Text Mining Applied to Molecular Biology
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genome-Wide Association Identifies Candidate Genes That Influence The
Genome-wide association identifies candidate genes that influence the human electroencephalogram Colin A. Hodgkinsona,1, Mary-Anne Enocha, Vibhuti Srivastavaa, Justine S. Cummins-Omana, Cherisse Ferriera, Polina Iarikovaa, Sriram Sankararamanb, Goli Yaminia, Qiaoping Yuana, Zhifeng Zhoua, Bernard Albaughc, Kenneth V. Whitea, Pei-Hong Shena, and David Goldmana aLaboratory of Neurogenetics, National Institute on Alcohol Abuse and Alcoholism, Rockville, MD 20852; bComputer Science Department, University of California, Berkeley, CA 94720; and cCenter for Human Behavior Studies, Weatherford, OK 73096 Edited* by Raymond L. White, University of California, Emeryville, CA, and approved March 31, 2010 (received for review July 23, 2009) Complex psychiatric disorders are resistant to whole-genome reflects rhythmic electrical activity of the brain. EEG patterns analysis due to genetic and etiological heterogeneity. Variation in dynamically and quantitatively index cortical activation, cognitive resting electroencephalogram (EEG) is associated with common, function, and state of consciousness. EEG traits were among the complex psychiatric diseases including alcoholism, schizophrenia, original intermediate phenotypes in neuropsychiatry, having been and anxiety disorders, although not diagnostic for any of them. EEG first recorded in humans in 1924 by Hans Berger, who documented traits for an individual are stable, variable between individuals, and the α rhythm, seen maximally during states of relaxation with eyes moderately to highly heritable. Such intermediate phenotypes closed, and supplanted by faster β waves during mental activity. appear to be closer to underlying molecular processes than are EEG can be used clinically for the evaluation and differential di- clinical symptoms, and represent an alternative approach for the agnosis of epilepsy and sleep disorders, differentiation of en- identification of genetic variation that underlies complex psychiat- cephalopathy from catatonia, assessment of depth of anesthesia, ric disorders. -

Identification of Common Key Genes in Breast, Lung and Prostate Cancer and Exploration of Their Heterogeneous Expression
1680 ONCOLOGY LETTERS 15: 1680-1690, 2018 Identification of common key genes in breast, lung and prostate cancer and exploration of their heterogeneous expression RICHA K. MAKHIJANI1, SHITAL A. RAUT1 and HEMANT J. PUROHIT2 1Department of Computer Science and Engineering, Visvesvaraya National Institute of Technology, Nagpur, Maharashtra 440010; 2Environmental Genomics Division, National Environmental Engineering Research Institute, Nagpur, Maharashtra 440020, India Received March 8, 2017; Accepted October 20, 2017 DOI: 10.3892/ol.2017.7508 Abstract. Cancer is one of the leading causes of mortality cancer and may be used as biomarkers for the development of worldwide, and in particular, breast cancer in women, prostate novel diagnostic and therapeutic strategies. cancer in men, and lung cancer in both women and men. The present study aimed to identify a common set of genes which Introduction may serve as indicators of important molecular and cellular processes in breast, prostate and lung cancer. Six microarray The highest rates of cancer-related mortality are associated gene expression profile datasets [GSE45827, GSE48984, with breast, prostate and lung cancer, as reported by the World GSE19804, GSE10072, GSE55945 and GSE26910 (two Health Organization (1), the World Cancer Report (2) and datasets for each cancer)] and one RNA-Seq expression dataset Cancer facts and figures (3) A plethora of cancer microarray (GSE62944 including all three cancer types), were downloaded and RNA sequencing (RNA-Seq) studies are publicly avail- from the Gene Expression Omnibus database. Differentially able in databases, including the Gene Expression Omnibus expressed genes (DEGs) were identified in each individual (GEO) (4), Array Express (5) and The Cancer Genome Atlas cancer type using the LIMMA statistical package in R, and (TCGA; http://cancergenome.nih.gov/). -
Primepcr™Assay Validation Report
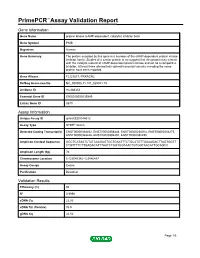
PrimePCR™Assay Validation Report Gene Information Gene Name protein kinase (cAMP-dependent, catalytic) inhibitor beta Gene Symbol PKIB Organism Human Gene Summary The protein encoded by this gene is a member of the cAMP-dependent protein kinase inhibitor family. Studies of a similar protein in rat suggest that this protein may interact with the catalytic subunit of cAMP-dependent protein kinase and act as a competitive inhibitor. At least three alternatively spliced transcript variants encoding the same protein have been reported. Gene Aliases FLJ23817, PRKACN2 RefSeq Accession No. NC_000006.11, NT_025741.15 UniGene ID Hs.486354 Ensembl Gene ID ENSG00000135549 Entrez Gene ID 5570 Assay Information Unique Assay ID qHsaCED0044612 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000368452, ENST00000368448, ENST00000258014, ENST00000354275, ENST00000368446, ENST00000392491, ENST00000392490 Amplicon Context Sequence GGCTCATAATCTATCAAGAGTGCTGAATTTCTGCATGTTGAAAGACTTAGTGGTT CTGTTTTCTTGAGACATTTAATCTGGTGGTAACTGTGGTAACATTGCAGCC Amplicon Length (bp) 76 Chromosome Location 6:123046342-123046447 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 95 R2 0.9996 cDNA Cq 22.83 cDNA Tm (Celsius) 76.5 gDNA Cq 24.52 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report PKIB, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve -

Location Analysis of Estrogen Receptor Target Promoters Reveals That
Location analysis of estrogen receptor ␣ target promoters reveals that FOXA1 defines a domain of the estrogen response Jose´ e Laganie` re*†, Genevie` ve Deblois*, Ce´ line Lefebvre*, Alain R. Bataille‡, Franc¸ois Robert‡, and Vincent Gigue` re*†§ *Molecular Oncology Group, Departments of Medicine and Oncology, McGill University Health Centre, Montreal, QC, Canada H3A 1A1; †Department of Biochemistry, McGill University, Montreal, QC, Canada H3G 1Y6; and ‡Laboratory of Chromatin and Genomic Expression, Institut de Recherches Cliniques de Montre´al, Montreal, QC, Canada H2W 1R7 Communicated by Ronald M. Evans, The Salk Institute for Biological Studies, La Jolla, CA, July 1, 2005 (received for review June 3, 2005) Nuclear receptors can activate diverse biological pathways within general absence of large scale functional data linking these putative a target cell in response to their cognate ligands, but how this binding sites with gene expression in specific cell types. compartmentalization is achieved at the level of gene regulation is Recently, chromatin immunoprecipitation (ChIP) has been used poorly understood. We used a genome-wide analysis of promoter in combination with promoter or genomic DNA microarrays to occupancy by the estrogen receptor ␣ (ER␣) in MCF-7 cells to identify loci recognized by transcription factors in a genome-wide investigate the molecular mechanisms underlying the action of manner in mammalian cells (20–24). This technology, termed 17-estradiol (E2) in controlling the growth of breast cancer cells. ChIP-on-chip or location analysis, can therefore be used to deter- We identified 153 promoters bound by ER␣ in the presence of E2. mine the global gene expression program that characterize the Motif-finding algorithms demonstrated that the estrogen re- action of a nuclear receptor in response to its natural ligand. -

Mouse Eaf2 Conditional Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Eaf2 Conditional Knockout Project (CRISPR/Cas9) Objective: To create a Eaf2 conditional knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Eaf2 gene (NCBI Reference Sequence: NM_001113401 ; Ensembl: ENSMUSG00000022838 ) is located on Mouse chromosome 16. 6 exons are identified, with the ATG start codon in exon 1 and the TGA stop codon in exon 6 (Transcript: ENSMUST00000114829). Exon 4 will be selected as conditional knockout region (cKO region). Deletion of this region should result in the loss of function of the Mouse Eaf2 gene. To engineer the targeting vector, homologous arms and cKO region will be generated by PCR using BAC clone RP24-400K10 as template. Cas9, gRNA and targeting vector will be co-injected into fertilized eggs for cKO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Mice homozygous for a null allele exhibit premature death, enlarged heart and prostate associate with hypertrophy, and increased incidence of tumors. Exon 4 starts from about 43.13% of the coding region. The knockout of Exon 4 will result in frameshift of the gene. The size of intron 3 for 5'-loxP site insertion: 2311 bp, and the size of intron 4 for 3'-loxP site insertion: 7170 bp. The size of effective cKO region: ~646 bp. The cKO region does not have any other known gene. Page 1 of 8 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele gRNA region 5' gRNA region 3' 1 4 6 Targeting vector Targeted allele Constitutive KO allele (After Cre recombination) Legends Exon of mouse Eaf2 Homology arm cKO region loxP site Page 2 of 8 https://www.alphaknockout.com Overview of the Dot Plot Window size: 10 bp Forward Reverse Complement Sequence 12 Note: The sequence of homologous arms and cKO region is aligned with itself to determine if there are tandem repeats. -

Genomic Variants Reveal Differential Evolutionary Constraints on Human Transglutaminases and Point Towards Unrecognized Significance of Transglutaminase 2
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by University of Debrecen Electronic Archive RESEARCH ARTICLE Genomic variants reveal differential evolutionary constraints on human transglutaminases and point towards unrecognized significance of transglutaminase 2 Kiruphagaran Thangaraju1, RoÂbert KiraÂly1, MaÂte A. DemeÂny1, JaÂnos AndraÂs MoÂtyaÂn1, a1111111111 Mo nika Fuxreiter1,2, LaÂszlo FeÂsuÈs1,3* a1111111111 a1111111111 1 Department of Biochemistry and Molecular Biology, Faculty of Medicine, University of Debrecen, a1111111111 Debrecen, Hungary, 2 MTA-DE Momentum Laboratory of Protein Dynamics, Faculty of Medicine, University a1111111111 of Debrecen, Debrecen, Hungary, 3 MTA-DE Stem cell, Apoptosis and Genomics Research Group of Hungarian Academy of Sciences, Faculty of Medicine, University of Debrecen, Debrecen, Hungary * [email protected] OPEN ACCESS Abstract Citation: Thangaraju K, KiraÂly R, DemeÂny MA, AndraÂs MoÂtyaÂn J, Fuxreiter M, FeÂsuÈs L (2017) Transglutaminases (TGMs) catalyze Ca2+-dependent transamidation of proteins with speci- Genomic variants reveal differential evolutionary constraints on human transglutaminases and point fied roles in blood clotting (F13a) and in cornification (TGM1, TGM3). The ubiquitous TGM2 towards unrecognized significance of has well described enzymatic and non-enzymatic functions but in-spite of numerous studies transglutaminase 2. PLoS ONE 12(3): e0172189. its physiological function in humans has not been defined. We compared data on non-syn- doi:10.1371/journal.pone.0172189 onymous single nucleotide variations (nsSNVs) and loss-of-function variants on TGM1-7 Editor: Richard L. Eckert, University of Maryland and F13a from the Exome aggregation consortium dataset, and used computational and School of Medicine, UNITED STATES biochemical analysis to reveal the roles of damaging nsSNVs of TGM2. -

Integrating Single-Step GWAS and Bipartite Networks Reconstruction Provides Novel Insights Into Yearling Weight and Carcass Traits in Hanwoo Beef Cattle
animals Article Integrating Single-Step GWAS and Bipartite Networks Reconstruction Provides Novel Insights into Yearling Weight and Carcass Traits in Hanwoo Beef Cattle Masoumeh Naserkheil 1 , Abolfazl Bahrami 1 , Deukhwan Lee 2,* and Hossein Mehrban 3 1 Department of Animal Science, University College of Agriculture and Natural Resources, University of Tehran, Karaj 77871-31587, Iran; [email protected] (M.N.); [email protected] (A.B.) 2 Department of Animal Life and Environment Sciences, Hankyong National University, Jungang-ro 327, Anseong-si, Gyeonggi-do 17579, Korea 3 Department of Animal Science, Shahrekord University, Shahrekord 88186-34141, Iran; [email protected] * Correspondence: [email protected]; Tel.: +82-31-670-5091 Received: 25 August 2020; Accepted: 6 October 2020; Published: 9 October 2020 Simple Summary: Hanwoo is an indigenous cattle breed in Korea and popular for meat production owing to its rapid growth and high-quality meat. Its yearling weight and carcass traits (backfat thickness, carcass weight, eye muscle area, and marbling score) are economically important for the selection of young and proven bulls. In recent decades, the advent of high throughput genotyping technologies has made it possible to perform genome-wide association studies (GWAS) for the detection of genomic regions associated with traits of economic interest in different species. In this study, we conducted a weighted single-step genome-wide association study which combines all genotypes, phenotypes and pedigree data in one step (ssGBLUP). It allows for the use of all SNPs simultaneously along with all phenotypes from genotyped and ungenotyped animals. Our results revealed 33 relevant genomic regions related to the traits of interest. -

Screening and Analysis of Pathogenic Genes During DMBA- Induced Buccal Mucosa Carcinogenesis in Golden Hamsters
1619-1624.qxd 23/4/2010 09:56 Ì ™ÂÏ›‰·1619 ONCOLOGY REPORTS 23: 1619-1624, 2010 Screening and analysis of pathogenic genes during DMBA- induced buccal mucosa carcinogenesis in golden hamsters KAI YANG1, GUODONG ZHANG1, JIE MEI1, DAN CHEN1 and MINGJUN WU2 1Department of Oral and Maxillofacial Surgery, the First Affiliated Hospital, Chongqing Medical University, Chongqing; 2Institute of Life Sciences of Chongqing Medical University, Chongqing 400016, P.R. China Received December 18, 2009; Accepted February 9, 2010 DOI: 10.3892/or_00000803 Abstract. We designed to screen pathogenic genes related to Introduction the occurrence and development of oral buccal mucosa cancer by whole genome microarray and analyze the mechanisms There are 274,000 new cases of oral cancer in the world every of carcinogenesis. The golden hamster model of buccal year (1). Squamous cell carcinoma accounts for 90% of mucosa cancer was established by induction with DMBA. oral cancer, and buccal carcinoma is one of the most common cRNAs labeled with Cy3 were synthesized and hybridized oral cancers (2). Although the treatment of oral cancers has with Agilent Whole Rat Genome Arrays containing 41,000 markedly improved in recent decades, the 5-year survival genes/ESTs. A Venn diagram analysis was performed to rate for buccal carcinoma and other oral cancers is only 55- screen the continuously abnormally expressed genes. Our 60% (2-4). Therefore, it is of great significance to explore results show 5,255 significantly differentially expressed the molecular mechanisms of the occurrence and develop- genes in golden hamster pouch mucosa during the ment of oral mucosa carcinomas and search for effective progression of normal buccal mucosa to squamous cell therapeutic targets. -

Role of Coiled-Coil Registry Shifts in Activation of Human Bicaudal D2 for Dynein Recruitment Upon Cargo-Binding
bioRxiv preprint doi: https://doi.org/10.1101/638536; this version posted May 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Role of coiled-coil registry shifts in activation of human Bicaudal D2 for dynein recruitment upon cargo-binding Crystal R. Noell,1,4 Jia Ying Loh, 1,4 Erik W. Debler,2 Kyle M. Loftus,1 Heying Cui,1 Blaine B. Russ,1 Puja Goyal,1,* Sozanne R. Solmaz1,3,* 1Department of Chemistry, State University of New York at Binghamton, Binghamton, NY 13902 2Department of Biochemistry & Molecular Biology, Thomas Jefferson University, Philadelphia, PA 19107. 3Lead contact. 4First author equally contributed *To whom the correspondence should be addressed: Sozanne R Solmaz Department of Chemistry, State University of New York at Binghamton PO Box 6000 Binghamton NY 13902 [email protected] +1 607 777 2089 Puja Goyal Department of Chemistry, State University of New York at Binghamton PO Box 6000 Binghamton NY 13902 [email protected] +1 607 777 4308 1 bioRxiv preprint doi: https://doi.org/10.1101/638536; this version posted May 15, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Highlights • Stable, bona fide BicD2 coiled-coils with distinct registries can be formed. • We provide evidence that a human disease mutation causes a coiled-coil registry shift. • A coiled-coil registry shift could relieve BicD2-autoinhibition upon cargo-binding. • The ability to undergo registry shifts may be an inherent property of coiled-coils. -

Co-Occurrence Based Meta-Analysis of Scientific Texts
Vol. 21 no. 9 2005, pages 2049–2058 BIOINFORMATICS ORIGINAL PAPER doi:10.1093/bioinformatics/bti268 Data and text mining Co-occurrence based meta-analysis of scientific texts: retrieving biological relationships between genes R. Jelier1,∗ G. Jenster2, L. C. J. Dorssers3, C. C. van der Eijk1, E. M. van Mulligen1, B. Mons1 and J. A. Kors1 1Department of Medical Informatics, 2Department of Urology and 3Department of Pathology, Erasmus MC—University Medical Center, Rotterdam, The Netherlands Received on July 15, 2004; revised on December 22, 2004; accepted on January 6, 2005 Advance Access publication January 18, 2005 ABSTRACT are studied, the number of relevant publications will frequently be Motivation: The advent of high-throughput experiments in molecu- prohibitively large. This renders the traditional approach of manu- lar biology creates a need for methods to efficiently extract and use ally searching bibliographic databases for every gene and reading information for large numbers of genes. Recently, the associative scientific articles inadequate. It is therefore an important challenge concept space (ACS) has been developed for the representation of at this time to make the available information both accessible and information extracted from biomedical literature. The ACS is a Euc- interpretable for molecular biologists. lidean space in which thesaurus concepts are positioned and the An interesting current development is the use of annotations of distances between concepts indicates their relatedness. The ACS genes with gene ontology (GO) terms (Ashburner et al., 2000; Camon uses co-occurrence of concepts as a source of information. In this et al., 2004) for the analysis of the results of microarray experiments paper we evaluate how well the system can retrieve functionally (Zhang et al., 2004; Al Shahrour et al., 2004). -

Molecular Effects of Isoflavone Supplementation Human Intervention Studies and Quantitative Models for Risk Assessment
Molecular effects of isoflavone supplementation Human intervention studies and quantitative models for risk assessment Vera van der Velpen Thesis committee Promotors Prof. Dr Pieter van ‘t Veer Professor of Nutritional Epidemiology Wageningen University Prof. Dr Evert G. Schouten Emeritus Professor of Epidemiology and Prevention Wageningen University Co-promotors Dr Anouk Geelen Assistant professor, Division of Human Nutrition Wageningen University Dr Lydia A. Afman Assistant professor, Division of Human Nutrition Wageningen University Other members Prof. Dr Jaap Keijer, Wageningen University Dr Hubert P.J.M. Noteborn, Netherlands Food en Consumer Product Safety Authority Prof. Dr Yvonne T. van der Schouw, UMC Utrecht Dr Wendy L. Hall, King’s College London This research was conducted under the auspices of the Graduate School VLAG (Advanced studies in Food Technology, Agrobiotechnology, Nutrition and Health Sciences). Molecular effects of isoflavone supplementation Human intervention studies and quantitative models for risk assessment Vera van der Velpen Thesis submitted in fulfilment of the requirements for the degree of doctor at Wageningen University by the authority of the Rector Magnificus Prof. Dr M.J. Kropff, in the presence of the Thesis Committee appointed by the Academic Board to be defended in public on Friday 20 June 2014 at 13.30 p.m. in the Aula. Vera van der Velpen Molecular effects of isoflavone supplementation: Human intervention studies and quantitative models for risk assessment 154 pages PhD thesis, Wageningen University, Wageningen, NL (2014) With references, with summaries in Dutch and English ISBN: 978-94-6173-952-0 ABSTRact Background: Risk assessment can potentially be improved by closely linked experiments in the disciplines of epidemiology and toxicology. -

Null Mutation of the Fascin2 Gene by TALEN Leading to Progressive Hearing Loss and Retinal Degeneration in C57BL/6J Mice
G3: Genes|Genomes|Genetics Early Online, published on August 6, 2018 as doi:10.1534/g3.118.200405 Null mutation of the Fascin2 gene by TALEN leading to progressive hearing loss and retinal degeneration in C57BL/6J mice Xiang Liu1,3,§, Mengmeng Zhao1,2,§, Yi Xie1,2, §, Ping Li1, Oumei Wang1 , Bingxin Zhou1,2, Linlin Yang1,4, Yao Nie1, Lin Cheng1, Xicheng Song1,5, Changzhu Jin1,3,**, Fengchan Han1,2,* 1Key Laboratory for Genetic Hearing Disorders in Shandong, Binzhou Medical University, 346 Guanhai Road, Yantai264003, Shandong, P. R. China 2Department of Biochemistry and Molecular Biology, Binzhou Medical University, 346 Guanhai Road, Yantai264003, Shandong, P. R. China 3Department of Human Anatomy and Histology and Embryology, Binzhou Medical University, 346 Guanhai Road, Yantai264003, Shandong, P. R. China 4Department of Otorhinolaryngology-Head and Neck Surgery, Yuhuangding Hospital, 20 East Yuhuangding Road, Yantai 264000, Shandong, P.R .China § These authors contribute equally *The First Corresponding Author: [email protected] **The Second Corresponding Author: [email protected] Key Laboratory for Genetic Hearing Disorders in Shandong, Department of Biochemistry and Molecular Biology, Binzhou Medical University, 346 Guanhai Road, Yantai264003, Shandong, P. R. China © The Author(s) 2013. Published by the Genetics Society of America. Abstract Fascin2 (FSCN2) is an actin cross-linking protein that is mainly localized in retinas and in the stereocilia of hair cells. Earlier studies showed that a deletion mutation in human FASCIN2 (FSCN2) gene could cause autosomal dominant retinitis pigmentosa. Recent studies have indicated that a missense mutation in mouse Fscn2 gene (R109H) can contribute to the early onset of hearing loss in DBA/2J mice.