KIF3B (NM 004798) Human Recombinant Protein Product Data
Total Page:16
File Type:pdf, Size:1020Kb

Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Transcriptomic and Proteomic Profiling Provides Insight Into
BASIC RESEARCH www.jasn.org Transcriptomic and Proteomic Profiling Provides Insight into Mesangial Cell Function in IgA Nephropathy † † ‡ Peidi Liu,* Emelie Lassén,* Viji Nair, Celine C. Berthier, Miyuki Suguro, Carina Sihlbom,§ † | † Matthias Kretzler, Christer Betsholtz, ¶ Börje Haraldsson,* Wenjun Ju, Kerstin Ebefors,* and Jenny Nyström* *Department of Physiology, Institute of Neuroscience and Physiology, §Proteomics Core Facility at University of Gothenburg, University of Gothenburg, Gothenburg, Sweden; †Division of Nephrology, Department of Internal Medicine and Department of Computational Medicine and Bioinformatics, University of Michigan, Ann Arbor, Michigan; ‡Division of Molecular Medicine, Aichi Cancer Center Research Institute, Nagoya, Japan; |Department of Immunology, Genetics and Pathology, Uppsala University, Uppsala, Sweden; and ¶Integrated Cardio Metabolic Centre, Karolinska Institutet Novum, Huddinge, Sweden ABSTRACT IgA nephropathy (IgAN), the most common GN worldwide, is characterized by circulating galactose-deficient IgA (gd-IgA) that forms immune complexes. The immune complexes are deposited in the glomerular mesangium, leading to inflammation and loss of renal function, but the complete pathophysiology of the disease is not understood. Using an integrated global transcriptomic and proteomic profiling approach, we investigated the role of the mesangium in the onset and progression of IgAN. Global gene expression was investigated by microarray analysis of the glomerular compartment of renal biopsy specimens from patients with IgAN (n=19) and controls (n=22). Using curated glomerular cell type–specific genes from the published literature, we found differential expression of a much higher percentage of mesangial cell–positive standard genes than podocyte-positive standard genes in IgAN. Principal coordinate analysis of expression data revealed clear separation of patient and control samples on the basis of mesangial but not podocyte cell–positive standard genes. -

Mitochondria Are Transported Along Microtubules in Membrane
Shen et al. Cell Death and Disease (2018) 9:81 DOI 10.1038/s41419-017-0145-x Cell Death & Disease ARTICLE Open Access Mitochondria are transported along microtubules in membrane nanotubes to rescue distressed cardiomyocytes from apoptosis Jing Shen1,2,3,4, Jiang-Hui Zhang1,2,3,4,HanXiao1,2,3,4,Ji-MinWu1,2,3,4,Kang-MinHe1,2,3,4,Zhi-ZhenLv1,2,3,4, Zi-Jian Li1,2,3,4, Ming Xu1,2,3,4 and You-Yi Zhang1,2,3,4 Abstract Membrane nanotubes (MNTs) act as “highways” between cells to facilitate the transfer of multiple signals and play an important role in many diseases. Our previous work reported on the transfer of mitochondria via MNTs between cardiomyocytes (CMs) and cardiac myofibroblasts (MFs); however, the elucidation of the underlying mechanism and pathophysiological significance of this transfer requires additional study. In this study, we determined that the mean movement velocity of mitochondria in MNTs between CMs and MFs was approximately 17.5 ± 2.1 nm/s. Meanwhile, treatment with microtubule polymerisation inhibitors nocodazole or colcemid in cell culture decreased mitochondrial velocity, and knockdown of the microtubule motor protein kinesin family member 5B (KIF5B) led to a similar effect, indicating that mitochondrial movement was dependent on microtubules and the motor protein KIF5B. Furthermore, we showed that hypoxia/reoxygenation-induced CM 1234567890 1234567890 apoptosis was attenuated by coculture with intact or hypoxia/reoxygenation-treated MFs, which transferred mitochondria to CMs. This rescue was prevented either by separating the cells using Transwell culture or by impairing mitochondrial transfer with nocodazole or colcemid treatment. -

Supplementary Table 1 Genes Tested in Qrt-PCR in Nfpas
Supplementary Table 1 Genes tested in qRT-PCR in NFPAs Gene Bank accession Gene Description number ABI assay ID a disintegrin-like and metalloprotease with thrombospondin type 1 motif 7 ADAMTS7 NM_014272.3 Hs00276223_m1 Rho guanine nucleotide exchange factor (GEF) 3 ARHGEF3 NM_019555.1 Hs00219609_m1 BCL2-associated X protein BAX NM_004324 House design Bcl-2 binding component 3 BBC3 NM_014417.2 Hs00248075_m1 B-cell CLL/lymphoma 2 BCL2 NM_000633 House design Bone morphogenetic protein 7 BMP7 NM_001719.1 Hs00233476_m1 CCAAT/enhancer binding protein (C/EBP), alpha CEBPA NM_004364.2 Hs00269972_s1 coxsackie virus and adenovirus receptor CXADR NM_001338.3 Hs00154661_m1 Homo sapiens Dicer1, Dcr-1 homolog (Drosophila) (DICER1) DICER1 NM_177438.1 Hs00229023_m1 Homo sapiens dystonin DST NM_015548.2 Hs00156137_m1 fms-related tyrosine kinase 3 FLT3 NM_004119.1 Hs00174690_m1 glutamate receptor, ionotropic, N-methyl D-aspartate 1 GRIN1 NM_000832.4 Hs00609557_m1 high-mobility group box 1 HMGB1 NM_002128.3 Hs01923466_g1 heterogeneous nuclear ribonucleoprotein U HNRPU NM_004501.3 Hs00244919_m1 insulin-like growth factor binding protein 5 IGFBP5 NM_000599.2 Hs00181213_m1 latent transforming growth factor beta binding protein 4 LTBP4 NM_001042544.1 Hs00186025_m1 microtubule-associated protein 1 light chain 3 beta MAP1LC3B NM_022818.3 Hs00797944_s1 matrix metallopeptidase 17 MMP17 NM_016155.4 Hs01108847_m1 myosin VA MYO5A NM_000259.1 Hs00165309_m1 Homo sapiens nuclear factor (erythroid-derived 2)-like 1 NFE2L1 NM_003204.1 Hs00231457_m1 oxoglutarate (alpha-ketoglutarate) -
Drosophila and Human Transcriptomic Data Mining Provides Evidence for Therapeutic
Drosophila and human transcriptomic data mining provides evidence for therapeutic mechanism of pentylenetetrazole in Down syndrome Author Abhay Sharma Institute of Genomics and Integrative Biology Council of Scientific and Industrial Research Delhi University Campus, Mall Road Delhi 110007, India Tel: +91-11-27666156, Fax: +91-11-27662407 Email: [email protected] Nature Precedings : hdl:10101/npre.2010.4330.1 Posted 5 Apr 2010 Running head: Pentylenetetrazole mechanism in Down syndrome 1 Abstract Pentylenetetrazole (PTZ) has recently been found to ameliorate cognitive impairment in rodent models of Down syndrome (DS). The mechanism underlying PTZ’s therapeutic effect is however not clear. Microarray profiling has previously reported differential expression of genes in DS. No mammalian transcriptomic data on PTZ treatment however exists. Nevertheless, a Drosophila model inspired by rodent models of PTZ induced kindling plasticity has recently been described. Microarray profiling has shown PTZ’s downregulatory effect on gene expression in fly heads. In a comparative transcriptomics approach, I have analyzed the available microarray data in order to identify potential mechanism of PTZ action in DS. I find that transcriptomic correlates of chronic PTZ in Drosophila and DS counteract each other. A significant enrichment is observed between PTZ downregulated and DS upregulated genes, and a significant depletion between PTZ downregulated and DS dowwnregulated genes. Further, the common genes in PTZ Nature Precedings : hdl:10101/npre.2010.4330.1 Posted 5 Apr 2010 downregulated and DS upregulated sets show enrichment for MAP kinase pathway. My analysis suggests that downregulation of MAP kinase pathway may mediate therapeutic effect of PTZ in DS. Existing evidence implicating MAP kinase pathway in DS supports this observation. -

The Role of Kinesin-2 in Navigating Microtubule Obstacles: Implications for the Regulation of Axonal Transport Gregory Hoeprich University of Vermont
University of Vermont ScholarWorks @ UVM Graduate College Dissertations and Theses Dissertations and Theses 2016 The Role Of Kinesin-2 In Navigating Microtubule Obstacles: Implications For The Regulation Of Axonal Transport Gregory Hoeprich University of Vermont Follow this and additional works at: https://scholarworks.uvm.edu/graddis Part of the Biophysics Commons Recommended Citation Hoeprich, Gregory, "The Role Of Kinesin-2 In Navigating Microtubule Obstacles: Implications For The Regulation Of Axonal Transport" (2016). Graduate College Dissertations and Theses. 558. https://scholarworks.uvm.edu/graddis/558 This Dissertation is brought to you for free and open access by the Dissertations and Theses at ScholarWorks @ UVM. It has been accepted for inclusion in Graduate College Dissertations and Theses by an authorized administrator of ScholarWorks @ UVM. For more information, please contact [email protected]. THE ROLE OF KINESIN-2 IN NAVIGATING MICROTUBULE OBSTACLES: IMPLICATIONS FOR THE REGULATION OF AXONAL TRANSPORT A Dissertation Presented by Gregory Joseph Hoeprich to The Faculty of the Graduate College of The University of Vermont In Partial Fulfilment of the Requirements For the Degree of Doctor of Philosophy Specializing in Molecular Physiology and Biophysics May, 2016 Defense Date: March 8, 2016 Dissertation Examination Committee: Christopher Berger, Ph.D., Advisor Victor May, Ph.D., Chairperson David Warshaw, Ph.D. Teresa Ruiz, Ph.D. Jason Stumpff, Ph.D. Cynthia J. Forehand, Ph.D., Dean of the Graduate College ABSTRACT Neurons are specialized cells that transmit information through electrical and chemical signals using structural processes known as dendrites and axons. Dendrites receive information for the cell to interpret while the exceedingly long axon transmits the processed information to its target destination. -

Download 20190410); Fragmentation for 20 S
ARTICLE https://doi.org/10.1038/s41467-020-17387-y OPEN Multi-layered proteomic analyses decode compositional and functional effects of cancer mutations on kinase complexes ✉ Martin Mehnert 1 , Rodolfo Ciuffa1, Fabian Frommelt 1, Federico Uliana1, Audrey van Drogen1, ✉ ✉ Kilian Ruminski1,3, Matthias Gstaiger1 & Ruedi Aebersold 1,2 fi 1234567890():,; Rapidly increasing availability of genomic data and ensuing identi cation of disease asso- ciated mutations allows for an unbiased insight into genetic drivers of disease development. However, determination of molecular mechanisms by which individual genomic changes affect biochemical processes remains a major challenge. Here, we develop a multilayered proteomic workflow to explore how genetic lesions modulate the proteome and are trans- lated into molecular phenotypes. Using this workflow we determine how expression of a panel of disease-associated mutations in the Dyrk2 protein kinase alter the composition, topology and activity of this kinase complex as well as the phosphoproteomic state of the cell. The data show that altered protein-protein interactions caused by the mutations are asso- ciated with topological changes and affected phosphorylation of known cancer driver pro- teins, thus linking Dyrk2 mutations with cancer-related biochemical processes. Overall, we discover multiple mutation-specific functionally relevant changes, thus highlighting the extensive plasticity of molecular responses to genetic lesions. 1 Department of Biology, Institute of Molecular Systems Biology, ETH Zurich, -
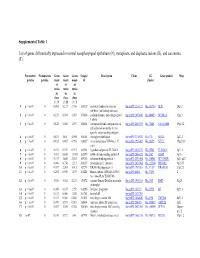
Supplemental Table 1 List of Genes Differentially Expressed In
Supplemental Table 1 List of genes differentially expressed in normal nasopharyngeal epithelium (N), metaplastic and displastic lesions (R), and carcinoma (T). Parametric Permutation Geom Geom Geom Unique Description Clone UG Gene symbol Map p-value p-value mean mean mean id cluster of of of ratios ratios ratios in in in class class class 1 : N 2 : R 3 : T 1 p < 1e-07 0 0.061 0.123 2.708 169329 secretory leukocyte protease IncytePD:2510171 Hs.251754 SLPI 20q12 inhibitor (antileukoproteinase) 2 p < 1e-07 0 0.125 0.394 1.863 163628 sodium channel, nonvoltage-gated IncytePD:1453049 Hs.446415 SCNN1A 12p13 1 alpha 3 p < 1e-07 0 0.122 0.046 1.497 160401 carcinoembryonic antigen-related IncytePD:2060355 Hs.73848 CEACAM6 19q13.2 cell adhesion molecule 6 (non- specific cross reacting antigen) 4 p < 1e-07 0 0.675 1.64 5.594 165101 monoglyceride lipase IncytePD:2174920 Hs.6721 MGLL 3q21.3 5 p < 1e-07 0 0.182 0.487 0.998 166827 nei endonuclease VIII-like 1 (E. IncytePD:1926409 Hs.28355 NEIL1 15q22.33 coli) 6 p < 1e-07 0 0.194 0.339 0.915 162931 hypothetical protein FLJ22418 IncytePD:2816379 Hs.36563 FLJ22418 1p11.1 7 p < 1e-07 0 1.313 0.645 13.593 162399 S100 calcium binding protein P IncytePD:2060823 Hs.2962 S100P 4p16 8 p < 1e-07 0 0.157 1.445 2.563 169315 selenium binding protein 1 IncytePD:2591494 Hs.334841 SELENBP1 1q21-q22 9 p < 1e-07 0 0.046 0.738 1.213 160115 prominin-like 1 (mouse) IncytePD:2070568 Hs.112360 PROML1 4p15.33 10 p < 1e-07 0 0.787 2.264 3.013 167294 HRAS-like suppressor 3 IncytePD:1969263 Hs.37189 HRASLS3 11q12.3 11 p < 1e-07 0 0.292 0.539 1.493 168221 Homo sapiens cDNA FLJ13510 IncytePD:64451 Hs.37896 2 fis, clone PLACE1005146. -

KIF3B Antibody (N-Term) Blocking Peptide Synthetic Peptide Catalog # Bp17242a
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 KIF3B Antibody (N-term) Blocking Peptide Synthetic peptide Catalog # BP17242a Specification KIF3B Antibody (N-term) Blocking Peptide KIF3B Antibody (N-term) Blocking Peptide - - Background Product Information The protein encoded by this gene acts as a Primary Accession O15066 heterodimerwith kinesin family member 3A to aid in chromosome movement duringmitosis and meiosis. The encoded protein is a plus KIF3B Antibody (N-term) Blocking Peptide - Additional Information end-directedmicrotubule motor and can interact with the SMC3 subunit of thecohesin complex. In addition, the encoded protein may Gene ID 9371 be involvedin the intracellular movement of membranous organelles. Thisprotein and Other Names kinesin family member 3A form the kinesin II Kinesin-like protein KIF3B, HH0048, subfamilyof the kinesin superfamily. Microtubule plus end-directed kinesin motor 3B, Kinesin-like protein KIF3B, N-terminally KIF3B Antibody (N-term) Blocking Peptide processed, KIF3B, KIAA0359 - References Format Peptides are lyophilized in a solid powder Reed, A.A., et al. Am. J. Physiol. Renal Physiol. format. Peptides can be reconstituted in 298 (2), F365-F380 (2010) :Keil, R., et al. J. solution using the appropriate buffer as Cell. Sci. 122 (PT 8), 1174-1183 (2009) needed. :Schonteich, E., et al. J. Cell. Sci. 121 (PT 22), 3824-3833 (2008) :Wu, Y., et al. Hum. Mol. Storage Genet. 15(22):3280-3292(2006)Haraguchi, K., Maintain refrigerated at 2-8°C for up to 6 et al. J. Biol. Chem. 281(7):4094-4099(2006) months. For long term storage store at -20°C. -

Kif26b, a Kinesin Family Gene, Regulates Adhesion of the Embryonic Kidney Mesenchyme
Kif26b, a kinesin family gene, regulates adhesion of the embryonic kidney mesenchyme Yukako Uchiyamaa,b,1, Masaji Sakaguchia,b,c,1, Takeshi Terabayashia,b, Toshiaki Inenagaa, Shuji Inouea,b, Chiyoko Kobayashia, Naoko Oshimad, Hiroshi Kiyonarid, Naomi Nakagatae, Yuya Satof, Kiyotoshi Sekiguchif, Hiroaki Mikig, Eiichi Arakic, Sayoko Fujimuraa, Satomi S. Tanakaa, and Ryuichi Nishinakamuraa,b,2 aDepartment of Kidney Development, Institute of Molecular Embryology and Genetics, Kumamoto University, Kumamoto 860-0811, Japan; bGlobal COE “Cell Fate Regulation Research and Education Unit,” Kumamoto University, Kumamoto 860-0811, Japan; cDepartment of Metabolic Medicine, Graduate School of Medical Sciences, Kumamoto University, Kumamoto 860-8556, Japan; dLaboratory for Animal Resources and Genetic Engineering, RIKEN Center for Developmental Biology, Kobe 650-0047, Japan; eDivision of Reproductive Engineering, Center for Animal Resources and Development, Kumamoto University, Kumamoto 860-0811, Japan; fLaboratory of Extracellular Matrix Biochemistry, Institute for Protein Research, Osaka University, Osaka 565-0871, Japan; and gLaboratory of Intracellular Signaling, Institute for Protein Research, Osaka University, Osaka 565-0871, Japan Edited by Eric N. Olson, University of Texas Southwestern, Dallas, TX, and approved April 15, 2010 (received for review November 27, 2009) The kidney develops through reciprocal interactions between two Results precursor tissues: the metanephric mesenchyme and the ureteric Kif26b Is Expressed in the Metanephric Mesenchyme During Nephro- fi bud. We previously demonstrated that the zinc nger protein Sall1 genesis. Mouse full-length Kif26b encodes a 2,112-aa protein is essential for ureteric bud attraction toward the mesenchyme. that shows 87% amino acid homology with human KIF26B and Here, we show that Kif26b, a kinesin family gene, is a downstream has a well conserved motor domain (96% identical to human Sall1 target of and that disruption of this gene causes kidney agen- KIF26B) in the N terminus (GenBank accession no. -

Upregulation of LINC00659 Expression Predicts a Poor Prognosis and Promotes Migration and Invasion of Gastric Cancer Cells
ONCOLOGY LETTERS 22: 557, 2021 Upregulation of LINC00659 expression predicts a poor prognosis and promotes migration and invasion of gastric cancer cells PIHAI GONG1,2, YING XU2, MIN LIU2, XIAOHUI SHEN1, YUHANG MAO2, YIPING LI3, KUN ZHANG4, SHENLING YU2 and HONG FAN1 1Department of Medical Genetics and Developmental Biology, Medical School of Southeast University, Key Laboratory of Developmental Genes and Human Diseases, Ministry of Education, Southeast University, Nanjing, Jiangsu 210009; 2School of Life Science and Technology, Southeast University, Nanjing, Jiangsu 210018; 3Department of Pathology, Medical School of Southeast University, Nanjing, Jiangsu 210009; 4Department of Medicine, The Third Affiliated Hospital of Harbin Medical University, Harbin, Heilongjiang 154000, P.R. China Received August 25, 2020; Accepted February 25, 2021 DOI: 10.3892/ol.2021.12818 Abstract. Long non‑coding RNAs (lncRNAs) serve an according to the Global Cancer Statistics 2018 (1). The 5‑year important role in the progression of cancer. LINC00659 was overall survival (OS) rate of patients with GC is <20%, and recently identified as a novel oncogenic lncRNA involved this low survival rate is partly due to a lack of effective early in colon cancer cell proliferation via modulating the cell diagnostic methods and prognostic indicators (2). In different cycle. However, the function of LINC00659 in other types of types of cancer, genomic instability is associated with chro‑ cancer, especially in gastric cancer (GC), remains unknown. mosomal aberrations, which affect numerous genes, further In the present study, bioinformatics analysis combined with promoting tumor progression (3). Comparative genomic cell experiments were performed to explore the function of hybridization (CGH) data have shown that gains of DNA copy LINC00659 in GC.