YWHAZ Promotes Ovarian Cancer Metastasis by Modulating Glycolysis
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

RT² Profiler PCR Array (384-Well Format) Human Apoptosis 384HT
RT² Profiler PCR Array (384-Well Format) Human Apoptosis 384HT Cat. no. 330231 PAHS-3012ZE For pathway expression analysis Format For use with the following real-time cyclers RT² Profiler PCR Array, Applied Biosystems® models 7900HT (384-well block), Format E ViiA™ 7 (384-well block); Bio-Rad CFX384™ RT² Profiler PCR Array, Roche® LightCycler® 480 (384-well block) Format G Description The Human Apoptosis RT² Profiler PCR Array profiles the expression of 370 key genes involved in apoptosis, or programmed cell death. The array includes the TNF ligands and their receptors; members of the bcl-2 family, BIR (baculoviral IAP repeat) domain proteins, CARD domain (caspase recruitment domain) proteins, death domain proteins, TRAF (TNF receptor-associated factor) domain proteins and caspases. These 370 genes are also grouped by their functional contribution to apoptosis, either anti-apoptosis or induction of apoptosis. Using real-time PCR, you can easily and reliably analyze expression of a focused panel of genes related to apoptosis with this array. For further details, consult the RT² Profiler PCR Array Handbook. Sample & Assay Technologies Shipping and storage RT² Profiler PCR Arrays in formats E and G are shipped at ambient temperature, on dry ice, or blue ice packs depending on destination and accompanying products. For long term storage, keep plates at –20°C. Note: Ensure that you have the correct RT² Profiler PCR Array format for your real-time cycler (see table above). Note: Open the package and store the products appropriately immediately -

The Study of Two Transmembrane Autophagy Proteins and the Autophagy Receptor, P62
The study of two transmembrane autophagy proteins and the autophagy receptor, p62 Gautam M. Runwal St. John’s College, Cambridge September 2018 This dissertation is submitted for the degree of Doctor of Philosophy Title: The study of two transmembrane autophagy proteins and the autophagy receptor, p62 Submitted by : Gautam M Runwal Abstract Autophagy is an evolutionarily conserved process across eukaryotes that is responsible for degradation of cargo such as aggregate-prone proteins, pathogens, damaged organelles, macromolecules etc. via its delivery to lysosomes. The process is known to involve the formation of a double-membraned structure, called autophagosome, that engulfs the cargo destined for degradation and delivers its contents by fusing with lysosomes. This process involves several proteins at its core which include two transmembrane proteins, ATG9 and VMP1. While ATG9 and VMP1 has been discovered for about a decade and half, the trafficking and function of these proteins remain relatively unclear. My work in this thesis identifies and characterises a novel trafficking route for ATG9 and VMP1 and shows that both these proteins traffic via the dynamin-independent ARF6-associated pathway. Moreover, I also show that these proteins physically interact with each other. In addition, the tools developed during these studies helped me identify a new role for the most common autophagy receptor protein, p62. I show that p62 can specifically associate with and sequester LC3-I in autophagy- impaired cells (ATG9 and ATG16 null cells) leading to formation of LC3-positive structures that can be misinterpreted as mature autophagosomes. Perturbations in the levels of p62 were seen to affect the formation of these LC3-positive structures in cells. -

Mir-451A Suppresses Cell Proliferation, Metastasis and EMT Via Targeting YWHAZ in Hepatocellular Carcinoma
European Review for Medical and Pharmacological Sciences 2019; 23: 5158-5167 MiR-451a suppresses cell proliferation, metastasis and EMT via targeting YWHAZ in hepatocellular carcinoma G.-Y. WEI1, M. HU2, L. ZHAO1, W.-S. GUO1 1Department of General Surgery, Henan Traditional Chinese Medicine Hospital, Zhengzhou, China 2Department of Infection Management, Henan Traditional Chinese Medicine Hospital, Zhengzhou, China Abstract. – OBJECTIVE: MicroRNAs (miR- lines. Moreover, miR-451a inhibited the prolif- NAs) have been identified to participate in eration, invasion and migration of HCC cells via the progression of hepatocellular carcinoma targeting YWHAZ. Our findings indicated that (HCC). However, the function of miR-451a in miR-451a could serve as a novel target for HCC HCC remains unknown. The aim of this study diagnosis and biological therapy. was to determine the function of miR-451a by construction of several experiments in HCC tis- Key Words: sues and cells. MiR-451a, Hepatocellular carcinoma (HCC), Prolifer- PATIENTS AND METHODS: The expression ation, Metastasis, YWHAZ. level of miR-451a in 69 paired of HCC and ad- jacent normal tissue samples was detected us- ing quantitative Real-time polymerase chain re- Introduction action (qRT-PCR). MiR-451a expression in HCC derived cell lines was detected as well. By trans- fecting with miR-451a mimics or inhibitor, the Hepatocellular carcinoma (HCC) accounts expression level of miR-451a in HepG2 or Huh-7 for 85-90% of primary liver cancer. HCC is the cells was up- or down-regulated. MTT (3-(4,5-di- fifth most common morbidity and third most methylthiazol-2-yl)-2,5-diphenyl tetrazolium bro- common mortality cancer worldwide1. -

Oncorequisite Role of an Aldehyde Dehydrogenase in the Pathogenesis of T-Cell Acute Lymphoblastic Leukemia
Acute Lymphoblastic Leukemia SUPPLEMENTARY APPENDIX Oncorequisite role of an aldehyde dehydrogenase in the pathogenesis of T-cell acute lymphoblastic leukemia Chujing Zhang, 1 Stella Amanda, 1 Cheng Wang, 2 Tze King Tan, 1 Muhammad Zulfaqar Ali, 1 Wei Zhong Leong, 1 Ley Moy Ng, 1 Shojiro Kitajima, 3 Zhenhua Li, 4 Allen Eng Juh Yeoh, 1,4 Shi Hao Tan 1 and Takaomi Sanda 1,5 1Cancer Science Institute of Singapore, National University of Singapore, Singapore; 2Department of Anatomy, National University of Singapore, Singapore; 3Institute for Advanced Biosciences, Keio University, Tsuruoka, Japan; 4VIVA-NUS CenTRAL, Department of Pedi - atrics, National University of Singapore, Singapore and 5Department of Medicine, Yong Loo Lin School of Medicine, National University of Singapore, Singapore. ©2021 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol. 2019.245639 Received: December 18, 2019. Accepted: May 14, 2020. Pre-published: May 15, 2020. Correspondence: TAKAOMI SANDA - [email protected] Zhang et al Roles of ALDH in T-ALL Supplementary Information Supplementary Materials and Methods Cell culture and reagents All human leukemia cell lines were cultured in RPMI-1640 medium (BioWest) supplemented with 10% FBS (BioWest). 293T cells were maintained in DMEM medium (BioWest) supplemented with 10% FBS and penicillin/streptomycin (Life Technologies). All-trans retinaldehyde was purchased from Sigma-Aldrich. Glucose free and Glutamine free RPMI- 1640 media were purchased from ThermoFisher. Patient derived xenograft (PDX) sample T-ALL patient-derived PDX sample (DFCI15) was kindly provided by Alejandro Gutierrez (Boston Children’s Hospital, Boston). Mouse studies were conducted according to the recommendations from the Institutional Animal Care and Use Committee (IACUC) and all protocols were approved by the Committee at the National University of Singapore (NUS). -

13509 Atg9a (D4O9D) Rabbit Mab
Revision 3 C 0 2 - t Atg9A (D4O9D) Rabbit mAb a e r o t S Orders: 877-616-CELL (2355) [email protected] 9 Support: 877-678-TECH (8324) 0 5 Web: [email protected] 3 www.cellsignal.com 1 # 3 Trask Lane Danvers Massachusetts 01923 USA For Research Use Only. Not For Use In Diagnostic Procedures. Applications: Reactivity: Sensitivity: MW (kDa): Source/Isotype: UniProt ID: Entrez-Gene Id: WB, IP H M R Mk Endogenous 100-110 Rabbit IgG Q7Z3C6 79065 Product Usage Information 3. Levine, B. and Yuan, J. (2005) J Clin Invest 115, 2679-88. 4. Klionsky, D.J. et al. (2003) Dev Cell 5, 539-45. Application Dilution 5. Noda, T. et al. (2000) J Cell Biol 148, 465-80. 6. Young, A.R. et al. (2006) J Cell Sci 119, 3888-900. Western Blotting 1:1000 7. Yamada, T. et al. (2005) J Biol Chem 280, 18283-90. Immunoprecipitation 1:50 8. Orsi, A. et al. (2012) Mol Biol Cell 23, 1860-73. 9. Webber, J.L. and Tooze, S.A. (2010) EMBO J 29, 27-40. Storage 10. Takahashi, Y. et al. (2011) Autophagy 7, 61-73. Supplied in 10 mM sodium HEPES (pH 7.5), 150 mM NaCl, 100 µg/ml BSA, 50% glycerol and less than 0.02% sodium azide. Store at –20°C. Do not aliquot the antibody. Specificity / Sensitivity Atg9A (D4O9D) Rabbit mAb recognizes endogenous levels of total Atg9A protein. Species Reactivity: Human, Mouse, Rat, Monkey Source / Purification Monoclonal antibody is produced by immunizing animals with a synthetic peptide corresponding to residues surrounding Gly780 of human Atg9A protein. -

Prognostic Significance of Autophagy-Relevant Gene Markers in Colorectal Cancer
ORIGINAL RESEARCH published: 15 April 2021 doi: 10.3389/fonc.2021.566539 Prognostic Significance of Autophagy-Relevant Gene Markers in Colorectal Cancer Qinglian He 1, Ziqi Li 1, Jinbao Yin 1, Yuling Li 2, Yuting Yin 1, Xue Lei 1 and Wei Zhu 1* 1 Department of Pathology, Guangdong Medical University, Dongguan, China, 2 Department of Pathology, Dongguan People’s Hospital, Southern Medical University, Dongguan, China Background: Colorectal cancer (CRC) is a common malignant solid tumor with an extremely low survival rate after relapse. Previous investigations have shown that autophagy possesses a crucial function in tumors. However, there is no consensus on the value of autophagy-associated genes in predicting the prognosis of CRC patients. Edited by: This work screens autophagy-related markers and signaling pathways that may Fenglin Liu, Fudan University, China participate in the development of CRC, and establishes a prognostic model of CRC Reviewed by: based on autophagy-associated genes. Brian M. Olson, Emory University, United States Methods: Gene transcripts from the TCGA database and autophagy-associated gene Zhengzhi Zou, data from the GeneCards database were used to obtain expression levels of autophagy- South China Normal University, China associated genes, followed by Wilcox tests to screen for autophagy-related differentially Faqing Tian, Longgang District People's expressed genes. Then, 11 key autophagy-associated genes were identified through Hospital of Shenzhen, China univariate and multivariate Cox proportional hazard regression analysis and used to Yibing Chen, Zhengzhou University, China establish prognostic models. Additionally, immunohistochemical and CRC cell line data Jian Tu, were used to evaluate the results of our three autophagy-associated genes EPHB2, University of South China, China NOL3, and SNAI1 in TCGA. -

Emerging Roles of P53 Family Members in Glucose Metabolism
International Journal of Molecular Sciences Review Emerging Roles of p53 Family Members in Glucose Metabolism Yoko Itahana and Koji Itahana * Cancer and Stem Cell Biology Program, Duke-NUS Medical School, 8 College Road, Singapore 169857, Singapore; [email protected] * Correspondence: [email protected]; Tel.: +65-6516-2554; Fax: +65-6221-2402 Received: 19 January 2018; Accepted: 22 February 2018; Published: 8 March 2018 Abstract: Glucose is the key source for most organisms to provide energy, as well as the key source for metabolites to generate building blocks in cells. The deregulation of glucose homeostasis occurs in various diseases, including the enhanced aerobic glycolysis that is observed in cancers, and insulin resistance in diabetes. Although p53 is thought to suppress tumorigenesis primarily by inducing cell cycle arrest, apoptosis, and senescence in response to stress, the non-canonical functions of p53 in cellular energy homeostasis and metabolism are also emerging as critical factors for tumor suppression. Increasing evidence suggests that p53 plays a significant role in regulating glucose homeostasis. Furthermore, the p53 family members p63 and p73, as well as gain-of-function p53 mutants, are also involved in glucose metabolism. Indeed, how this protein family regulates cellular energy levels is complicated and difficult to disentangle. This review discusses the roles of the p53 family in multiple metabolic processes, such as glycolysis, gluconeogenesis, aerobic respiration, and autophagy. We also discuss how the dysregulation of the p53 family in these processes leads to diseases such as cancer and diabetes. Elucidating the complexities of the p53 family members in glucose homeostasis will improve our understanding of these diseases. -

6-Phosphofructo-2-Kinase/Fructose-2
Published OnlineFirst August 7, 2019; DOI: 10.1158/1078-0432.CCR-18-3448 Translational Cancer Mechanisms and Therapy Clinical Cancer Research 6-Phosphofructo-2-Kinase/Fructose-2,6- Biphosphatase-2 Regulates TP53-Dependent Paclitaxel Sensitivity in Ovarian and Breast Cancers Hailing Yang1, Zhang Shu1,2,Yongying Jiang3, Weiqun Mao1, Lan Pang1, Abena Redwood4, Sabrina L. Jeter-Jones4, Nicholas B. Jennings5, Argentina Ornelas6, Jinhua Zhou1, Cristian Rodriguez-Aguayo1,7, Geoffrey Bartholomeusz1, LaKesla R. Iles1, Niki M. Zacharias8, Steven W. Millward6, Gabriel Lopez-Berestein1,7, Xiao-Feng Le1, Ahmed A. Ahmed9,10, Helen Piwnica-Worms4, Anil K. Sood5,7, Robert C. Bast1, and Zhen Lu1 Abstract Purpose: Paclitaxel is an integral component of primary Results: Knockdown of PFKFB2 inhibited clonogenic therapy for breast and epithelial ovarian cancers, but less than growth and enhanced paclitaxel sensitivity in ovarian and half of these cancers respond to the drug. Enhancing the breast cancer cell lines with wild-type TP53 (wtTP53). response to primary therapy with paclitaxel could improve Silencing PFKFB2 significantly inhibited tumor growth and outcomes for women with both diseases. enhanced paclitaxel sensitivity in four xenografts derived Experimental Design: Twelve kinases that regulate from two ovarian and two breast cancer cell lines, and metabolism were depleted in multiple ovarian and breast prolonged survival in a triple-negative breast cancer PDX. cancer cell lines to determine whether they regulate sensi- Transfection of siPFKFB2 increased the glycolysis rate, but tivity to paclitaxel in Sulforhodamine B assays. The effects decreased the flow of intermediates through the pentose– of 6-phosphofructo-2-kinase/fructose-2,6-bisphosphatase phosphate pathway in cancer cells with wtTP53,decreasing 2(PFKFB2) depletion on cell metabolomics, extracellular NADPH. -

Identification and Validation of a Six-Gene Signature Associated With
Cai et al. BMC Cancer (2020) 20:1133 https://doi.org/10.1186/s12885-020-07598-3 RESEARCH ARTICLE Open Access Identification and validation of a six-gene signature associated with glycolysis to predict the prognosis of patients with cervical cancer Luya Cai1†, Chuan Hu2†, Shanshan Yu3, Lixiao Liu1, Xiaobo Yu1, Jiahua Chen1, Xuan Liu1, Fan Lin4, Cheng Zhang4, Wenfeng Li3 and Xiaojian Yan1* Abstract Background: Cervical cancer (CC) is one of the most common gynaecological cancers. The gene signature is believed to be reliable for predicting cancer patient survival. However, there is no relevant study on the relationship between the glycolysis-related gene (GRG) signature and overall survival (OS) of patients with CC. Methods: We extracted the mRNA expression profiles of 306 tumour and 13 normal tissues from the University of California Santa Cruz (UCSC) Database. Then, we screened out differentially expressed glycolysis-related genes (DEGRGs) among these mRNAs. All patients were randomly divided into training cohort and validation cohort according to the ratio of 7: 3. Next, univariate and multivariate Cox regression analyses were carried out to select the GRG with predictive ability for the prognosis of the training cohort. Additionally, risk score model was constructed and validated it in the validation cohort. Results: Six mRNAs were obtained that were associated with patient survival. The filtered mRNAs were classified into the protective type (GOT1) and the risk type (HSPA5, ANGPTL4, PFKM, IER3 and PFKFB4). Additionally, by constructing the prognostic risk score model, we found that the OS of the high-risk group was notably poorer, which showed good predictive ability both in training cohort and validation cohort. -
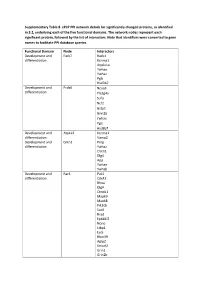
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4 -

Glycolytic Reliance Promotes Anabolism in Photoreceptors
bioRxiv preprint doi: https://doi.org/10.1101/101964; this version posted January 21, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Glycolytic reliance promotes anabolism in photoreceptors Yashodhan Chinchore, Tedi Begaj, David Wu, Eugene Drokhlyansky, Constance L. Cepko* *Departments of Genetics and Ophthalmology, Howard Hughes Medical Institute, Harvard Medical School, Boston, Massachusetts 02115, USA [email protected] bioRxiv preprint doi: https://doi.org/10.1101/101964; this version posted January 21, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Sensory neurons capture information from the environment and convert it 2 into signals that can greatly impact the survival of an organism. These systems 3 are thus under heavy selective pressure, including for the most efficient use of 4 energy to support their sensitivity and efficiency1. In this regard, the 5 vertebrate photoreceptor cells face a dual challenge. They not only need to 6 preserve their membrane excitability via ion pumps by ATP hydrolysis2 but 7 also maintain a highly membrane rich organelle, the outer segment, which is 8 the primary site of phototransduction, creating a considerable biosynthetic 9 demand. How photoreceptors manage carbon allocation to balance their 10 catabolic and anabolic demands is poorly understood. One metabolic feature 11 of the retina is its ability to convert the majority of its glucose into lactate3,4 12 even in the presence of oxygen. -

Characterization of Genes in the CFTR-Mediated Innate Immune Response
The University of Maine DigitalCommons@UMaine Honors College 5-2012 Characterization of Genes in the CFTR-Mediated Innate Immune Response Eric Peterman Follow this and additional works at: https://digitalcommons.library.umaine.edu/honors Part of the Cell and Developmental Biology Commons, and the Molecular Biology Commons Recommended Citation Peterman, Eric, "Characterization of Genes in the CFTR-Mediated Innate Immune Response" (2012). Honors College. 71. https://digitalcommons.library.umaine.edu/honors/71 This Honors Thesis is brought to you for free and open access by DigitalCommons@UMaine. It has been accepted for inclusion in Honors College by an authorized administrator of DigitalCommons@UMaine. For more information, please contact [email protected]. CHARACTERIZATION OF GENES IN THE CFTR-MEDIATED INNATE IMMUNE RESPONSE by Eric Peterman A Thesis Submitted in Partial Fulfillment of the Requirements for a Degree with Honors (Biochemistry, Molecular and Cellular Biology) The Honors College University of Maine May 2012 Advisory Committee: Carol Kim, Professor, Molecular & Biomedical Sciences Robert Gundersen, Associate Professor, Molecular & Biomedical Sciences John Singer, Professor, Molecular & Biomedical Sciences Julie Gosse, Assistant Professor, Molecular & Biomedical Sciences Keith Hutchison, Professor, Molecular & Biomedical Sciences Mark Haggerty, Lecturer, Rezendes Preceptor for Civic Engagement Abstract: Recently, the Kim Lab has shown that the cystic fibrosis transmembrane conductance regulator (cftr) gene is responsible for mediating resistance to Pseudomonas aeruginosa in a zebrafish infection model. Using the Gene Expression Omnibus, an NCBI functional genomics data repository, it was determined that Smad3, a transcription factor in the TGF-β signaling pathway, is upregulated in the presence of P. aeruginosa. It was found that in our zebrafish model, the Smad3 paralogs Smad3a and Smad3b are upregulated following microinjection of a cftr antisense morpholino oligomer.