Folding and Stability of Ankyrin Repeats Control Biological Protein Function
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

INK4 Locus of the Tumor-Resistant Rodent, the Naked Mole Rat, Expresses a Functional P15/P16 Hybrid Isoform
INK4 locus of the tumor-resistant rodent, the naked mole rat, expresses a functional p15/p16 hybrid isoform Xiao Tiana,1, Jorge Azpuruaa,1,2, Zhonghe Kea, Adeline Augereaub, Zhengdong D. Zhangc, Jan Vijgc, Vadim N. Gladyshevb, Vera Gorbunovaa,3, and Andrei Seluanova,3 aDepartment of Biology, University of Rochester, Rochester, NY 14627; bBrigham and Women’s Hospital, Harvard Medical School, Boston, MA 02115; and cAlbert Einstein College of Medicine, Bronx, NY 10461 Edited* by Eviatar Nevo, Institute of Evolution, Haifa, Israel, and approved December 1, 2014 (received for review September 21, 2014) The naked mole rat (Heterocephalus glaber) is a long-lived and because, in mammals, it encodes three distinct tumor suppressors: tumor-resistant rodent. Tumor resistance in the naked mole rat p16INK4a, p15INK4b, and p19ARF (p14ARF in human) (9). These is mediated by the extracellular matrix component hyaluronan three proteins coordinate a signaling network that depends on of very high molecular weight (HMW-HA). HMW-HA triggers hy- the activities of the retinoblastoma protein (RB) and the p53 persensitivity of naked mole rat cells to contact inhibition, which is tumor suppressor protein. The p16INK4a and p15INK4b pro- associated with induction of the INK4 (inhibitors of cyclin depen- teins are cyclin-dependent kinase inhibitors that directly in- dent kinase 4) locus leading to cell-cycle arrest. The INK4a/b locus is hibit the binding of cyclins to their target cyclin-dependent among the most frequently mutated in human cancer. This locus kinases (10). p16INK4a is involved in establishing replicative encodes three distinct tumor suppressors: p15INK4b, p16INK4a, and senescence, oncogene-induced senescence, and stress-induced INK4b ARF (alternate reading frame). -

Nucleocytoplasmic Shuttling of the Glucocorticoid Receptor Is Influenced by Tetratricopeptide Repeat-Containing Proteins Gisela I
© 2020. Published by The Company of Biologists Ltd | Journal of Cell Science (2020) 133, jcs238873. doi:10.1242/jcs.238873 RESEARCH ARTICLE Nucleocytoplasmic shuttling of the glucocorticoid receptor is influenced by tetratricopeptide repeat-containing proteins Gisela I. Mazaira1,*, Pablo C. Echeverria2,* and Mario D. Galigniana1,3,‡ ABSTRACT FKBP52 (also known as FKBP4), FKBP51 (FKBP5) and CyP40 It has been demonstrated that tetratricopeptide-repeat (TPR) domain (PPID), and the immunophilin-like proteins PP5, FKBPL (WISp39) proteins regulate the subcellular localization of glucocorticoid receptor and FKBP37 (ARA9/XAP2/AIP) are counted among the best (GR). This study analyses the influence of the TPR domain of high characterized members of the TPR domain family of proteins molecular weight immunophilins in the retrograde transport and associated with nuclear receptor family members via Hsp90 (Pratt nuclear retention of GR. Overexpression of the TPR peptide et al., 2004a; Storer et al., 2011). TPR domains are composed of prevented efficient nuclear accumulation of the GR by disrupting the loosely conserved 34 amino acid sequence motifs that are found in formation of complexes with the dynein-associated immunophilin arrays comprising between 2 and 20 repeats, wherein the repeats pack FKBP52 (also known as FKBP4), the adaptor transporter importin-β1 against each other giving rise to coiled and superhelical structures (KPNB1), the nuclear pore-associated glycoprotein Nup62 and nuclear (Perez-Riba and Itzhaki, 2019). Originally identified in components of matrix-associated structures. We also show that nuclear import of GR the anaphase-promoting complex (Sikorski et al., 1991), TPR was impaired, whereas GR nuclear export was enhanced. domains are now known to mediate specific protein interactions in Interestingly, the CRM1 (exportin-1) inhibitor leptomycin-B abolished numerous cellular contexts. -

Ankrd9 Is a Metabolically-Controlled Regulator of Impdh2 Abundance and Macro-Assembly
ANKRD9 IS A METABOLICALLY-CONTROLLED REGULATOR OF IMPDH2 ABUNDANCE AND MACRO-ASSEMBLY by Dawn Hayward A dissertation submitted to The Johns Hopkins University in conformity with the requirements of the degree of Doctor of Philosophy Baltimore, Maryland April 2019 ABSTRACT Members of a large family of Ankyrin Repeat Domains proteins (ANKRD) regulate numerous cellular processes by binding and changing properties of specific protein targets. We show that interactions with a target protein and the functional outcomes can be markedly altered by cells’ metabolic state. ANKRD9 facilitates degradation of inosine monophosphate dehydrogenase 2 (IMPDH2), the rate-limiting enzyme in GTP biosynthesis. Under basal conditions ANKRD9 is largely segregated from the cytosolic IMPDH2 by binding to vesicles. Upon nutrient limitation, ANKRD9 loses association with vesicles and assembles with IMPDH2 into rod-like structures, in which IMPDH2 is stable. Inhibition of IMPDH2 with Ribavirin favors ANKRD9 binding to rods. The IMPDH2/ANKRD9 assembly is reversed by guanosine, which restores association of ANKRD9 with vesicles. The conserved Cys109Cys110 motif in ANKRD9 is required for the vesicles-to-rods transition as well as binding and regulation of IMPDH2. ANKRD9 knockdown increases IMPDH2 levels and prevents formation of IMPDH2 rods upon nutrient limitation. Thus, the status of guanosine pools affects the mode of ANKRD9 action towards IMPDH2. Advisor: Dr. Svetlana Lutsenko, Department of Physiology, Johns Hopkins University School of Medicine Second reader: -

Repeat Proteins Challenge the Concept of Structural Domains
844 Biochemical Society Transactions (2015) Volume 43, part 5 Repeat proteins challenge the concept of structural domains Rocıo´ Espada*1, R. Gonzalo Parra*1, Manfred J. Sippl†, Thierry Mora‡, Aleksandra M. Walczak§ and Diego U. Ferreiro*2 *Protein Physiology Lab, Dep de Qu´ımica Biologica, ´ Facultad de Ciencias Exactas y Naturales, UBA-CONICET-IQUIBICEN, Buenos Aires, C1430EGA, Argentina †Center of Applied Molecular Engineering, Division of Bioinformatics, Department of Molecular Biology, University of Salzburg, 5020 Salzburg, Austria ‡Laboratoire de physique statistique, CNRS, UPMC and Ecole normale superieure, ´ 24 rue Lhomond, 75005 Paris, France §Laboratoire de physique theorique, ´ CNRS, UPMC and Ecole normale superieure, ´ 24 rue Lhomond, 75005 Paris, France Abstract Structural domains are believed to be modules within proteins that can fold and function independently. Some proteins show tandem repetitions of apparent modular structure that do not fold independently, but rather co-operate in stabilizing structural forms that comprise several repeat-units. For many natural repeat-proteins, it has been shown that weak energetic links between repeats lead to the breakdown of co-operativity and the appearance of folding sub-domains within an apparently regular repeat array. The quasi-1D architecture of repeat-proteins is crucial in detailing how the local energetic balances can modulate the folding dynamics of these proteins, which can be related to the physiological behaviour of these ubiquitous biological systems. Introduction and between repeats, challenging the concept of structural It was early on noted that many natural proteins typically domain. collapse stretches of amino acid chains into compact units, defining structural domains [1]. These domains typically correlate with biological activities and many modern proteins can be described as composed by novel ‘domain arrange- Definition of the repeat-units ments’ [2]. -

The Involvement of Ubiquitination Machinery in Cell Cycle Regulation and Cancer Progression
International Journal of Molecular Sciences Review The Involvement of Ubiquitination Machinery in Cell Cycle Regulation and Cancer Progression Tingting Zou and Zhenghong Lin * School of Life Sciences, Chongqing University, Chongqing 401331, China; [email protected] * Correspondence: [email protected] Abstract: The cell cycle is a collection of events by which cellular components such as genetic materials and cytoplasmic components are accurately divided into two daughter cells. The cell cycle transition is primarily driven by the activation of cyclin-dependent kinases (CDKs), which activities are regulated by the ubiquitin-mediated proteolysis of key regulators such as cyclins, CDK inhibitors (CKIs), other kinases and phosphatases. Thus, the ubiquitin-proteasome system (UPS) plays a pivotal role in the regulation of the cell cycle progression via recognition, interaction, and ubiquitination or deubiquitination of key proteins. The illegitimate degradation of tumor suppressor or abnormally high accumulation of oncoproteins often results in deregulation of cell proliferation, genomic instability, and cancer occurrence. In this review, we demonstrate the diversity and complexity of the regulation of UPS machinery of the cell cycle. A profound understanding of the ubiquitination machinery will provide new insights into the regulation of the cell cycle transition, cancer treatment, and the development of anti-cancer drugs. Keywords: cell cycle regulation; CDKs; cyclins; CKIs; UPS; E3 ubiquitin ligases; Deubiquitinases (DUBs) Citation: Zou, T.; Lin, Z. The Involvement of Ubiquitination Machinery in Cell Cycle Regulation and Cancer Progression. 1. Introduction Int. J. Mol. Sci. 2021, 22, 5754. https://doi.org/10.3390/ijms22115754 The cell cycle is a ubiquitous, complex, and highly regulated process that is involved in the sequential events during which a cell duplicates its genetic materials, grows, and di- Academic Editors: Kwang-Hyun Bae vides into two daughter cells. -

The Uvomorulin-Anchorage Protein a Catenin Is a Vinculin
Proc. Nail. Acad. Sci. USA Vol. 88, pp. 9156-9160, October 1991 Cell Biology The uvomorulin-anchorage protein a catenin is a vinculin homologue KURT HERRENKNECHT*, MASAYUKI OZAWA*t, CHRISTOPH ECKERSKORN*, FRIEDRICH LOTTSPEICHt, MARTIN LENTER*, AND ROLF KEMLER*§ *Max-Planck-Institut ffir Immunbiologie, FG Molekulare Embryologie, D-7800 Freiburg, Federal Republic of Germany; and tMax-Planck-Institut ffr Biochemie, D-8033 Martinsried, Federal Republic of Germany Communicated by Franqois Jacob, July 18, 1991 (receivedfor review June 25, 1991) ABSTRACT The cytoplasmic region of the Ca2+- domain is well conserved in other cadherins, it is possible that dependent cell-adhesion molecule (CAM) uvomorulin associ- catenins may also complex with other members of this gene ates with distinct cytoplasmic proteins with molecular masses family (13, 14). Here we have produced antibodies against a of 102, 88, and 80 kDa termed a, (3, and ycatenin, respectively. catenin and show that a catenin is indeed associated with This complex formation links uvomorulin to the actin filament cadherins from human, mouse, and Xenopus. We have network, which seems to be of primary importance for its cloned and sequenced¶ the cDNA coding for a catenin and cell-adhesion properties. We show here that antibodies against have established the primary protein structure. Sequence a catenin also immunoprecipitate complexes that contain hu- comparison reveals homology to vinculin, a well-known man N-cadherin, mouse P-cadherin, chicken A-CAM (adhe- adherens-type and focal contact protein. rens junction-specific CAM; also called N-cadherin) or Xeno- pus U-cadherin, demonstrating that a catenin is complexed with other cadherins. -

A Structural Guide to Proteins of the NF-Kb Signaling Module
Downloaded from http://cshperspectives.cshlp.org/ on September 26, 2021 - Published by Cold Spring Harbor Laboratory Press A Structural Guide to Proteins of the NF-kB Signaling Module Tom Huxford1 and Gourisankar Ghosh2 1Department of Chemistry and Biochemistry, San Diego State University, 5500 Campanile Drive, San Diego, California 92182-1030 2Department of Chemistry and Biochemistry, University of California, San Diego, 9500 Gilman Drive, La Jolla, California 92093-0375 Correspondence: [email protected] The prosurvival transcription factor NF-kB specifically binds promoter DNA to activate target gene expression. NF-kB is regulated through interactions with IkB inhibitor proteins. Active proteolysis of these IkB proteins is, in turn, under the control of the IkB kinase complex (IKK). Together, these three molecules form the NF-kB signaling module. Studies aimed at charac- terizing the molecular mechanisms of NF-kB, IkB, and IKK in terms of their three-dimen- sional structures have lead to a greater understanding of this vital transcription factor system. F-kB is a master transcription factor that from the perspective of their three-dimensional Nresponds to diverse cell stimuli by activat- structures. ing the expression of stress response genes. Multiple signals, including cytokines, growth factors, engagement of the T-cell receptor, and NF-kB bacterial and viral products, induce NF-kB Introduction to NF-kB transcriptional activity (Hayden and Ghosh 2008). A point of convergence for the myriad NF-kB was discovered in the laboratory of of NF-kB inducing signals is the IkB kinase David Baltimore as a nuclear activity with bind- complex (IKK). Active IKK in turn controls ing specificity toward a ten-base-pair DNA transcription factor NF-kB by regulating pro- sequence 50-GGGACTTTCC-30 present within teolysis of the IkB inhibitor protein (Fig. -
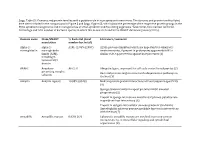
Supp. Table S2: Domains and Protein Families with a Putative Role in Host-Symbiont Interactions
Supp. Table S2: Domains and protein families with a putative role in host-symbiont interactions. The domains and protein families listed here were included in the comparisons in Figure 5 and Supp. Figure S5, which show the percentage of the respective protein groups in the Riftia symbiont metagenome and in metagenomes of other symbiotic and free-living organisms. % bacterial, total number bacterial: Percentage and total number of bacterial species in which this domain is found in the SMART database (January 2019). Domain name Pfam/SMART % bacterial (total Literature/comment annotation number bacterial) Alpha-2- alpha-2- A2M: 42.05% (2057) A2Ms: protease inhibitors which are important for eukaryotic macroglobulin macroglobulin innate immunity, if present in prokaryotes apparently fulfill a family (A2M), similar role, e.g. protection against host proteases (1) including N- terminal MG1 domain ANAPC Anaphase- APC2: 0 Ubiquitin ligase, important for cell cycle control in eukaryotes (2) promoting complex Bacterial proteins might interact with ubiquitination pathways in subunits the host (3) Ankyrin Ankyrin repeats 10.88% (8348) Mediate protein-protein interactions without sequence specificity (4) Sponge symbiont ankyrin-repeat proteins inhibit amoebal phagocytosis (5) Present in sponge microbiome metatranscriptomes, putative role in symbiont-host interactions (6) Present in obligate intracellular amoeba symbiont Candidatus Amoebophilus asiaticus genome, probable function in interactions with the host (7) Armadillo Armadillo repeats 0.83% (67) -

Consensus Tetratricopeptide Repeat Proteins Are Complex Superhelical 2 Nanosprings
bioRxiv preprint doi: https://doi.org/10.1101/2021.03.27.437344; this version posted August 30, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Consensus tetratricopeptide repeat proteins are complex superhelical 2 nanosprings 1, ∗ 1 1 2 2 3 Marie Synakewicz, Rohan S. Eapen, Albert Perez-Riba, Daniela Bauer, Andreas Weißl, 3 3 2 1, ∗ 4, ∗ 4 Gerhard Fischer, Marko Hyv¨onen, Matthias Rief, Laura S. Itzhaki, and Johannes Stigler 1 5 Department of Pharmacology, University of Cambridge, 6 Tennis Court Road, Cambridge, CB2 1PD, United Kingdom 2 7 Physik-Department, Technische Universit¨atM¨unchen, 8 James-Franck-Straße 1, 85748 Garching, Germany 3 9 Department of Biochemistry, University of Cambridge, 10 Tennis Court Road, Cambridge, CB2 1GA, United Kingdom 4 11 Gene Center Munich, Ludwig-Maximilians-Universit¨atM¨unchen, 12 Feodor-Lynen-Straße 25, 81377 M¨unchen,Germany 13 (Dated: August 20, 2021) Abstract. Tandem-repeat proteins comprise small secondary structure motifs that stack to form one- dimensional arrays with distinctive mechanical properties that are proposed to direct their cellular functions. Here, we use single-molecule optical tweezers to study the folding of consensus-designed tetratricopeptide repeats (CTPRs) | superhelical arrays of short helix-turn-helix motifs. We find that CTPRs display a spring-like mechanical response in which individual repeats undergo rapid equilibrium fluctuations between folded and unfolded conformations. We rationalise the force response using Ising models and dissect the folding pathway of CTPRs under mechanical load, revealing how the repeat arrays form from the centre towards both termini simultaneously. -

Small Glutamine-Rich Tetratricopeptide Repeat
www.nature.com/scientificreports OPEN Small Glutamine-Rich Tetratricopeptide Repeat- Containing Protein Alpha (SGTA) Received: 04 January 2016 Accepted: 07 June 2016 Ablation Limits Offspring Viability Published: 30 June 2016 and Growth in Mice Lisa K. Philp1, Tanya K. Day1, Miriam S. Butler1, Geraldine Laven-Law1, Shalini Jindal1, Theresa E. Hickey1, Howard I. Scher2, Lisa M. Butler1,3,* & Wayne D. Tilley1,3,* Small glutamine-rich tetratricopeptide repeat-containing protein α (SGTA) has been implicated as a co-chaperone and regulator of androgen and growth hormone receptor (AR, GHR) signalling. We investigated the functional consequences of partial and full Sgta ablation in vivo using Cre-lox Sgta- null mice. Sgta+/− breeders generated viable Sgta−/− offspring, but at less than Mendelian expectancy. Sgta−/− breeders were subfertile with small litters and higher neonatal death (P < 0.02). Body size was significantly and proportionately smaller in male and femaleSgta −/− (vs WT, Sgta+/− P < 0.001) from d19. Serum IGF-1 levels were genotype- and sex-dependent. Food intake, muscle and bone mass and adiposity were unchanged in Sgta−/−. Vital and sex organs had normal relative weight, morphology and histology, although certain androgen-sensitive measures such as penis and preputial size, and testis descent, were greater in Sgta−/−. Expression of AR and its targets remained largely unchanged, although AR localisation was genotype- and tissue-dependent. Generally expression of other TPR- containing proteins was unchanged. In conclusion, this thorough investigation of SGTA-null mutation reports a mild phenotype of reduced body size. The model’s full potential likely will be realised by genetic crosses with other models to interrogate the role of SGTA in the many diseases in which it has been implicated. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Eukaryotic Genome Annotation
Comparative Features of Multicellular Eukaryotic Genomes (2017) (First three statistics from www.ensembl.org; other from original papers) C. elegans A. thaliana D. melanogaster M. musculus H. sapiens Species name Nematode Thale Cress Fruit Fly Mouse Human Size (Mb) 103 136 143 3,482 3,555 # Protein-coding genes 20,362 27,655 13,918 22,598 20,338 (25,498 (13,601 original (30,000 (30,000 original est.) original est.) original est.) est.) Transcripts 58,941 55,157 34,749 131,195 200,310 Gene density (#/kb) 1/5 1/4.5 1/8.8 1/83 1/97 LINE/SINE (%) 0.4 0.5 0.7 27.4 33.6 LTR (%) 0.0 4.8 1.5 9.9 8.6 DNA Elements 5.3 5.1 0.7 0.9 3.1 Total repeats 6.5 10.5 3.1 38.6 46.4 Exons % genome size 27 28.8 24.0 per gene 4.0 5.4 4.1 8.4 8.7 average size (bp) 250 506 Introns % genome size 15.6 average size (bp) 168 Arabidopsis Chromosome Structures Sorghum Whole Genome Details Characterizing the Proteome The Protein World • Sequencing has defined o Many, many proteins • How can we use this data to: o Define genes in new genomes o Look for evolutionarily related genes o Follow evolution of genes ▪ Mixing of domains to create new proteins o Uncover important subsets of genes that ▪ That deep phylogenies • Plants vs. animals • Placental vs. non-placental animals • Monocots vs. dicots plants • Common nomenclature needed o Ensure consistency of interpretations InterPro (http://www.ebi.ac.uk/interpro/) Classification of Protein Families • Intergrated documentation resource for protein super families, families, domains and functional sites o Mitchell AL, Attwood TK, Babbitt PC, et al.