Assignment 1 Part I
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Role of NF-Κb and C/EBP Factors During Pathogen-Mediated
The role of NF-B and C/EBP factors during pathogen-mediated activation of bovine interleukin 8 and beta-defensin in mammary epithelial cells Inaugural dissertation for the academic degree Doctor rerum naturalium of Mathematisch-Naturwissenschaftlichen Fakultät Universität Rostock By Shuzhen Liu (M. Sc.), born on March-02-1972, in Shanxi Province, China From the Forschungsinstitut für die Biologie landwirtschaftlicher Nutztiere in Dummerstorf Rostock 2009 URN: urn:nbn:de:gbv:28-diss2009-0185-5 Dean: Prof. Dr. Hendrik Schubert Reviewers: 1. Prof. Dr. Hans-Martin Seyfert Research Unit molecular biology, Research Institute for the Biology of Farm Animals, Wilhelm-Stahl-Allee 2, D-18196 Dummerstorf, Germany 2. Prof. Dr. Dieter G. Weiss Division of animal Physiology, Institute of Cell Biology and Biosystems Technology, University of Rostock, Albert-Einstein-Strasse 3, 18059 Rostock, Germany 3. PD Dr. Ulrike Gimsa Research Unit Behavioural Physiology, Research Institute for the Biology of Farm Animals, Wilhelm-Stahl-Allee 2, D-18196 Dummerstorf, Germany Date of defense: October 19th, 2009 Table of Contents TABLE OF CONTENTS 1. INTRODUCTION..................................................................................................................1 1.1 Mastitis as a challenge in general immunology..................................................................1 1.2 Innate immunity of the bovine mammary gland ................................................................2 1.3 Toll-like receptors (TLRs): main receptors perceiving the pathogen -

Genetic Analysis of SARS-Cov-2 and the Common Golden Nucleotides to Human Gene
Genetic analysis of SARS-CoV-2 and the common golden nucleotides to human gene Hamed Babaee ( [email protected] ) Payame Noor University https://orcid.org/0000-0001-7093-6400 Research Article Keywords: Coronavirus, COVID-19, SARS-CoV-2, RNA sequencing, Human genome Posted Date: August 30th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-854956/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/11 Abstract Background The coronavirus disease pandemic began in 2019 in Wuhan, China, and continues into 2021, as new mutant viruses appear and require new solutions and treatments. Methods and Results In this study, by genetic analysis of data from 24 SARS-CoV-2 samples from different countries and their alignment with each other and the Wuhan reference virus as well as the human genome, the disease factor is looked at from another angle. The result is the identication of genetic differences in viruses, and the nding of a unique 17-nucleotide sequence between human genes, viruses, and enzymes that can contribute to the onset and progression of the disease. Conclusions The role of this sequence in DNA replication and the production of new proteins and its alignment with the EPPK1 gene that may cause disease and its various symptoms are likely. Introduction In 2019, the world was again struck by a new viral contagious disease called COVID-19 or Coronavirus disease. Apparently, this disease originated from a seafood wholesale market in Wuhan, China from an unknown animal source. In the beginning, China had the highest case report and mortality rates which were mitigated by severe quarantine measures. -

CLONING, EXPRESSION ANALYSIS, and TRANSFORMATION VECTOR CONSTRUCTION of DAM HOMOLOGS in PEACH and POPLAR Yuhui Xie Clemson University, [email protected]
Clemson University TigerPrints All Theses Theses 1-2011 CLONING, EXPRESSION ANALYSIS, AND TRANSFORMATION VECTOR CONSTRUCTION OF DAM HOMOLOGS IN PEACH AND POPLAR Yuhui Xie Clemson University, [email protected] Follow this and additional works at: https://tigerprints.clemson.edu/all_theses Part of the Plant Biology Commons Recommended Citation Xie, Yuhui, "CLONING, EXPRESSION ANALYSIS, AND TRANSFORMATION VECTOR CONSTRUCTION OF DAM HOMOLOGS IN PEACH AND POPLAR" (2011). All Theses. 1146. https://tigerprints.clemson.edu/all_theses/1146 This Thesis is brought to you for free and open access by the Theses at TigerPrints. It has been accepted for inclusion in All Theses by an authorized administrator of TigerPrints. For more information, please contact [email protected]. CLONING, EXPRESSION ANALYSIS, AND TRANSFORMATION VECTOR CONSTRUCTION OF DAM HOMOLOGS IN PEACH AND POPLAR _______________________________________________ A Thesis Presented to the Graduate School of Clemson University ________________________________________________ In Partial Fulfillment of the Requirement for the Degree Master of Science Plant and Environmental Sciences _________________________________________________ by Yuhui Xie August 2011 ________________________________________________ Accepted by: Dr. Douglas G. Bielenberg, Committee Chair Dr. Haiying Liang Dr. Hong Luo ABSTRACT Genetic fine mapping and sequencing of the EVG locus in peach [Prunus persica (L.) Batsch] identified six tandem arrayed Dormancy-Associated MADS-box (DAM) genes as candidates for regulating growth cessation and terminal bud formation in the non-dormant evergrowing (evg) mutant. Since the mutant is lacking expression of six genes in the mapped locus, further functional analysis is needed to narrow the list of gene candidates for the non-dormant evg phenotype. Here I report three sets of experiments designed to functionally test DAM genes in peach and their homologs in a model tree, hybrid poplar. -

Enzymes in Cloning Part I
˹̀/˺̀/˺̊˼̊ ENZYMES IN CLONING PART I Dr.Sarookhani ˺ ˹̀/˺̀/˺̊˼̊ Cloning --aa definition •• From the Greek --klon,klon, a twig •• An aggregate of the asexually produced progeny of an individual;a group of replicas of all or part of a macromolecule (such as DNA or an antibody) •• An individual grown from a single somatic cell of its parent & genetically identical to it •• Clone: a collection of molecules or cells, all identical to an original molecule or cell Dr.Sarookhani ˻ ˹̀/˺̀/˺̊˼̊ Different types of Cloning 1. Reproductive Cloning 2. Therapeutic Cloning 3. Recombinant DNA Technology or DNA Cloning Dr.Sarookhani ˼ ˹̀/˺̀/˺̊˼̊ DNA CLONING A method for identifying and purifying a particular DNA fragment (clone) of interest from a complex mixture of DNA fragments, and then producing large numbers of the fragment (clone) of interest. Dr.Sarookhani ̊ ˹̀/˺̀/˺̊˼̊ What is genetic engineering • Genetic engineering, also known as recombinant DNA technology, means altering the genes in a living organism to produce a Genetically Modified Organism (GMO) with a new genotype. • Various kinds of genetic modification are possible: inserting a foreign gene from one species into another, forming a transgenic organism; altering an existing gene so that its product is changed; or changing gene expression so that it is translated more often or not at all. Dr.Sarookhani ̋ ˹̀/˺̀/˺̊˼̊ Dr.Sarookhani ̌ ˹̀/˺̀/˺̊˼̊ Dr.Sarookhani ̀ ˹̀/˺̀/˺̊˼̊ Genomic Library Dr.Sarookhani ́ ˹̀/˺̀/˺̊˼̊ Dr.Sarookhani ̂ ˹̀/˺̀/˺̊˼̊ Basic steps in genetic engineering 1. -

Cyclic Digestion and Ligation-Mediated PCR Used For
www.nature.com/scientificreports OPEN Cyclic Digestion and Ligation- Mediated PCR Used for Flanking Sequence Walking Dong Yu1,2,5, Tianshun Zhou2,4,5, Xuewu Sun2,3, Zhizhong Sun2, Xiabing Sheng1,2, Yanning Tan2, Ling Liu2,4, Ning Ouyang2,4, Ke Xu2, Kaibing Shi2, Guilong Yuan2, Jia Ding2, Meijuan Duan3* & Dingyang Yuan1,2,3,4* Ligation-mediated PCR (LM-PCR) is a classical method for isolating fanking sequences; however, it has a common limitation of reduced success rate owing to the circularization or multimerization of target restriction fragments including the known sequence. To address this limitation, we developed a novel LM-PCR method, termed Cyclic Digestion and Ligation-Mediated PCR (CDL-PCR). The novelty of this approach involves the design of new adapters that cannot be digested after being ligated with the restriction fragment, and cyclic digestion and ligation may be manipulated to block the circularization or multimerization of the target restriction fragments. Moreover, to improve the generality and fexibility of CDL-PCR, an adapter precursor sequence was designed, which could be digested to prepare 12 diferent adapters at low cost. Using this method, the fanking sequences of T-DNA insertions were obtained from transgenic rice and Arabidopsis thaliana. The experimental results demonstrated that CDL-PCR is an efcient and fexible method for identifying the fanking sequences in transgenic rice and Arabidopsis thaliana. Identifcation of fanking sequences has ofen been employed to determine the location of T-DNA insertion in genomic DNA. Methods to obtain fanking sequencea include inverse PCR1, randomly primed PCR2–5, and ligation-mediated PCR6–8. Inverse PCR, the earliest fanking cloning technique, has a low efciency and is limited by the rate of self-ligation and amplifcation range of DNA polymerases9. -

The Cloning Guide the First Step Towards a Succesful Igem Project Igem Bonn NRP-UEA Igem
The Cloning Guide The first step towards a succesful iGEM Project iGEM Bonn NRP-UEA iGEM iGEM Manchester-Graz iGEM Evry iGEM Vanderbilt iGEM Minnesota Carnegie Mellon iGEM iGEM Sydney UIUC iGEM iGEM York iGEM Toulouse iGEM Pasteur UCLA iGEM iGEM Stockholm Preface The cloning guide which is lying before you or on your desktop is a document brought to you by iGEM TU Eindhoven in collaboration with numerous iGEM (International Genetically Engi- neered Machine) teams during iGEM 2015. An important part in the iGEM competition is the collabration of your own team with other teams. These collaborations are fully in line with the dedications of the iGEM Foundation on education and competition, the advancement of syn- thetic biology, and the development of an open community and collaboration. Collaborations between groups of (undergrad and overgrad) students can lead to nice products, as we have tried to provide one for you in 2015. In order to compile a cloning guide, several iGEM teams from all over the world have been con- tacted to cooperate on this. Finally fifiteen teams contributed in a fantastic way and a cloning guide consisting of the basics about nine different cloning methods and experiences of teams working with them has been realized. Without the help of all collaborating teams this guide could never have been realized, so we will thank all teams in advance. This guide may be of great help when new iGEM teams (edition 2016 and later) are at the point of designing their project. How to assemble the construct for you project, is an important choice which is possibly somewhat easier to make after reading this guide. -

Psp73 Vector Technical Bulletin #TB041
tb041.0906.qxp 9/25/2006 10:44 AM Page a Technical Bulletin pSP73 Vector INSTRUCTIONS FOR USE OF PRODUCT P2221. PRINTED IN USA. Revised 9/06 Part# TB041 AF9TB041 0906TB041 tb041.0906.qxp 9/25/2006 10:46 AM Page 1 pSP73 Vector All technical literature is available on the Internet at: www.promega.com/tbs/ Please visit the web site to verify that you are using the most current version of this Technical Bulletin. Please contact Promega Technical Services if you have questions on use of this system. E-mail: [email protected] I. Description..........................................................................................................1 II. Product Components and Storage Conditions ............................................1 III. pSP73 Vector Multiple Cloning Region and Circle Map..........................2 IV. pSP73 Vector Restriction Sites........................................................................4 V. Related Products ................................................................................................6 VI. Reference .............................................................................................................7 I. Description The pSP73 Vector (1) offers a wide range of restriction sites, providing greater versatility in cloning and transcription of RNA in vitro. The pSP73 Vector contains the SP6 and T7 RNA polymerase promoters and a unique multiple cloning region, which includes restriction sites for BglII, EcoRV, ClaI, EcoRI, SacI, KpnI, SmaI, BamHI, XbaI, AccI, SalI, PstI, SphI, HindIII, PvuII and XhoI. The sequences of Promega vectors are available online at www.promega.com/vectors/ and are also available from the GenBank® database. II. Product Components and Storage Conditions Product Size Cat.# pSP73 Vector 20µg P2221 Storage Conditions: Store the pSP73 Vector at –20°C. Promega Corporation · 2800 Woods Hollow Road · Madison, WI 53711-5399 USA Toll Free in USA 800-356-9526 · Phone 608-274-4330 · Fax 608-277-2516 · www.promega.com Printed in USA. -

Bpuami: a Novel Saci Neoschizomer from Bacillus Pumilus Discovered in an Isolate from Amazon Basin, Recognizing 5'-Gag↓Ctc-3'
Brazilian Journal of Microbiology (2006) 37:96-100 ISSN 1517-8382 BPUAMI: A NOVEL SACI NEOSCHIZOMER FROM BACILLUS PUMILUS DISCOVERED IN AN ISOLATE FROM AMAZON BASIN, RECOGNIZING 5'-GAG↓CTC-3' Jocelei M. Chies1,2,*; Ana C. de O. Dias1; Hélio M. M. Maia3; Spartaco Astolfi-Filho2,3 1Centro de Biotecnologia, Universidade Federal do Rio Grande do Sul, RS, Brasil; 2Curso de Pós-Graduação em Biologia Molecular, Universidade de Brasília, Brasília, DF, Brasil; 3Centro de Apoio Multidisciplinar, Universidade Federal do Amazonas, Manaus, AM, Brasil Submitted: June 27, 2005; Returned to authors for corrections: November 16, 2005; Approved: January 12, 2006 ABSTRACT A strain of Bacillus pumilus was isolated and identified from water samples collected from a small affluent of the Amazon River. Type II restriction endonuclease activity was detected in these bacteria. The enzyme was purified and the molecular weight of the native protein estimated by gel filtration and SDS-PAGE. The optimum pH, temperature and salt requirements were determined. Quality control assays showed the complete absence of “nonspecific nucleases.” Restriction cleavage analysis and DNA sequencing of restriction fragments allowed the unequivocal demonstration of 5´GAG↓CTC3´ as the recognition sequence. This enzyme was named BpuAmI and is apparently a neoschizomer of the prototype restriction endonuclease SacI. This is the first report of an isoschizomer and/or neoschizomer of the prototype SacI identified in the genus Bacillus. Key words: type II restriction endonuclease, BpuAmI, SacI, neoschizomers, Bacillus pumilus INTRODUCTION thousands of taxonomically diverse bacteria for enzymes with new characteristics. However, to our knowledge, there has Restriction endonucleases are enzymes which recognize not been any report to date of an isoschizomer and/or short DNA sequences and cleave DNA in both strands. -

Psp72 Vector INSTRUCTIONS for USE of PRODUCT P2191
tb040.0906.qxp 9/25/2006 10:12 AM Page a Technical Bulletin pSP72 Vector INSTRUCTIONS FOR USE OF PRODUCT P2191. PRINTED IN USA. Revised 9/06 Part# TB040 AF9TB040 0906TB040 tb040.0906.qxp 9/25/2006 10:12 AM Page 1 pSP72 Vector All technical literature is available on the Internet at: www.promega.com/tbs/ Please visit the web site to verify that you are using the most current version of this Technical Bulletin. Please contact Promega Technical Services if you have questions on use of this system. E-mail: [email protected] I. Description..........................................................................................................1 II. Product Components and Storage Conditions ............................................1 III. pSP72 Vector Multiple Cloning Region and Circle Map..........................2 IV. pSP72 Vector Restriction Sites........................................................................4 V. Related Products ................................................................................................6 VI. Reference .............................................................................................................6 I. Description The pSP72 Vector (1) can be used as a standard cloning vector and can also be used for transcription of RNA in vitro. The pSP72 Vector contains the SP6 and T7 RNA polymerase promoters flanking a unique multiple cloning region, which includes restriction sites for XhoI, PvuII, HindIII, SphI, PstI, SalI, AccI, XbaI, BamHI, SmaI, KpnI, SacI, EcoRI, ClaI, EcoRV and BglII. The sequences of Promega vectors are available online at: www.promega.com/vectors/ and are also available from the GenBank® database. II. Product Components and Storage Conditions Product Size Cat.# pSP72 Vector 20µg P2191 Storage Conditions: Store the pSP72 Vector at –20°C. Promega Corporation · 2800 Woods Hollow Road · Madison, WI 53711-5399 USA Toll Free in USA 800-356-9526 · Phone 608-274-4330 · Fax 608-277-2516 · www.promega.com Printed in USA. -

Restriction Endonucleases
Restriction Endonucleases TECHNICAL GUIDE UPDATE 2017/18 be INSPIRED drive DISCOVERY stay GENUINE RESTRICTION ENZYMES FROM NEB Cut Smarter with Restriction Enzymes from NEB® Looking to bring CONVENIENCE to your workflow? Simplify Reaction Setup and Double Activity of DNA Modifying Enzymes in CutSmart Buffer: Digestion with CutSmart® Buffer Clone Smarter! Activity Enzyme Required Supplements Over 210 restriction enzymes are 100% active in a single buffer, in CutSmart Phosphatases: CutSmart Buffer, making it significantly easier to set up your Alkaline Phosphatase (CIP) + + + double digest reactions. Since CutSmart Buffer includes BSA, there Antarctic Phosphatase + + + Requires Zn2+ Quick CIP + + + are fewer tubes and pipetting steps to worry about. Additionally, Shrimp Alkaline Phosphatase (rSAP) + + + many DNA modifying enzymes are 100% active in CutSmart Ligases: T4 DNA Ligase + + + Requires ATP Buffer, eliminating the need for subsequent purification. E. coli DNA Ligase + + + Requires NAD T3 DNA Ligase + + + Requires ATP + PEG For more information, visit www.NEBCutSmart.com T7 DNA Ligase + + + Requires ATP + PEG Polymerases: T4 DNA Polymerase + + + DNA Polymerase I, Large (Klenow) Frag. + + + DNA Polymerase I + + + DNA Polymerase Klenow Exo– + + + Bst DNA Polymerase + + + ™ phi29 DNA Polymerase + + + Speed up Digestions with Time-Saver T7 DNA Polymerase (unmodified) + + + Qualified Restriction Enzymes Transferases/Kinases: T4 Polynucleotide Kinase + + + Requires ATP + DTT T4 PNK (3´ phosphatase minus) + + + Requires ATP + DTT > 190 of our restriction enzymes are able to digest DNA in CpG Methyltransferase (M. SssI) + + + 5–15 minutes, and can safely be used overnight with no loss of GpC Methyltransferase (M. CviPI) + Requires DTT T4 Phage β-glucosyltransferase + + + sample. For added convenience and flexibility, most of these are Nucleases, other: supplied with CutSmart Buffer. -
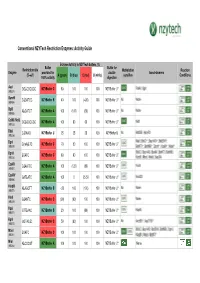
Conventional Nzytech Restriction Enzymes: Activity Guide
Conventional NZYTech Restriction Enzymes: Activity Guide Enzyme Activity in NZYTech buffers (%) Buffer Buffer for Restriction site Methylation Reaction Enzyme provided for double Isoschizomers (5’ →→→3’) A (green) B (blue) C (red) U (white) sensitive Conditions 100% activity digestion AscI GG ↓↓↓CGCGCC NZYBuffer C 80 100 100 100 NZYBuffer U * PalAI, SgsI (MB231) BamHI G↓↓↓GATCC NZYBuffer B 40 100 (<20) 100 NZYBuffer U * No None (MB064) BglII A↓↓↓GATCT NZYBuffer A 100 (100) (60) 100 NZYBuffer U * No None (MB065) CciNI (NotI) GC ↓↓↓GGCCGC NZYBuffer A 100 80 60 100 NZYBuffer U * NotI (MB153) DdeI C↓↓↓TNAG NZYBuffer U 25 25 25 100 NZYBuffer U No BstDEI, HpyF3I (MB236) MalI, BfuCI ♦♦♦, Bsp143I ♦♦♦, BstENII ♦♦♦, DpnI G(mA)↓↓↓TC NZYBuffer C 70 50 100 100 NZYBuffer U * (MB078) BstKTI ♦♦♦, BstMBI ♦♦♦, DpnII ♦♦♦, Kzo9I ♦♦♦, NdeII ♦♦♦ BfuCI, Bsp143I, BssMI, BstKTI, BstMBI, DpnII ↓↓↓GATC NZYBuffer C 80 80 100 100 NZYBuffer U * (MB233) Kzo9I, MboI, NdeII, Sau3AI EcoRI G↓↓↓AATTC NZYBuffer A 100 (120) (80) 100 NZYBuffer U * FunII (MB067) EcoRV GAT ↓↓↓ATC NZYBuffer A 100 0 25-50 100 NZYBuffer U * Eco32I (MB068) HindIII A↓↓↓AGCTT NZYBuffer B <20 100 (100) 100 NZYBuffer U * No None (MB070) HinfI G↓↓↓ANTC NZYBuffer C (90) (90) 100 100 NZYBuffer U * None (MB239) HpaI GTT ↓↓↓AAC NZYBuffer B 20 100 (80) 100 NZYBuffer U * KspAI (MB071) KpnI GGTAC ↓↓↓C NZYBuffer C 50 (80) 100 100 NZYBuffer U * No Acc65I ♦♦♦, Asp718I ♦♦♦ (MB072) BfuCI, Bsp143I, BstENII, BstMBI, DpnII, MboI ↓↓↓GATC NZYBuffer C 100 100 100 100 NZYBuffer U * (MB241) Kzo9II, NdeII, BstKTI ♦♦♦ MluI -

James Alan Tunaley
Structure and Activity Investigations of the Cell Fate Determinant, SpoIIE, from Bacillus subtilis. James Alan Tunaley Thesis Submitted for the Degree of Doctor of Philosophy University of York Department of Chemistry September 2013 Abstract For many years the Gram positive bacterium Bacillus subtilis has been a model organism for prokaryotic cell and molecular biology. The asymmetric cell division which B. subtilis undergoes during sporulation is a simple system by which to study the process of cell differentiation. Sporulation is governed by a series of genetic temporal and spatial controls. Gene regulation brought about by a series of σ factors and transcriptional regulators is coupled to key morphological stages or checkpoints. σF initiates the first step in a cascade of complex genetic control which eventually produces a resilient endospore. The activation of σF, the first compartment-specific sigma factor, in the forespore and its regulation through interaction between three proteins; SpoIIAA, SpoIIAB and SpoIIE, is of particular interest. SpoIIE, a protein phosphatase which binds to the asymmetric division septum, is a crucial factor in the selective activation of σF in the forespore. Of three putative domains in SpoIIE only the C-terminal PP2C phosphatase domain has been structurally characterised. The central domain, domain II, of SpoIIE has been assigned a role in interaction with the cell division machinery; however mutational studies have shown that, in addition, this domain is also responsible for the regulation of phosphatase activity. This work describes the isolation and characterisation of three new fragments of SpoIIE containing elements of the central cytoplasmic domain of SpoIIE. These include a fragment found to accurately represent the N-terminal solubility limit of domain II which shows a high degree of oligomeric character.