CDG Therapies: from Bench to Bedside
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

A Proteomic Approach to Uncover Neuroprotective Mechanisms of Oleocanthal Against Oxidative Stress
International Journal of Molecular Sciences Article A Proteomic Approach to Uncover Neuroprotective Mechanisms of Oleocanthal against Oxidative Stress Laura Giusti 1,†, Cristina Angeloni 2,†, Maria Cristina Barbalace 3, Serena Lacerenza 4, Federica Ciregia 5, Maurizio Ronci 6 ID , Andrea Urbani 7, Clementina Manera 4, Maria Digiacomo 4 ID , Marco Macchia 4, Maria Rosa Mazzoni 4, Antonio Lucacchini 1 ID and Silvana Hrelia 3,* ID 1 Department of Clinical and Experimental Medicine, University of Pisa, 56126 Pisa, Italy; [email protected] (L.G.); [email protected] (A.L.) 2 School of Pharmacy, University of Camerino, 62032 Camerino, Italy; [email protected] 3 Department for Life Quality Studies, Alma Mater Studiorum, University of Bologna, 47921 Rimini, Italy; [email protected] 4 Department of Pharmacy, University of Pisa, 56126 Pisa, Italy; [email protected] (S.L.); [email protected] (C.M.); [email protected] (M.D.); [email protected] (M.M.); [email protected] (M.R.M.) 5 Department of Rheumatology, GIGA Research, Centre Hospitalier Universitaire (CHU) de Liège, University of Liège, 4000 Liège, Belgium; [email protected] 6 Department of Medical, Oral and Biotechnological Sciences, University G. d’Annunzio of Chieti-Pescara, 65127 Pescara, Italy; [email protected] 7 Institute of Biochemistry and Clinical Biochemistry, Catholic University, 00198 Rome, Italy; [email protected] * Correspondence: [email protected]; Tel.: +39-051-209-1235 † These authors contributed equally to this work. Received: 3 July 2018; Accepted: 1 August 2018; Published: 8 August 2018 Abstract: Neurodegenerative diseases represent a heterogeneous group of disorders that share common features like abnormal protein aggregation, perturbed Ca2+ homeostasis, excitotoxicity, impairment of mitochondrial functions, apoptosis, inflammation, and oxidative stress. -

Congenital Disorders of Glycosylation from a Neurological Perspective
brain sciences Review Congenital Disorders of Glycosylation from a Neurological Perspective Justyna Paprocka 1,* , Aleksandra Jezela-Stanek 2 , Anna Tylki-Szyma´nska 3 and Stephanie Grunewald 4 1 Department of Pediatric Neurology, Faculty of Medical Science in Katowice, Medical University of Silesia, 40-752 Katowice, Poland 2 Department of Genetics and Clinical Immunology, National Institute of Tuberculosis and Lung Diseases, 01-138 Warsaw, Poland; [email protected] 3 Department of Pediatrics, Nutrition and Metabolic Diseases, The Children’s Memorial Health Institute, W 04-730 Warsaw, Poland; [email protected] 4 NIHR Biomedical Research Center (BRC), Metabolic Unit, Great Ormond Street Hospital and Institute of Child Health, University College London, London SE1 9RT, UK; [email protected] * Correspondence: [email protected]; Tel.: +48-606-415-888 Abstract: Most plasma proteins, cell membrane proteins and other proteins are glycoproteins with sugar chains attached to the polypeptide-glycans. Glycosylation is the main element of the post- translational transformation of most human proteins. Since glycosylation processes are necessary for many different biological processes, patients present a diverse spectrum of phenotypes and severity of symptoms. The most frequently observed neurological symptoms in congenital disorders of glycosylation (CDG) are: epilepsy, intellectual disability, myopathies, neuropathies and stroke-like episodes. Epilepsy is seen in many CDG subtypes and particularly present in the case of mutations -

NICU Gene List Generator.Xlsx
Neonatal Crisis Sequencing Panel Gene List Genes: A2ML1 - B3GLCT A2ML1 ADAMTS9 ALG1 ARHGEF15 AAAS ADAMTSL2 ALG11 ARHGEF9 AARS1 ADAR ALG12 ARID1A AARS2 ADARB1 ALG13 ARID1B ABAT ADCY6 ALG14 ARID2 ABCA12 ADD3 ALG2 ARL13B ABCA3 ADGRG1 ALG3 ARL6 ABCA4 ADGRV1 ALG6 ARMC9 ABCB11 ADK ALG8 ARPC1B ABCB4 ADNP ALG9 ARSA ABCC6 ADPRS ALK ARSL ABCC8 ADSL ALMS1 ARX ABCC9 AEBP1 ALOX12B ASAH1 ABCD1 AFF3 ALOXE3 ASCC1 ABCD3 AFF4 ALPK3 ASH1L ABCD4 AFG3L2 ALPL ASL ABHD5 AGA ALS2 ASNS ACAD8 AGK ALX3 ASPA ACAD9 AGL ALX4 ASPM ACADM AGPS AMELX ASS1 ACADS AGRN AMER1 ASXL1 ACADSB AGT AMH ASXL3 ACADVL AGTPBP1 AMHR2 ATAD1 ACAN AGTR1 AMN ATL1 ACAT1 AGXT AMPD2 ATM ACE AHCY AMT ATP1A1 ACO2 AHDC1 ANK1 ATP1A2 ACOX1 AHI1 ANK2 ATP1A3 ACP5 AIFM1 ANKH ATP2A1 ACSF3 AIMP1 ANKLE2 ATP5F1A ACTA1 AIMP2 ANKRD11 ATP5F1D ACTA2 AIRE ANKRD26 ATP5F1E ACTB AKAP9 ANTXR2 ATP6V0A2 ACTC1 AKR1D1 AP1S2 ATP6V1B1 ACTG1 AKT2 AP2S1 ATP7A ACTG2 AKT3 AP3B1 ATP8A2 ACTL6B ALAS2 AP3B2 ATP8B1 ACTN1 ALB AP4B1 ATPAF2 ACTN2 ALDH18A1 AP4M1 ATR ACTN4 ALDH1A3 AP4S1 ATRX ACVR1 ALDH3A2 APC AUH ACVRL1 ALDH4A1 APTX AVPR2 ACY1 ALDH5A1 AR B3GALNT2 ADA ALDH6A1 ARFGEF2 B3GALT6 ADAMTS13 ALDH7A1 ARG1 B3GAT3 ADAMTS2 ALDOB ARHGAP31 B3GLCT Updated: 03/15/2021; v.3.6 1 Neonatal Crisis Sequencing Panel Gene List Genes: B4GALT1 - COL11A2 B4GALT1 C1QBP CD3G CHKB B4GALT7 C3 CD40LG CHMP1A B4GAT1 CA2 CD59 CHRNA1 B9D1 CA5A CD70 CHRNB1 B9D2 CACNA1A CD96 CHRND BAAT CACNA1C CDAN1 CHRNE BBIP1 CACNA1D CDC42 CHRNG BBS1 CACNA1E CDH1 CHST14 BBS10 CACNA1F CDH2 CHST3 BBS12 CACNA1G CDK10 CHUK BBS2 CACNA2D2 CDK13 CILK1 BBS4 CACNB2 CDK5RAP2 -
Technical Note, Appendix: an Analysis of Blood Processing Methods to Prepare Samples for Genechip® Expression Profiling (Pdf, 1
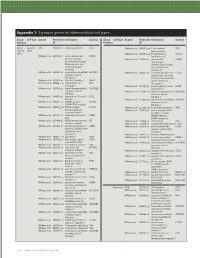
Appendix 1: Signature genes for different blood cell types. Blood Cell Type Source Probe Set Description Symbol Blood Cell Type Source Probe Set Description Symbol Fraction ID Fraction ID Mono- Lympho- GSK 203547_at CD4 antigen (p55) CD4 Whitney et al. 209813_x_at T cell receptor TRG nuclear cytes gamma locus cells Whitney et al. 209995_s_at T-cell leukemia/ TCL1A Whitney et al. 203104_at colony stimulating CSF1R lymphoma 1A factor 1 receptor, Whitney et al. 210164_at granzyme B GZMB formerly McDonough (granzyme 2, feline sarcoma viral cytotoxic T-lymphocyte- (v-fms) oncogene associated serine homolog esterase 1) Whitney et al. 203290_at major histocompatibility HLA-DQA1 Whitney et al. 210321_at similar to granzyme B CTLA1 complex, class II, (granzyme 2, cytotoxic DQ alpha 1 T-lymphocyte-associated Whitney et al. 203413_at NEL-like 2 (chicken) NELL2 serine esterase 1) Whitney et al. 203828_s_at natural killer cell NK4 (H. sapiens) transcript 4 Whitney et al. 212827_at immunoglobulin heavy IGHM Whitney et al. 203932_at major histocompatibility HLA-DMB constant mu complex, class II, Whitney et al. 212998_x_at major histocompatibility HLA-DQB1 DM beta complex, class II, Whitney et al. 204655_at chemokine (C-C motif) CCL5 DQ beta 1 ligand 5 Whitney et al. 212999_x_at major histocompatibility HLA-DQB Whitney et al. 204661_at CDW52 antigen CDW52 complex, class II, (CAMPATH-1 antigen) DQ beta 1 Whitney et al. 205049_s_at CD79A antigen CD79A Whitney et al. 213193_x_at T cell receptor beta locus TRB (immunoglobulin- Whitney et al. 213425_at Homo sapiens cDNA associated alpha) FLJ11441 fis, clone Whitney et al. 205291_at interleukin 2 receptor, IL2RB HEMBA1001323, beta mRNA sequence Whitney et al. -

Human Metabolome Technologies
Human Metabolome Human Metabolome Technologies Inc. (HMT) is a leading metabolomics service provider company established on July 2003 based on capillary electrophoresis mass spectrometry (CE-MS) technologies. Technologies The company was listed on the Mothers section of Tokyo Stock Exchange in December 2013. Our main Commissioned metabolome analysis services business is commissioned metabolomics analysis using capillary electrophoresis time-of-flight mass spectrometry (CE-TOFMS): We have a time-tested track record in a number of different fields including medical sciences, pharmaceuticals, food products, fermentation, and cosmetics. With an aim of contributing in a wide range of fields, we will now set our sights on untapped areas such as the environment, energy, and chemical industries. History Jul 2003 Founded in Suehiromachi in Tsuruoka, Yamagata Prefecture with capital of 10 million yen. Jun 2004 Concluded a joint research agreement with Ajinomoto Co., Inc. May 2009 Commenced the “HMT Research Grant for Young Leaders” Aug 2012 Launched a cancer research specialized package, “C-SCOPE.” Oct 2012 Established a sales subsidiary, “Human Metabolome Technologies America, Inc.” in Massachusetts, USA Sep 2013 Registered the patent “The biomarker for depression, the measuring method for the biomarker of depression, and the program and storage for the diagnostic method” (patent number 5372213) in Japan Dec 2013 Listed on the Mothers section of the Tokyo Stock Exchange Jan 2016 Established a biomarker business company “HMT Biomedical Co., Ltd.” in Yokohama, Kanagawa, Japan May 2017 Established a sales subsidiary, “Human Metabolome Technologies Europe B.V.” in Leiden, Netherlands Apr 2018 Launched functional lipidomics specialized package “Mediator Scan” Human Metabolome Technologies Inc. -

PIGA Gene Phosphatidylinositol Glycan Anchor Biosynthesis Class A
PIGA gene phosphatidylinositol glycan anchor biosynthesis class A Normal Function The PIGA gene provides instructions for making a protein called phosphatidylinositol glycan class A. This protein takes part in a series of steps that produce a molecule called GPI anchor. Specifically, phosphatidylinositol glycan class A is involved in the first step of the sequence, which produces an intermediate molecule called N- acetylglucosaminyl phosphatidylinositol, or GlcNAc-PI. This step takes place in the endoplasmic reticulum of the cell, a structure involved in protein processing and transport. The PIGA protein forms a complex with several other proteins, and this complex helps to start the reaction that produces GlcNAc-PI. The GPI anchor, the ultimate product of the sequence, attaches many different proteins to the cell membrane, thereby ensuring that these proteins are available when needed at the surface of the cell. Health Conditions Related to Genetic Changes Paroxysmal nocturnal hemoglobinuria Some gene mutations are acquired during a person's lifetime and are present only in certain cells. These changes, which are called somatic mutations, are not inherited. In people with paroxysmal nocturnal hemoglobinuria, somatic mutations of the PIGA gene occur in blood-forming cells called hematopoietic stem cells. Hematopoietic stem cells produce red blood cells (erythrocytes) that carry oxygen, white blood cells (leukocytes) that protect the body from infection, and platelets (thrombocytes) that are involved in blood clotting. Individuals with paroxysmal nocturnal hemoglobinuria have one or more PIGA gene mutations in their hematopoietic stem cells, which leads to abnormal blood cells. As the abnormal hematopoietic stem cells multiply, populations of abnormal blood cells are formed, alongside normal blood cells produced by normal hematopoietic stem cells. -

Deoxyadenosine Triphosphate As a Mediator of Deoxyguanosine Toxicity in Cultured T Lymphoblasts
Deoxyadenosine triphosphate as a mediator of deoxyguanosine toxicity in cultured T lymphoblasts. G J Mann, R M Fox J Clin Invest. 1986;78(5):1261-1269. https://doi.org/10.1172/JCI112710. Research Article The mechanism by which 2'-deoxyguanosine is toxic for lymphoid cells is relevant both to the severe cellular immune defect of inherited purine nucleoside phosphorylase (PNP) deficiency and to attempts to exploit PNP inhibitors therapeutically. We have studied the cell cycle and biochemical effects of 2'-deoxyguanosine in human lymphoblasts using the PNP inhibitor 8-aminoguanosine. We show that cytostatic 2'-deoxyguanosine concentrations cause G1-phase arrest in PNP-inhibited T lymphoblasts, regardless of their hypoxanthine guanine phosphoribosyltransferase status. This effect is identical to that produced by 2'-deoxyadenosine in adenosine deaminase-inhibited T cells. 2'-Deoxyguanosine elevates both the 2'-deoxyguanosine-5'-triphosphate (dGTP) and 2'-deoxyadenosine-5'-triphosphate (dATP) pools; subsequently pyrimidine deoxyribonucleotide pools are depleted. The time course of these biochemical changes indicates that the onset of G1-phase arrest is related to increase of the dATP rather than the dGTP pool. When dGTP elevation is dissociated from dATP elevation by coincubation with 2'-deoxycytidine, dGTP does not by itself interrupt transit from the G1 to the S phase. It is proposed that dATP can mediate both 2'-deoxyguanosine and 2'-deoxyadenosine toxicity in T lymphoblasts. Find the latest version: https://jci.me/112710/pdf Deoxyadenosine Triphosphate as a Mediator of Deoxyguanosine Toxicity in Cultured T Lymphoblasts G. J. Mann and R. M. Fox Ludwig Institute for Cancer Research (Sydney Branch), University ofSydney, Sydney, New South Wales 2006, Australia Abstract urine of PNP-deficient individuals, with elevation of plasma inosine and guanosine and mild hypouricemia (3). -

Biological Chemistry Department
MINISTRY OF HEALTH OF UKRAINE ZAPORIZHZHIA STATE MEDICAL UNIVERSITY Biological Chemistry Department Biological chemistry A manual for independent work at home and in class preparation for licensing examination “KROK 1” on semantic modules 6, 7 of module 2 for students of International Faculty (the second year of study) Zaporizhzhia 2017 UDC 577.1(075) BBC 28.902я73 B60 Ratified on meeting of the Central methodical committee of Zaporozhye State Medical University (protocol N 3 from 02_03_17) and it is recommended for the use in educational process for foreign students. Reviewers: Prihodko O. B., Head of Department of Medical Biology, Parasitology and Genetics. Dr. Hab, assoc. professor; Belenichev I. F., Head of Department of Pharmacology and Medicinal Preparations, Dr. Hab., professor Authors: Aleksandrova K. V., Romanenko M. I., Krisanova N. V., Ivanchenko D. G., Rudko N. P., Levich S. V. Biological chemistry : a manual for independent work at home and in class preparation for licensing examination "KROK 1" on semantic modules 6, 7 of module 2 for students of International Faculty (the second year of study) / K. V. Aleksandrova, М. І. Romanenko, N. V. Krisanova, D. G. Ivanchenko, N. P. Rudko, S. V. Levich. – Zaporizhzhia : ZSMU, 2017. – 213 p. This manual is recommended for II year students of International Faculty of specialty "General medicine" studying biological chemistry, as additional material to prepare for practical training semantic modules 6, 7 of module 2 and licensing exam "KROK 1: General medical training". Біологічна хімія : навч.-метод. посіб. для самостійної роботи при підготовці до ліцензійного іспиту "КРОК 1" змістових модулів 6, 7 модулю 2 для студентів 2 курсу міжнар. -

Epigenetic Mechanisms Are Involved in the Oncogenic Properties of ZNF518B in Colorectal Cancer
Epigenetic mechanisms are involved in the oncogenic properties of ZNF518B in colorectal cancer Francisco Gimeno-Valiente, Ángela L. Riffo-Campos, Luis Torres, Noelia Tarazona, Valentina Gambardella, Andrés Cervantes, Gerardo López-Rodas, Luis Franco and Josefa Castillo SUPPLEMENTARY METHODS 1. Selection of genomic sequences for ChIP analysis To select the sequences for ChIP analysis in the five putative target genes, namely, PADI3, ZDHHC2, RGS4, EFNA5 and KAT2B, the genomic region corresponding to the gene was downloaded from Ensembl. Then, zoom was applied to see in detail the promoter, enhancers and regulatory sequences. The details for HCT116 cells were then recovered and the target sequences for factor binding examined. Obviously, there are not data for ZNF518B, but special attention was paid to the target sequences of other zinc-finger containing factors. Finally, the regions that may putatively bind ZNF518B were selected and primers defining amplicons spanning such sequences were searched out. Supplementary Figure S3 gives the location of the amplicons used in each gene. 2. Obtaining the raw data and generating the BAM files for in silico analysis of the effects of EHMT2 and EZH2 silencing The data of siEZH2 (SRR6384524), siG9a (SRR6384526) and siNon-target (SRR6384521) in HCT116 cell line, were downloaded from SRA (Bioproject PRJNA422822, https://www.ncbi. nlm.nih.gov/bioproject/), using SRA-tolkit (https://ncbi.github.io/sra-tools/). All data correspond to RNAseq single end. doBasics = TRUE doAll = FALSE $ fastq-dump -I --split-files SRR6384524 Data quality was checked using the software fastqc (https://www.bioinformatics.babraham. ac.uk /projects/fastqc/). The first low quality removing nucleotides were removed using FASTX- Toolkit (http://hannonlab.cshl.edu/fastxtoolkit/). -

Whole-Exome Sequencing Identifies Causative Mutations in Families
BASIC RESEARCH www.jasn.org Whole-Exome Sequencing Identifies Causative Mutations in Families with Congenital Anomalies of the Kidney and Urinary Tract Amelie T. van der Ven,1 Dervla M. Connaughton,1 Hadas Ityel,1 Nina Mann,1 Makiko Nakayama,1 Jing Chen,1 Asaf Vivante,1 Daw-yang Hwang,1 Julian Schulz,1 Daniela A. Braun,1 Johanna Magdalena Schmidt,1 David Schapiro,1 Ronen Schneider,1 Jillian K. Warejko,1 Ankana Daga,1 Amar J. Majmundar,1 Weizhen Tan,1 Tilman Jobst-Schwan,1 Tobias Hermle,1 Eugen Widmeier,1 Shazia Ashraf,1 Ali Amar,1 Charlotte A. Hoogstraaten,1 Hannah Hugo,1 Thomas M. Kitzler,1 Franziska Kause,1 Caroline M. Kolvenbach,1 Rufeng Dai,1 Leslie Spaneas,1 Kassaundra Amann,1 Deborah R. Stein,1 Michelle A. Baum,1 Michael J.G. Somers,1 Nancy M. Rodig,1 Michael A. Ferguson,1 Avram Z. Traum,1 Ghaleb H. Daouk,1 Radovan Bogdanovic,2 Natasa Stajic,2 Neveen A. Soliman,3,4 Jameela A. Kari,5,6 Sherif El Desoky,5,6 Hanan M. Fathy,7 Danko Milosevic,8 Muna Al-Saffar,1,9 Hazem S. Awad,10 Loai A. Eid,10 Aravind Selvin,11 Prabha Senguttuvan,12 Simone Sanna-Cherchi,13 Heidi L. Rehm,14 Daniel G. MacArthur,14,15 Monkol Lek,14,15 Kristen M. Laricchia,15 Michael W. Wilson,15 Shrikant M. Mane,16 Richard P. Lifton,16,17 Richard S. Lee,18 Stuart B. Bauer,18 Weining Lu,19 Heiko M. Reutter ,20,21 Velibor Tasic,22 Shirlee Shril,1 and Friedhelm Hildebrandt1 Due to the number of contributing authors, the affiliations are listed at the end of this article. -

Glycomic and Transcriptomic Response of GSC11 Glioblastoma Stem Cells to STAT3 Phosphorylation Inhibition and Serum- Induced Differentiation
See discussions, stats, and author profiles for this publication at: https://www.researchgate.net/publication/41720955 Glycomic and Transcriptomic Response of GSC11 Glioblastoma Stem Cells to STAT3 Phosphorylation Inhibition and Serum- Induced Differentiation Article in Journal of Proteome Research · March 2010 Impact Factor: 4.25 · DOI: 10.1021/pr900793a · Source: PubMed CITATIONS READS 21 107 11 authors, including: Yongjie Ji Waldemar Priebe University of Texas MD Anderson Cancer C… University of Texas MD Anderson Cancer C… 14 PUBLICATIONS 550 CITATIONS 275 PUBLICATIONS 6,260 CITATIONS SEE PROFILE SEE PROFILE Frederick F Lang Charles A Conrad University of Texas MD Anderson Cancer C… University of Texas MD Anderson Cancer C… 254 PUBLICATIONS 10,474 CITATIONS 84 PUBLICATIONS 2,169 CITATIONS SEE PROFILE SEE PROFILE Available from: Frederick F Lang Retrieved on: 26 May 2016 Glycomic and Transcriptomic Response of GSC11 Glioblastoma Stem Cells to STAT3 Phosphorylation Inhibition and Serum-Induced Differentiation Huan He,†,‡ Carol L. Nilsson,*,†,# Mark R. Emmett,†,‡ Alan G. Marshall,†,‡ Roger A. Kroes,§ Joseph R. Moskal,§ Yongjie Ji,| Howard Colman,| Waldemar Priebe,⊥ Frederick F. Lang,| and Charles A. Conrad| Ion Cyclotron Resonance Program, National High Magnetic Field Laboratory, Florida State University, Tallahassee, Florida 32310, Department of Chemistry and Biochemistry, Florida State University, Tallahassee, Florida 32306-43903, Falk Center for Molecular Therapeutics, Department of Biomedical Engineering, Northwestern University, Evanston, Illinois 60201, Department of Neuro-oncology, The University of Texas M.D. Anderson Cancer Center, Houston, Texas 77030, and Department of Experimental Therapeutics, The University of Texas M.D. Anderson Cancer Center, Houston, Texas 77030 Received September 05, 2009 A glioblastoma stem cell (GSC) line, GSC11, grows as neurospheres in serum-free media supplemented with EGF (epidermal growth factor) and bFGF (basic fibroblast growth factor), and, if implanted in nude mice brains, will recapitulate high-grade glial tumors.