PPM1H Phosphatase Counteracts LRRK2 Signaling by Selectively
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Myotubularin-Related Protein (MTMR) 9 Determines the Enzymatic Activity, Substrate Specificity, and Role in Autophagy of MTMR8
Myotubularin-related protein (MTMR) 9 determines the enzymatic activity, substrate specificity, and role in autophagy of MTMR8 Jun Zoua,1, Chunfen Zhangb,1,2, Jasna Marjanovicc, Marina V. Kisselevab, Philip W. Majerusb,d,2, and Monita P. Wilsonb,2 aDepartment of Pathology and Immunology, bDivision of Hematology, Department of Internal Medicine, and dDepartment of Biochemistry and Molecular Biophysics, Washington University School of Medicine, St. Louis, MO 63110; and cDivision of Basic and Pharmaceutical Sciences, St. Louis College of Pharmacy, St. Louis, MO 63110 Contributed by Philip W. Majerus, May 1, 2012 (sent for review February 24, 2012) The myotubularins are a large family of inositol polyphosphate myotubularin proteins (16–21). One mechanism that regulates 3-phosphatases that, despite having common substrates, subsume the myotubularins is the formation of heterodimers between unique functions in cells that are disparate. The myotubularin catalytically active and inactive proteins. The interaction between family consists of 16 different proteins, 9 members of which different myotubularin proteins has a significant effect on en- possess catalytic activity, dephosphorylating phosphatidylinositol zymatic activity. For example, the association of myotubularin 3-phosphate [PtdIns(3)P] and phosphatidylinositol 3,5-bisphos- (MTM1) with MTMR12 results in a threefold increase in the 3- phate [PtdIns(3,5)P2] at the D-3 position. Seven members are in- phosphatase activity of MTM1, alters the subcellular localiza- active because they lack the conserved cysteine residue in the tion of MTM1 from the plasma membrane to the cytosol, and CX5R motif required for activity. We studied a subfamily of homol- attenuates the filopodia formation seen with MTM1 overex- ogous myotubularins, including myotubularin-related protein 6 pression (21, 22). -

Myotubularin-Related Phosphatase 5 Is a Critical Determinant of Autophagy in Neurons
bioRxiv preprint doi: https://doi.org/10.1101/2021.07.20.453106; this version posted July 20, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Myotubularin-related phosphatase 5 is a critical determinant of autophagy in neurons Jason P. Chua*1,8, Karan Bedi2,3,4, Michelle T. Paulsen2,4, Mats Ljungman2,4, Elizabeth M. H. Tank1, Erin S. Kim1, Jennifer M. Colón-Mercado7, Michael E. Ward7, Lois S. Weisman5,6, and Sami J. Barmada*1,8 1Department of Neurology, University of Michigan, Ann Arbor, MI, USA 2Department of Radiation Oncology, University of Michigan, Ann Arbor, MI, USA 3Department of Biostatistics, University of Michigan, Ann Arbor, MI, USA 4Rogel Cancer Center and Center for RNA Biomedicine, University of Michigan, Ann Arbor, MI, USA 5Life Sciences Institute, University of Michigan, Ann Arbor, MI, USA 6Department of Cell and Developmental Biology, University of Michigan, Ann Arbor, MI, USA 7National Institute of Neurological Disorders and Stroke, NIH, Bethesda, MD, USA 8Lead contact *Correspondence: [email protected] (J.P.C.), [email protected] (S.J.B.) 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.07.20.453106; this version posted July 20, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. ABSTRACT Autophagy is a conserved, multi-step process of capturing proteolytic cargo in autophagosomes for lysosome degradation. The capacity to remove toxic proteins that accumulate in neurodegenerative disorders attests to the disease-modifying potential of the autophagy pathway. -

Supplemental Figure 1. Vimentin
Double mutant specific genes Transcript gene_assignment Gene Symbol RefSeq FDR Fold- FDR Fold- FDR Fold- ID (single vs. Change (double Change (double Change wt) (single vs. wt) (double vs. single) (double vs. wt) vs. wt) vs. single) 10485013 BC085239 // 1110051M20Rik // RIKEN cDNA 1110051M20 gene // 2 E1 // 228356 /// NM 1110051M20Ri BC085239 0.164013 -1.38517 0.0345128 -2.24228 0.154535 -1.61877 k 10358717 NM_197990 // 1700025G04Rik // RIKEN cDNA 1700025G04 gene // 1 G2 // 69399 /// BC 1700025G04Rik NM_197990 0.142593 -1.37878 0.0212926 -3.13385 0.093068 -2.27291 10358713 NM_197990 // 1700025G04Rik // RIKEN cDNA 1700025G04 gene // 1 G2 // 69399 1700025G04Rik NM_197990 0.0655213 -1.71563 0.0222468 -2.32498 0.166843 -1.35517 10481312 NM_027283 // 1700026L06Rik // RIKEN cDNA 1700026L06 gene // 2 A3 // 69987 /// EN 1700026L06Rik NM_027283 0.0503754 -1.46385 0.0140999 -2.19537 0.0825609 -1.49972 10351465 BC150846 // 1700084C01Rik // RIKEN cDNA 1700084C01 gene // 1 H3 // 78465 /// NM_ 1700084C01Rik BC150846 0.107391 -1.5916 0.0385418 -2.05801 0.295457 -1.29305 10569654 AK007416 // 1810010D01Rik // RIKEN cDNA 1810010D01 gene // 7 F5 // 381935 /// XR 1810010D01Rik AK007416 0.145576 1.69432 0.0476957 2.51662 0.288571 1.48533 10508883 NM_001083916 // 1810019J16Rik // RIKEN cDNA 1810019J16 gene // 4 D2.3 // 69073 / 1810019J16Rik NM_001083916 0.0533206 1.57139 0.0145433 2.56417 0.0836674 1.63179 10585282 ENSMUST00000050829 // 2010007H06Rik // RIKEN cDNA 2010007H06 gene // --- // 6984 2010007H06Rik ENSMUST00000050829 0.129914 -1.71998 0.0434862 -2.51672 -

Genes Retina/RPE Choroid Sclera
Supplementary Materials: Genes Retina/RPE Choroid Sclera Fold Change p-value Fold Change p-value Fold Change p-value PPFIA2 NS NS 2.35 1.3X10-3 1.5 1.6X10-3 PTPRF 1.24 2.65X10-5 6.42 7X10-4 1.11 1X10-4 1.19 2.65X10-5 NS NS 1.11 3.3X10-3 PTPRR 1.44 2.65X10-5 3.04 4.7X10-3 NS NS Supplementary Table S1. Genes Differentially Expressed Related to Candidate Genes from Association. Genes selected for follow up validation by real time quantitative PCR. Multiple values for each gene indicate multiple probes within the same gene. NS indicates the fold change was not statistically significant. Gene/SNP Assay ID rs4764971 C__30866249_10 rs7134216 C__30023434_10 rs17306116 C__33218892_10 rs3803036 C__25749934_20 rs824311 C___8342112_10 PPFIA2 Hs00170308_m1 PTPRF Hs00160858_m1 PTPRR Hs00373136_m1 18S Hs03003631_g1 GAPDH Hs02758991_g1 Supplementary Table S2. Taqman® Genotyping and Gene Expression Assay Identification Numbers. SNP Chimp Orangutan Rhesus Marmoset Mouse Rat Cow Pig Guinea Pig Dog Elephant Opossum Chicken rs3803036 X X X X X X X X X X X X X rs1520562 X X X X X X rs1358228 X X X X X X X X X X X rs17306116 X X X X X X rs790436 X X X X X X X rs1558726 X X X X X X X X rs741525 X X X X X X X X rs7134216 X X X X X X rs4764971 X X X X X X X Supplementary Table S3. Conservation of Top SNPs from Association. X indicates SNP is conserved. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Molecular and Genetic Medicine
Bertazzi et al., J Mol Genet Med 2015, 8:2 Molecular and Genetic Medicine http://dx.doi.org/10.4172/1747-0862.1000116 Review Article Open Access Myotubularin MTM1 Involved in Centronuclear Myopathy and its Roles in Human and Yeast Cells Dimitri L. Bertazzi#, Johan-Owen De Craene# and Sylvie Friant* Department of Molecular and Cellular Genetics, UMR7156, Université de Strasbourg and CNRS, France #Authors contributed equally to this work. *Corresponding author: Friant S, Department of Molecular and Cellular Genetics, UMR7156, Université de Strasbourg and CNRS, 67084 Strasbourg, France, E-mail: [email protected] Received date: April 17, 2014; Accepted date: July 21, 2014; Published date: July 28, 2014 Copyright: © 2014 Bertazzi DL, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. Abstract Mutations in the MTM1 gene, encoding the phosphoinositide phosphatase myotubularin, are responsible for the X-linked centronuclear myopathy (XLCNM) or X-linked myotubular myopathy (XLMTM). The MTM1 gene was first identified in 1996 and its function as a PtdIns3P and PtdIns(,5)P2 phosphatase was discovered in 2000. In recent years, very important progress has been made to set up good models to study MTM1 and the XLCNM disease such as knockout or knockin mice, the Labrador Retriever dog, the zebrafish and the yeast Saccharomyces cerevisiae. These helped to better understand the cellular function of MTM1 and of its four conserved domains: PH-GRAM (Pleckstrin Homology-Glucosyltransferase, Rab-like GTPase Activator and Myotubularin), RID (Rac1-Induced recruitment Domain), PTP/DSP (Protein Tyrosine Phosphatase/Dual-Specificity Phosphatase) and SID (SET-protein Interaction Domain). -

A Peripheral Blood Gene Expression Signature to Diagnose Subclinical Acute Rejection
CLINICAL RESEARCH www.jasn.org A Peripheral Blood Gene Expression Signature to Diagnose Subclinical Acute Rejection Weijia Zhang,1 Zhengzi Yi,1 Karen L. Keung,2 Huimin Shang,3 Chengguo Wei,1 Paolo Cravedi,1 Zeguo Sun,1 Caixia Xi,1 Christopher Woytovich,1 Samira Farouk,1 Weiqing Huang,1 Khadija Banu,1 Lorenzo Gallon,4 Ciara N. Magee,5 Nader Najafian,5 Milagros Samaniego,6 Arjang Djamali ,7 Stephen I. Alexander,2 Ivy A. Rosales,8 Rex Neal Smith,8 Jenny Xiang,3 Evelyne Lerut,9 Dirk Kuypers,10,11 Maarten Naesens ,10,11 Philip J. O’Connell,2 Robert Colvin,8 Madhav C. Menon,1 and Barbara Murphy1 Due to the number of contributing authors, the affiliations are listed at the end of this article. ABSTRACT Background In kidney transplant recipients, surveillance biopsies can reveal, despite stable graft function, histologic features of acute rejection and borderline changes that are associated with undesirable graft outcomes. Noninvasive biomarkers of subclinical acute rejection are needed to avoid the risks and costs associated with repeated biopsies. Methods We examined subclinical histologic and functional changes in kidney transplant recipients from the prospective Genomics of Chronic Allograft Rejection (GoCAR) study who underwent surveillance biopsies over 2 years, identifying those with subclinical or borderline acute cellular rejection (ACR) at 3 months (ACR-3) post-transplant. We performed RNA sequencing on whole blood collected from 88 indi- viduals at the time of 3-month surveillance biopsy to identify transcripts associated with ACR-3, developed a novel sequencing-based targeted expression assay, and validated this gene signature in an independent cohort. -

Pharmacological Targeting of the Mitochondrial Phosphatase PTPMT1 by Dahlia Doughty Shenton Department of Biochemistry Duke
Pharmacological Targeting of the Mitochondrial Phosphatase PTPMT1 by Dahlia Doughty Shenton Department of Biochemistry Duke University Date: May 1 st 2009 Approved: ___________________________ Dr. Patrick J. Casey, Supervisor ___________________________ Dr. Perry J. Blackshear ___________________________ Dr. Anthony R. Means ___________________________ Dr. Christopher B. Newgard ___________________________ Dr. John D. York Dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Biochemistry in the Graduate School of Duke University 2009 ABSTRACT Pharmacological Targeting of the Mitochondrial Phosphatase PTPMT1 by Dahlia Doughty Shenton Department of Biochemistry Duke University Date: May 1 st 2009 Approved: ___________________________ Dr. Patrick J. Casey, Supervisor ___________________________ Dr. Perry J. Blackshear ___________________________ Dr. Anthony R. Means ___________________________ Dr. Christopher B. Newgard ___________________________ Dr. John D. York An abstract of a dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Biochemistry in the Graduate School of Duke University 2009 Copyright by Dahlia Doughty Shenton 2009 Abstract The dual specificity protein tyrosine phosphatases comprise the largest and most diverse group of protein tyrosine phosphatases and play integral roles in the regulation of cell signaling events. The dual specificity protein tyrosine phosphatases impact multiple -

Deciphering the Functional and Molecular Differences Between MTM1 and MTMR2 to Better Understand Two Neuromuscular Diseases Matthieu Raess
Deciphering the functional and molecular differences between MTM1 and MTMR2 to better understand two neuromuscular diseases Matthieu Raess To cite this version: Matthieu Raess. Deciphering the functional and molecular differences between MTM1 and MTMR2 to better understand two neuromuscular diseases. Genomics [q-bio.GN]. Université de Strasbourg, 2017. English. NNT : 2017STRAJ088. tel-03081300 HAL Id: tel-03081300 https://tel.archives-ouvertes.fr/tel-03081300 Submitted on 18 Dec 2020 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. UNIVERSITÉ DE STRASBOURG ÉCOLE DOCTORALE DES SCIENCES DE LA VIE ET DE LA SANTE (ED 414) Génétique Moléculaire, Génomique, Microbiologie (GMGM) – UMR 7156 & Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC) UMR 7104 – INSERM U 964 THÈSE présentée par : Matthieu RAESS soutenue le : 13 octobre 2017 pour obtenir le grade de : Docteur de l’université de Strasbourg Discipline/ Spécialité : Aspects moléculaires et cellulaires de la biologie Deciphering the functional and molecular differences between MTM1 and MTMR2 to better understand two neuromuscular diseases. THÈSE dirigée par : Mme FRIANT Sylvie Directrice de recherche, Université de Strasbourg & Mme COWLING Belinda Chargée de recherche, Université de Strasbourg RAPPORTEURS : Mme BOLINO Alessandra Directrice de recherche, Institut San Raffaele de Milan M. -

Supplementary Table 1
Supplementary Table 1. 492 genes are unique to 0 h post-heat timepoint. The name, p-value, fold change, location and family of each gene are indicated. Genes were filtered for an absolute value log2 ration 1.5 and a significance value of p ≤ 0.05. Symbol p-value Log Gene Name Location Family Ratio ABCA13 1.87E-02 3.292 ATP-binding cassette, sub-family unknown transporter A (ABC1), member 13 ABCB1 1.93E-02 −1.819 ATP-binding cassette, sub-family Plasma transporter B (MDR/TAP), member 1 Membrane ABCC3 2.83E-02 2.016 ATP-binding cassette, sub-family Plasma transporter C (CFTR/MRP), member 3 Membrane ABHD6 7.79E-03 −2.717 abhydrolase domain containing 6 Cytoplasm enzyme ACAT1 4.10E-02 3.009 acetyl-CoA acetyltransferase 1 Cytoplasm enzyme ACBD4 2.66E-03 1.722 acyl-CoA binding domain unknown other containing 4 ACSL5 1.86E-02 −2.876 acyl-CoA synthetase long-chain Cytoplasm enzyme family member 5 ADAM23 3.33E-02 −3.008 ADAM metallopeptidase domain Plasma peptidase 23 Membrane ADAM29 5.58E-03 3.463 ADAM metallopeptidase domain Plasma peptidase 29 Membrane ADAMTS17 2.67E-04 3.051 ADAM metallopeptidase with Extracellular other thrombospondin type 1 motif, 17 Space ADCYAP1R1 1.20E-02 1.848 adenylate cyclase activating Plasma G-protein polypeptide 1 (pituitary) receptor Membrane coupled type I receptor ADH6 (includes 4.02E-02 −1.845 alcohol dehydrogenase 6 (class Cytoplasm enzyme EG:130) V) AHSA2 1.54E-04 −1.6 AHA1, activator of heat shock unknown other 90kDa protein ATPase homolog 2 (yeast) AK5 3.32E-02 1.658 adenylate kinase 5 Cytoplasm kinase AK7 -
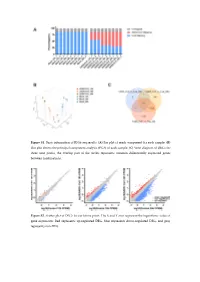
Figure S1. Basic Information of RNA-Seq Results. (A) Bar Plot of Reads Component for Each Sample
Figure S1. Basic information of RNA-seq results. (A) Bar plot of reads component for each sample. (B) Dot plot shows the principal component analysis (PCA) of each sample. (C) Venn diagram of DEGs for three time points, the overlap part of the circles represents common differentially expressed genes between combinations. Figure S2. Scatter plot of DEGs for each time point. The X and Y axes represent the logarithmic value of gene expression. Red represents up-regulated DEG, blue represents down-regulated DEG, and gray represents non-DEG. Table S1. Primers used for quantitative real-time PCR analysis of DEGs. Gene Primer Sequence Forward 5’-CTACGAGTGGATGGTCAAGAGC-3’ FOXO1 Reverse 5’-CCAGTTCCTTCATTCTGCACACG-3’ Forward 5’-GACGTCCGGCATCAGAGAAA-3’ IRS2 Reverse 5’-TCCACGGCTAATCGTCACAG-3’ Forward 5’-CACAACCAGGACCTCACACC-3’ IRS1 Reverse 5’-CTTGGCACGATAGAGAGCGT-3’ Forward 5’-AGGATACCACTCCCAACAGACCT-3’ IL6 Reverse 5’-CAAGTGCATCATCGTTGTTCATAC-3’ Forward 5’-TCACGTTGTACGCAGCTACC-3’ CCL5 Reverse 5’-CAGTCCTCTTACAGCCTTTGG-3’ Forward 5’-CTGTGCAGCCGCAGTGCCTACC-3’ BMP7 Reverse 5’-ATCCCTCCCCACCCCACCATCT-3’ Forward 5’-CTCTCCCCCTCGACTTCTGA-3’ BCL2 Reverse 5’-AGTCACGCGGAACACTTGAT-3’ Forward 5’-CTGTCGAACACAGTGGTACCTG-3’ FGF7 Reverse 5’-CCAACTGCCACTGTCCTGATTTC-3’ Forward 5’-GGGAGCCAAAAGGGTCATCA-3’ GAPDH Reverse 5’-CGTGGACTGTGGTCATGAGT-3’ Supplementary material: Differentially expressed genes log2(SADS-CoV_12h/ Qvalue (SADS-CoV _12h/ Gene Symbol Control_12h) Control_12h) PTGER4 -1.03693 6.79E-04 TMEM72 -3.08132 3.66E-04 IFIT2 -1.02918 2.11E-07 FRAT2 -1.09282 4.66E-05 -

Lipid Phosphatases Identified by Screening a Mouse Phosphatase Shrna Library Regulate T-Cell Differentiation and Protein Kinase
Lipid phosphatases identified by screening a mouse PNAS PLUS phosphatase shRNA library regulate T-cell differentiation and Protein kinase B AKT signaling Liying Guoa, Craig Martensb, Daniel Brunob, Stephen F. Porcellab, Hidehiro Yamanea, Stephane M. Caucheteuxa, Jinfang Zhuc, and William E. Paula,1 aCytokine Biology Unit, cMolecular and Cellular Immunoregulation Unit, Laboratory of Immunology, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD 20892; and bGenomics Unit, Research Technologies Section, Rocky Mountain Laboratories, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Hamilton, MT 59840 Contributed by William E. Paul, March 27, 2013 (sent for review December 18, 2012) Screening a complete mouse phosphatase lentiviral shRNA library production (10, 11). Conversely, constitutive expression of active using high-throughput sequencing revealed several phosphatases AKT leads to increased proliferation and enhanced Th1/Th2 cy- that regulate CD4 T-cell differentiation. We concentrated on two lipid tokine production (12). phosphatases, the myotubularin-related protein (MTMR)9 and -7. The amount of PI[3,4,5]P3 and the level of AKT activation are Silencing MTMR9 by shRNA or siRNA resulted in enhanced T-helper tightly controlled by several mechanisms, including breakdown of (Th)1 differentiation and increased Th1 protein kinase B (PKB)/AKT PI[3,4,5]P3, down-regulation of the amount and activity of PI3K, phosphorylation while silencing MTMR7 caused increased Th2 and and the dephosphorylation of AKT (13). PTEN is a major negative Th17 differentiation and increased AKT phosphorylation in these regulator of PI[3,4,5]P3. It removes the 3-phosphate from the cells.