • DNA or RNA – single-stranded or double-stranded – linear or circular • Compact genomes with little spacer DNA Chapter 2: Genes and genomes © 2002 by W. H. Freeman and Company Chapter 2: Genes and genomes © 2002 by W. H. Freeman and Company Prokaryotic genome • Usually circular double helix – occupies nucleoid region of cell – attached to plasma membrane • Genes are close together with little intergenic spacer • Operon – tandem cluster of coordinately regulated genes – transcribed as single mRNA • Introns very rare Chapter 2: Genes and genomes © 2002 by W. H. Freeman and Company Chapter 2: Genes and genomes © 2002 by W. H. Freeman and Company 1 Eukaryotic nuclear genomes • Each species has characteristic chromosome number • Genes are segments of nuclear chromosomes • Ploidy refers to number of complete sets of chromosomes –haploid (1n): one complete set of genes – diploid (2n) – polyploid (≥3n) • In diploids, chromosomes come in homologous pairs (homologs) In humans, somatic cells have – structurally similar 2n = 46 chromosomes. – same sequence of genes – may contain different alleles Chapter 2: Genes and genomes © 2002 by W. H. -

PEX5 Regulates Autophagy Via the Mtorc1-TFEB Axis During Starvation
Eun et al. Experimental & Molecular Medicine (2018) 50:4 DOI 10.1038/s12276-017-0007-8 Experimental & Molecular Medicine ARTICLE Open Access PEX5 regulates autophagy via the mTORC1-TFEB axis during starvation So Young Eun1,JoonNoLee2,In-KooNam2, Zhi-qiang Liu1,Hong-SeobSo 1, Seong-Kyu Choe1 and RaeKil Park2 Abstract Defects in the PEX5 gene impair the import of peroxisomal matrix proteins, leading to nonfunctional peroxisomes and other associated pathological defects such as Zellweger syndrome. Although PEX5 regulates autophagy process in a stress condition, the mechanisms controlling autophagy by PEX5 under nutrient deprivation are largely unknown. Herein, we show a novel function of PEX5 in the regulation of autophagy via Transcription Factor EB (TFEB). Under serum-starved conditions, when PEX5 is depleted, the mammalian target of rapamycin (mTORC1) inhibitor TSC2 is downregulated, which results in increased phosphorylation of the mTORC1 substrates, including 70S6K, S6K, and 4E- BP-1. mTORC1 activation further suppresses the nuclear localization of TFEB, as indicated by decreased mRNA levels of TFEB, LIPA, and LAMP1. Interestingly, peroxisomal mRNA and protein levels are also reduced by TFEB inactivation, indicating that TFEB might control peroxisome biogenesis at a transcriptional level. Conversely, pharmacological inhibition of mTOR resulting from PEX5 depletion during nutrient starvation activates TFEB by promoting nuclear localization of the protein. In addition, mTORC1 inhibition recovers the damaged-peroxisome biogenesis. These data suggest that PEX5 may be a critical regulator of lysosomal gene expression and autophagy through the mTOR-TFEB- autophagy axis under nutrient deprivation. 1234567890():,; 1234567890():,; Introduction Mitochondrial antiviral-signaling protein (MAVS) func- Peroxisome is an essential cellular organelle for per- tions as an antiviral signaling platform to induce the forming various metabolic activities, including oxidation interferon-independent signaling pathways4. -
Chapter 4 – Cell Structure
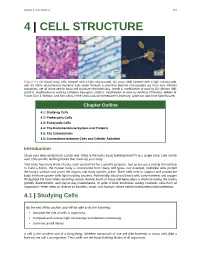
Chapter 4 | Cell Structure 107 4 | CELL STRUCTURE Figure 4.1 (a) Nasal sinus cells (viewed with a light microscope), (b) onion cells (viewed with a light microscope), and (c) Vibrio tasmaniensis bacterial cells (seen through a scanning electron microscope) are from very different organisms, yet all share certain basic cell structure characteristics. (credit a: modification of work by Ed Uthman, MD; credit b: modification of work by Umberto Salvagnin; credit c: modification of work by Anthony D'Onofrio, William H. Fowle, Eric J. Stewart, and Kim Lewis of the Lewis Lab at Northeastern University; scale-bar data from Matt Russell) Chapter Outline 4.1: Studying Cells 4.2: Prokaryotic Cells 4.3: Eukaryotic Cells 4.4: The Endomembrane System and Proteins 4.5: The Cytoskeleton 4.6: Connections between Cells and Cellular Activities Introduction Close your eyes and picture a brick wall. What is the wall's basic building block? It is a single brick. Like a brick wall, cells are the building blocks that make up your body. Your body has many kinds of cells, each specialized for a specific purpose. Just as we use a variety of materials to build a home, the human body is constructed from many cell types. For example, epithelial cells protect the body's surface and cover the organs and body cavities within. Bone cells help to support and protect the body. Immune system cells fight invading bacteria. Additionally, blood and blood cells carry nutrients and oxygen throughout the body while removing carbon dioxide. Each of these cell types plays a vital role during the body's growth, development, and day-to-day maintenance. -

The Association of Peroxisomes with the Developing Cell Plate in Dividing Onion Root Cells Depends on Actin Microfilaments and Myosin
Planta (2003) 218: 204–216 DOI 10.1007/s00425-003-1096-2 ORIGINAL ARTICLE David A. Collings Æ John D. I. Harper Æ Kevin C. Vaughn The association of peroxisomes with the developing cell plate in dividing onion root cells depends on actin microfilaments and myosin Received: 5 April 2003 / Accepted: 23 June 2003 / Published online: 21 August 2003 Ó Springer-Verlag 2003 Abstract We have investigated changes in the distribu- cling of excess membranes from secretory vesicles via the tion of peroxisomes through the cell cycle in onion b-oxidation pathway. Differences in aggregation, a (Allium cepa L.) root meristem cells with immunofluo- phenomenon which occurs in onion, some other mono- rescence and electron microscopy, and in leek (Allium cots and to a lesser extent in tobacco BY-2 suspension porrum L.) epidermal cells with immunofluorescence and cells, but which is not obvious in the roots of Arabidopsis peroxisomal-targeted green fluorescent protein. During thaliana (L.) Heynh., may reflect differences within the interphase and mitosis, peroxisomes distribute randomly primary cell walls of these plants. throughout the cytoplasm, but beginning late in ana- phase, they accumulate at the division plane. Initially, Keywords Actin microfilaments Æ Allium Æ peroxisomes occur within the microtubule phragmoplast Microtubule Æ Cell plate Æ Peroxisome Æ Phragmoplast in two zones on either side of the developing cell plate. However, as the phragmoplast expands outwards to Abbreviations BDM: 2,3-butanedione monoxime Æ DAPI: form an annulus, peroxisomes redistribute into a ring 4¢,6-diamidino-2-phenylindole Æ ER: endoplasmic reti- immediately inside the location of the microtubules. -

Hif-2A Promotes Degradation of Mammalian Peroxisomes by Selective Autophagy
Cell Metabolism Article Hif-2a Promotes Degradation of Mammalian Peroxisomes by Selective Autophagy Katharina M. Walter,1,2,10 Miriam J. Scho¨ nenberger,1,2,10 Martin Tro¨ tzmu¨ ller,3 Michael Horn,1 Hans-Peter Elsa¨ sser,4 Ann B. Moser,5 Miriam S. Lucas,6 Tobias Schwarz,6 Philipp A. Gerber,7 Phyllis L. Faust,8 Holger Moch,9 Harald C. Ko¨ feler,3 Wilhelm Krek,1,2,* and Werner J. Kovacs1,2,* 1Institute of Molecular Health Sciences 2Competence Center for Systems Physiology and Metabolic Diseases ETH Zurich, CH-8093 Zurich, Switzerland 3Core Facility for Mass Spectrometry, Center for Medical Research, Medical University of Graz, A-8010 Graz, Austria 4Department of Cytobiology, Philipps-University Marburg, D-35037 Marburg, Germany 5Kennedy Krieger Institute, Baltimore, MD 21205, USA 6ScopeM – Scientific Center for Optical and Electron Microscopy, ETH Zurich, CH-8093 Zurich, Switzerland 7Division of Endocrinology and Diabetes, University Hospital Zurich, CH-8091 Zurich, Switzerland 8Department of Pathology and Cell Biology, College of Physicians and Surgeons, Columbia University, New York, NY 10032, USA 9Institute of Surgical Pathology, University Hospital Zurich, CH-8091 Zurich, Switzerland 10Co-first Authors *Correspondence: [email protected] (W.K.), [email protected] (W.J.K.) http://dx.doi.org/10.1016/j.cmet.2014.09.017 SUMMARY expressed HIF-1b subunit and O2-regulated a subunits (HIF-1a and HIF-2a)(Keith et al., 2012). Under normoxia, HIF-a subunits Peroxisomes play a central role in lipid metabolism, are hydroxylated and targeted for proteasomal degradation by and their function depends on molecular oxygen. -

The Mitochondrial Genome. the Nucleoid
ISSN 0006-2979, Biochemistry (Moscow), 2016, Vol. 81, No. 10, pp. 1057-1065. © Pleiades Publishing, Ltd., 2016. Original Russian Text © A. A. Kolesnikov, 2016, published in Biokhimiya, 2016, Vol. 81, No. 10, pp. 1322-1331. REVIEW The Mitochondrial Genome. The Nucleoid A. A. Kolesnikov Lomonosov Moscow State University, Faculty of Biology, 119991 Moscow, Russia; E-mail: [email protected] Received May 30, 2016 Revision received July 1, 2016 Abstract—Mitochondrial DNA (mtDNA) in cells is organized in nucleoids containing DNA and various proteins. This review discusses questions of organization and structural dynamics of nucleoids as well as their protein components. The structures of mt-nucleoid from different organisms are compared. The currently accepted model of nucleoid organization is described and questions needing answers for better understanding of the fine mechanisms of the mitochondrial genetic apparatus functioning are discussed. DOI: 10.1134/S0006297916100047 Key words: mitochondrial nucleoid, mitochondrial genome It is now clear that understanding of genome organ- mtDNA). In fact, proteins associated with a nucleoid can ization of subcellular structures is necessary for under- influence the speed of accumulation of mutations. standing of cellular processes occurring through the cell In the last decade, reviews devoted to this subject cycle, including questions of storage, realization, and appeared regularly [5-9]. Generally, they cover the subject transmission of genetic information. A considerable of nucleoid organization in metazoan mitochondria amount of information about forms and sizes of mito- (mainly in human cells) and fungi (mainly yeast). chondrial DNA (mtDNA) throughout evolution has been Reviews touching plant cells are much less frequent [10, accumulated [1]. -

Autophagy Stimulus-Dependent Role of the Small Gtpase Ras2 in Peroxisome Degradation
biomolecules Communication Autophagy Stimulus-Dependent Role of the Small GTPase Ras2 in Peroxisome Degradation Fahd Boutouja 1,2 and Harald W. Platta 1,* 1 Biochemie Intrazellulärer Transportprozesse, Ruhr-Universität Bochum, 44801 Bochum, Germany; [email protected] 2 Institute of Pathobiochemistry, Johannes Gutenberg-University Mainz, 55099 Mainz, Germany * Correspondence: [email protected]; Tel.: +49-234-322-4968 Received: 17 October 2020; Accepted: 12 November 2020; Published: 14 November 2020 Abstract: The changing accessibility of nutrient resources induces the reprogramming of cellular metabolism in order to adapt the cell to the altered growth conditions. The nutrient-depending signaling depends on the kinases mTOR (mechanistic target of rapamycin), which is mainly activated by nitrogen-resources, and PKA (protein kinase A), which is mainly activated by glucose, as well as both of their associated factors. These systems promote protein synthesis and cell proliferation, while they inhibit degradation of cellular content by unselective bulk autophagy. Much less is known about their role in selective autophagy pathways, which have a more regulated cellular function. Especially, we were interested to analyse the central Ras2-module of the PKA-pathway in the context of peroxisome degradation. Yeast Ras2 is homologous to the mammalian Ras proteins, whose mutant forms are responsible for 33% of human cancers. In the present study, we were able to demonstrate a context-dependent role of Ras2 activity depending on the type of mTOR-inhibition and glucose-sensing situation. When mTOR was inhibited directly via the macrolide rapamycin, peroxisome degradation was still partially suppressed by Ras2, while inactivation of Ras2 resulted in an enhanced degradation of peroxisomes, suggesting a role of Ras2 in the inhibition of peroxisome degradation in glucose-grown cells. -
Basic Features of All Cells
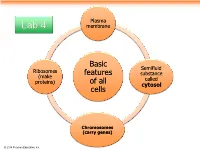
Plasma Lab 4 membrane Basic Semifluid Ribosomes features substance (make called proteins) of all cytosol cells Chromosomes (carry genes) © 2014 Pearson Education, Inc. Prokaryotic cells are characterized by having . No nucleus . DNA in an unbound region called the nucleoid . No membrane-bound organelles . Cytoplasm bound by the plasma membrane © 2014 Pearson Education, Inc. Eukaryotic cells are characterized by having • DNA in a nucleus that is bounded by a membranous nuclear envelope • Membrane-bound organelles • Cytoplasm in the region between the plasma membrane and nucleus Eukaryotic cells are generally much larger than prokaryotic cells © 2014 Pearson Education, Inc. Figure 6.8a ENDOPLASMIC RETICULUM (ER) Nuclear envelope Rough ER Smooth ER Nucleolus NUCLEUS Flagellum Chromatin Centrosome Plasma membrane CYTOSKELETON: Microfilaments Intermediate filaments Microtubules Ribosomes Microvilli Golgi apparatus Peroxisome Lysosome Mitochondrion © 2014 Pearson Education, Inc. Figure 6.8b Nuclear envelope NUCLEUS Nucleolus Rough ER Chromatin Smooth ER Ribosomes Golgi Central vacuole apparatus Microfilaments CYTOSKELETON Microtubules Mitochondrion Peroxisome Plasma Chloroplast membrane Cell wall Plasmodesmata Wall of adjacent cell © 2014 Pearson Education, Inc. © 2014 Pearson Education, Inc. © 2014 Pearson Education, Inc. © 2014 Pearson Education, Inc. Table 6.1 © 2014 Pearson Education, Inc. Cell Walls of Plants The cell wall is an extracellular structure that distinguishes plant cells from animal cells Plant cell walls Prokaryotes, are made -

Coli Chromosome Into a Nucleoid Filament
Strong intranucleoid interactions organize the Escherichia coli chromosome into a nucleoid filament Paul A. Wigginsa,1, Keith C. Cheverallsa,b, Joshua S. Martina,b, Robert Lintnera, and Jané Kondevb aWhitehead Institute for Biomedical Research, 9 Cambridge Center, Cambridge, MA 02142; and bMartin A. Fisher School of Physics, Brandeis University, 415 South Street, Waltham, MA 02453 Edited by Nancy E Kleckner, Harvard University, Cambridge, MA, and approved January 27, 2010 (received for review October 26, 2009) The stochasticity of chromosome organization was investigated by Our quantitative measurements of E. coli nucleoid structure con- fluorescently labeling genetic loci in live Escherichia coli cells. In firm this qualitative picture: The body of the nucleoid is linearly spite of the common assumption that the chromosome is well organized along the long-axis of the cell, implying that the modeled by an unstructured polymer, measurements of the locus nucleoid has a nearly-constant linear packing density, except distributions reveal that the E. coli chromosome is precisely orga- for a short region, genomically ter-proximate, which connects nized into a nucleoid filament with a linear order. Loci in the body the two arms of the chromosome (11). But in spite of the obser- of the nucleoid show a precision of positioning within the cell of vation of a linear chromosome organization in both E. coli and better than 10% of the cell length. The precision of interlocus dis- C. crescentus, the mechanism which gives rise to this characteristic tance of genomically-proximate loci was better than 4% of the cell structure is unknown. length. The measured dependence of the precision of interlocus To probe the mechanism of nucleoid organization, we measure distance on genomic distance singles out intranucleoid interactions and analyze the position fluctuations of single loci and the cor- as the mechanism responsible for chromosome organization. -

The Membrane Remodeling Protein Pex11p Activates the Gtpase Dnm1p During Peroxisomal Fission
The membrane remodeling protein Pex11p activates the GTPase Dnm1p during peroxisomal fission Chris Williamsa, Lukasz Opalinskia,b,1, Christiane Landgrafc, Joseph Costellod, Michael Schraderd, Arjen M. Krikkena,b, Kèvin Knoopsa, Anita M. Krama,b, Rudolf Volkmerc,e, and Ida J. van der Kleia,b,2 aMolecular Cell Biology, Groningen Biomolecular Sciences and Biotechnology Institute, and bKluyver Centre for Genomics of Industrial Fermentation, University of Groningen, 9747 AG Groningen, The Netherlands; cInstitut für Medizinische Immunologie, Charité-Universitätsmedizin Berlin, 10115 Berlin, Germany; dCollege of Life and Environmental Sciences, Biosciences, University of Exeter, Exeter EX4 4QD, United Kingdom; and eLeibniz-Institut für Molekulare Pharmakologie, 13125 Berlin, Germany Edited by Jennifer Lippincott-Schwartz, National Institutes of Health, Bethesda, MD, and approved April 14, 2015 (received for review October 9, 2014) The initial phase of peroxisomal fission requires the peroxisomal Fis1p and (in S. cerevisiae) the accessory proteins Mdv1p and membrane protein Peroxin 11 (Pex11p), which remodels the mem- Caf4p (12). Interestingly these proteins are also responsible for brane, resulting in organelle elongation. Here, we identify an ad- mitochondrial fission in yeast (13). ditional function for Pex11p, demonstrating that Pex11p also plays Dnm1p (Drp1 in mammals) (11, 14) is a large GTPase that a crucial role in the final step of peroxisomal fission: dynamin-like achieves membrane fission by forming oligomeric, ring-like struc- protein (DLP)-mediated membrane scission. First, we demonstrate tures around constricted sites on organelle membranes (15). Powered that yeast Pex11p is necessary for the function of the GTPase by GTP hydrolysis, these ring-like structures then tighten further Dynamin-related 1 (Dnm1p) in vivo. -

The Chloroplast Nucleoid in Ochromonas Danica I
J. Cell Sci. 16, 557-577 0974) 557 Printed in Great Britain THE CHLOROPLAST NUCLEOID IN OCHROMONAS DANICA I. THREE-DIMENSIONAL MORPHOLOGY IN LIGHT- AND DARK-GROWN CELLS SARAH P. GIBBS, D. CHENG AND T. SLANKIS Department of Biology, McGill University, Montreal HT,C 3G1, Canada SUMMARY The 3-dimensional structure of the plastid nucleoid was determined from serial sections of the plastid of dark-grown, greening, and light-grown cells of Ochromonas danica. In light- grown and greening cells, the chloroplast nucleoid forms a continuous cord or ring which closely follows the rim of each lateral lobe of the chloroplast and is continuous across the top and bottom of the bridge connecting the 2 chloroplast lobes. The nucleoid always lies just inside the chloroplast girdle bands where they loop around the rim of the plastid. It was demonstrated by electron-microscopic autoradiography of greening cells labelled with [3H]- thymidine that all the plastid DNA is localized in this peripheral ring-shaped nucleoid. In the proplastid of dark-grown cells, the nucleoid also forms a ring-shaped structure lying just inside the single girdle thylakoid, although frequent irregularities, such as gaps, are present. It is postulated that the girdle bands determine the shape of the chloroplast nucleoid, possibly by having specific attachment sites for the plastid DNA molecules. A survey of the literature shows that a peripheral ring-shaped chloroplast nucleoid is a characteristic feature of the 5 classes of algae whose chloroplasts possess girdle bands, namely the Raphidophyceae, Chryso- phyceae, Bacillariophyceae, Xanthophyceae, and Phaeophyceae, and has never been observed in plants whose plastids lack girdle bands. -

In Situ Imaging of Mitochondrial Translation Shows Weak Correlation
© 2017. Published by The Company of Biologists Ltd | Journal of Cell Science (2017) 130, 4193-4199 doi:10.1242/jcs.206714 TOOLS AND RESOURCES In situ imaging of mitochondrial translation shows weak correlation with nucleoid DNA intensity and no suppression during mitosis Christopher Estell1, Emmanouela Stamatidou2, Sarah El-Messeiry2,3 and Andrew Hamilton2,* ABSTRACT not been tested because no method exists to examine total de novo Although mitochondrial translation produces only 13 proteins, we show mitochondrial genome and expression simultaneously in situ here how this process can be visualised and detected in situ . Mitochondrial genome polymorphisms (heteroplasmy) is by fluorescence microscopy with a simple, rapid and inexpensive also common in mammalian cells (Larsson, 2010; He et al., 2010), procedure using non-canonical amino acid labelling and click partly due to the much higher mutation rate of mitochondrial DNA chemistry. This allows visualisation of the translational output in compared to nuclear DNA (Taylor et al., 2003) but, again, it is not different mitochondria within a cell, their position within that cell and a known how such sequence variation affects overall genome comparison of mitochondrial translation between cells. The most highly expression. Heterogeneity of mitochondrial membrane potential translationally active mitochondria were closest to the nucleus but were and activity of some mitochondrial enzymes have been observed also found at the distal end of long cellular projections. There were and associated with disease (Wikstrom et al., 2009), in a way similar substantial differences in translation between adjacent mitochondria to that of genetic polymorphism. However, the extent to which and this did not readily correlate with apparent mitochondrial genome such physiological heterogeneity arises from mitochondrial gene content.