Relatively Frequent Switching of Transcription Start Sites During Cerebellar Development Peter Zhang1†, Emmanuel Dimont2,3†, Thomas Ha1, Douglas J
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Information Integrative Analyses of Splicing in the Aging Brain: Role in Susceptibility to Alzheimer’S Disease
Supplementary Information Integrative analyses of splicing in the aging brain: role in susceptibility to Alzheimer’s Disease Contents 1. Supplementary Notes 1.1. Religious Orders Study and Memory and Aging Project 1.2. Mount Sinai Brain Bank Alzheimer’s Disease 1.3. CommonMind Consortium 1.4. Data Availability 2. Supplementary Tables 3. Supplementary Figures Note: Supplementary Tables are provided as separate Excel files. 1. Supplementary Notes 1.1. Religious Orders Study and Memory and Aging Project Gene expression data1. Gene expression data were generated using RNA- sequencing from Dorsolateral Prefrontal Cortex (DLPFC) of 540 individuals, at an average sequence depth of 90M reads. Detailed description of data generation and processing was previously described2 (Mostafavi, Gaiteri et al., under review). Samples were submitted to the Broad Institute’s Genomics Platform for transcriptome analysis following the dUTP protocol with Poly(A) selection developed by Levin and colleagues3. All samples were chosen to pass two initial quality filters: RNA integrity (RIN) score >5 and quantity threshold of 5 ug (and were selected from a larger set of 724 samples). Sequencing was performed on the Illumina HiSeq with 101bp paired-end reads and achieved coverage of 150M reads of the first 12 samples. These 12 samples will serve as a deep coverage reference and included 2 males and 2 females of nonimpaired, mild cognitive impaired, and Alzheimer's cases. The remaining samples were sequenced with target coverage of 50M reads; the mean coverage for the samples passing QC is 95 million reads (median 90 million reads). The libraries were constructed and pooled according to the RIN scores such that similar RIN scores would be pooled together. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Toxicogenomics Applications of New Functional Genomics Technologies in Toxicology
\-\w j Toxicogenomics Applications of new functional genomics technologies in toxicology Wilbert H.M. Heijne Proefschrift ter verkrijging vand egraa dva n doctor opgeza gva nd e rector magnificus vanWageninge n Universiteit, Prof.dr.ir. L. Speelman, in netopenbaa r te verdedigen op maandag6 decembe r200 4 des namiddagst e half twee ind eAul a - Table of contents Abstract Chapter I. page 1 General introduction [1] Chapter II page 21 Toxicogenomics of bromobenzene hepatotoxicity: a combined transcriptomics and proteomics approach[2] Chapter III page 48 Bromobenzene-induced hepatotoxicity atth etranscriptom e level PI Chapter IV page 67 Profiles of metabolites and gene expression in rats with chemically induced hepatic necrosis[4] Chapter V page 88 Liver gene expression profiles in relation to subacute toxicity in rats exposed to benzene[5] Chapter VI page 115 Toxicogenomics analysis of liver gene expression in relation to subacute toxicity in rats exposed totrichloroethylen e [6] Chapter VII page 135 Toxicogenomics analysis ofjoin t effects of benzene and trichloroethylene mixtures in rats m Chapter VII page 159 Discussion and conclusions References page 171 Appendices page 187 Samenvatting page 199 Dankwoord About the author Glossary Abbreviations List of genes Chapter I General introduction Parts of this introduction were publishedin : Molecular Biology in Medicinal Chemistry, Heijne etal., 2003 m NATO Advanced Research Workshop proceedings, Heijne eral., 2003 81 Chapter I 1. General introduction 1.1 Background /.1.1 Toxicologicalrisk -

Whole Exome Sequencing in Families at High Risk for Hodgkin Lymphoma: Identification of a Predisposing Mutation in the KDR Gene
Hodgkin Lymphoma SUPPLEMENTARY APPENDIX Whole exome sequencing in families at high risk for Hodgkin lymphoma: identification of a predisposing mutation in the KDR gene Melissa Rotunno, 1 Mary L. McMaster, 1 Joseph Boland, 2 Sara Bass, 2 Xijun Zhang, 2 Laurie Burdett, 2 Belynda Hicks, 2 Sarangan Ravichandran, 3 Brian T. Luke, 3 Meredith Yeager, 2 Laura Fontaine, 4 Paula L. Hyland, 1 Alisa M. Goldstein, 1 NCI DCEG Cancer Sequencing Working Group, NCI DCEG Cancer Genomics Research Laboratory, Stephen J. Chanock, 5 Neil E. Caporaso, 1 Margaret A. Tucker, 6 and Lynn R. Goldin 1 1Genetic Epidemiology Branch, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; 2Cancer Genomics Research Laboratory, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; 3Ad - vanced Biomedical Computing Center, Leidos Biomedical Research Inc.; Frederick National Laboratory for Cancer Research, Frederick, MD; 4Westat, Inc., Rockville MD; 5Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; and 6Human Genetics Program, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD, USA ©2016 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol.2015.135475 Received: August 19, 2015. Accepted: January 7, 2016. Pre-published: June 13, 2016. Correspondence: [email protected] Supplemental Author Information: NCI DCEG Cancer Sequencing Working Group: Mark H. Greene, Allan Hildesheim, Nan Hu, Maria Theresa Landi, Jennifer Loud, Phuong Mai, Lisa Mirabello, Lindsay Morton, Dilys Parry, Anand Pathak, Douglas R. Stewart, Philip R. Taylor, Geoffrey S. Tobias, Xiaohong R. Yang, Guoqin Yu NCI DCEG Cancer Genomics Research Laboratory: Salma Chowdhury, Michael Cullen, Casey Dagnall, Herbert Higson, Amy A. -

Research Article
z Available online at http://www.journalcra.com INTERNATIONAL JOURNAL OF CURRENT RESEARCH International Journal of Current Research Vol. 10, Issue, 05, pp.69389-69394, May, 2018 ISSN: 0975-833X RESEARCH ARTICLE PATHWAY MAPPING OF PROTEINS INVOLVED IN MULTIPLE SCLEROSIS AND IDENTIFICATION OF NOVEL PROTEINS *Ruchi Yadav and Samreen Rizvi AMITY Institute of Biotechnology, AMITY University, Uttar Pradesh Lucknow Campus, 226028, UP, India ARTICLE INFO ABSTRACT Article History: WASH in the urban areas Multiple sclerosis is an unpredictable potentially disabling disease that Received 14th February, 2018 causes disruption of the myelin that insulates and protects nerve cells of spinal cord and brain and Received in revised form permits electrical impulses to be conducted along the nerve fiber with speed and accuracy. So this 02nd March, 2018 disease disrupts central nervous system by damaging or demyelinating the insulated covers of nerve Accepted 09th April, 2018 cells. In this paper we have made network of interacting proteins of more than 300 interacting Published online 30th May, 2018 proteins identified from protein interaction database .Initially 14 proteins were identified from Uni Prot database that were involved in multiple sclerosis disease . Gene Cards database was used to Key words: study pathways of proteins involved in network and new network was made that classifies interacting proteins on the basis of pathways involves. We found Three proteins, APBB2, APBB3 and MMP15 Multiple Sclerosis, Pathway Mapping, having no information about pathway and disease in which they are involved. New Network was Protein Interaction, String Database, Gene designed to predicted pathway of these novel proteins, APBB2, APBB3 and MMP15, network Mania Database. -

Termination of RNA Polymerase II Transcription by the 5’-3’ Exonuclease Xrn2
TERMINATION OF RNA POLYMERASE II TRANSCRIPTION BY THE 5’-3’ EXONUCLEASE XRN2 by MICHAEL ANDRES CORTAZAR OSORIO B.S., Universidad del Valle – Colombia, 2011 A thesis submitted to the Faculty of the Graduate School of the University of Colorado in partial fulfillment of the requirements for the degree of Doctor of Philosophy Molecular Biology Program 2018 This thesis for the Doctor of Philosophy degree by Michael Andrés Cortázar Osorio has been approved for the Molecular Biology Program by Mair Churchill, Chair Richard Davis Jay Hesselberth Thomas Blumenthal James Goodrich David Bentley, Advisor Date: Aug 17, 2018 ii Cortázar Osorio, Michael Andrés (Ph.D., Molecular Biology) Termination of RNA polymerase II transcription by the 5’-3’ exonuclease Xrn2 Thesis directed by Professor David L. Bentley ABSTRACT Termination of transcription occurs when RNA polymerase (pol) II dissociates from the DNA template and releases a newly-made mRNA molecule. Interestingly, an active debate fueled by conflicting reports over the last three decades is still open on which of the two main models of termination of RNA polymerase II transcription does in fact operate at 3’ ends of genes. The torpedo model indicates that the 5’-3’ exonuclease Xrn2 targets the nascent transcript for degradation after cleavage at the polyA site and chases pol II for termination. In contrast, the allosteric model asserts that transcription through the polyA signal induces a conformational change of the elongation complex and converts it into a termination-competent complex. In this thesis, I propose a unified allosteric-torpedo mechanism. Consistent with a polyA site-dependent conformational change of the elongation complex, I found that pol II transitions at the polyA site into a mode of slow transcription elongation that is accompanied by loss of Spt5 phosphorylation in the elongation complex. -

Systems Biology Approaches to Mining High Throughput Biological Data
BioMed Research International Systems Biology Approaches to Mining High Throughput Biological Data Guest Editors: Fang-Xiang Wu, Min Li, Jishou Ruan, and Feng Luo Systems Biology Approaches to Mining High Throughput Biological Data BioMed Research International Systems Biology Approaches to Mining High Throughput Biological Data Guest Editors: Fang-Xiang Wu, Min Li, Jishou Ruan, and Feng Luo Copyright © 2015 Hindawi Publishing Corporation. All rights reserved. This is a special issue published in “BioMed Research International.” All articles are open access articles distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Contents Systems Biology Approaches to Mining High Throughput Biological Data, Fang-Xiang Wu, Min Li, Jishou Ruan, and Feng Luo Volume 2015, Article ID 504362, 2 pages ProSim: A Method for Prioritizing Disease Genes Based on Protein Proximity and Disease Similarity, Gamage Upeksha Ganegoda, Yu Sheng, and Jianxin Wang Volume 2015, Article ID 213750, 11 pages Differential Expression Analysis in RNA-Seq by a Naive Bayes Classifier with Local Normalization, YongchaoDou,XiaomeiGuo,LinglingYuan,DavidR.Holding,andChiZhang Volume 2015, Article ID 789516, 9 pages ?? -Profiles: A Nonlinear Clustering Method for Pattern Detection in High Dimensional, Data Kai Wang, Qing Zhao, Jianwei Lu, and Tianwei Yu Volume 2015, Article ID 918954, 10 pages Screening Ingredients from Herbs against Pregnane X Receptor in -

Cellular Phenotyping of Hippocampal Progenitors Exposed to Patient Serum Predicts Conversion to Alzheimer’S Disease
Supplementary Materials for: Cellular phenotyping of hippocampal progenitors exposed to patient serum predicts conversion to Alzheimer’s Disease Aleksandra Maruszak, Tytus Murphy, Benjamine Liu, Chiara de Lucia, Abdel Douiri, Alejo J Nevado, Charlotte E Teunissen, Pieter Jelle Visser, Jack Price, Simon Lovestone, Sandrine Thuret* *Corresponding author: [email protected] This file includes: Fig. S1. Receiver-operator characteristic-curve for the cross-validation model predicting conversion to Alzheimer’s disease. Fig. S2. Receiver-operator characteristic-curve for predicting conversion to Alzheimer’s disease using a panel of 207 proteins. Table S1. Comparison of AUC for logistic regression models for individual predictors. Table S2. 207 proteins significantly differentially expressed between MCI converters and non- converters. 1 Fig. S1. Receiver-operator characteristic-curve for the cross-validation model predicting conversion to Alzheimer’s disease. Area under the curve, AUC=0.93, Sensitivity 90.3%, Specificity 79.0%. 2 Fig. S2. Receiver-operator characteristic-curve for predicting conversion to Alzheimer’s disease using a panel of 207 proteins. Area under the curve, AUC=0.943 Sensitivity= 91.65%, Specificity= 81.68%. 3 Table S3. Comparison of AUC for logistic regression models for individual predictors. P- value refers to the model. AUC OR SE 95% CI R2 p Education (years) 0.7562 0.79 0.06 0.68-0.93 0.1433 0.001 Average cell count 0.8080 1.01 0.004 1.01-1.024 0.3069 <0.0001 (proliferation) %Ki67+ cells (proliferation) 0.5728 1.04 0.05 0.94-1.15 0.0104 0.3919 %CC3+ cells (differentiation) 0.7972 2.97 1.08 1.45-6.08 0.2154 0.0001 4 Table S4. -
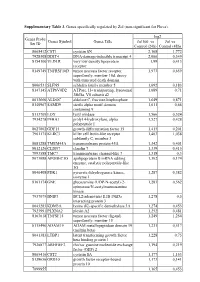
Supplementary Table 3. Genes Specifically Regulated by Zol (Non-Significant for Fluva)
Supplementary Table 3. Genes specifically regulated by Zol (non-significant for Fluva). log2 Genes Probe Genes Symbol Genes Title Zol100 vs Zol vs Set ID Control (24h) Control (48h) 8065412 CST1 cystatin SN 2,168 1,772 7928308 DDIT4 DNA-damage-inducible transcript 4 2,066 0,349 8154100 VLDLR very low density lipoprotein 1,99 0,413 receptor 8149749 TNFRSF10D tumor necrosis factor receptor 1,973 0,659 superfamily, member 10d, decoy with truncated death domain 8006531 SLFN5 schlafen family member 5 1,692 0,183 8147145 ATP6V0D2 ATPase, H+ transporting, lysosomal 1,689 0,71 38kDa, V0 subunit d2 8013660 ALDOC aldolase C, fructose-bisphosphate 1,649 0,871 8140967 SAMD9 sterile alpha motif domain 1,611 0,66 containing 9 8113709 LOX lysyl oxidase 1,566 0,524 7934278 P4HA1 prolyl 4-hydroxylase, alpha 1,527 0,428 polypeptide I 8027002 GDF15 growth differentiation factor 15 1,415 0,201 7961175 KLRC3 killer cell lectin-like receptor 1,403 1,038 subfamily C, member 3 8081288 TMEM45A transmembrane protein 45A 1,342 0,401 8012126 CLDN7 claudin 7 1,339 0,415 7993588 TMC7 transmembrane channel-like 7 1,318 0,3 8073088 APOBEC3G apolipoprotein B mRNA editing 1,302 0,174 enzyme, catalytic polypeptide-like 3G 8046408 PDK1 pyruvate dehydrogenase kinase, 1,287 0,382 isozyme 1 8161174 GNE glucosamine (UDP-N-acetyl)-2- 1,283 0,562 epimerase/N-acetylmannosamine kinase 7937079 BNIP3 BCL2/adenovirus E1B 19kDa 1,278 0,5 interacting protein 3 8043283 KDM3A lysine (K)-specific demethylase 3A 1,274 0,453 7923991 PLXNA2 plexin A2 1,252 0,481 8163618 TNFSF15 tumor necrosis -

A Meta-Analysis of the Effects of High-LET Ionizing Radiations in Human Gene Expression
Supplementary Materials A Meta-Analysis of the Effects of High-LET Ionizing Radiations in Human Gene Expression Table S1. Statistically significant DEGs (Adj. p-value < 0.01) derived from meta-analysis for samples irradiated with high doses of HZE particles, collected 6-24 h post-IR not common with any other meta- analysis group. This meta-analysis group consists of 3 DEG lists obtained from DGEA, using a total of 11 control and 11 irradiated samples [Data Series: E-MTAB-5761 and E-MTAB-5754]. Ensembl ID Gene Symbol Gene Description Up-Regulated Genes ↑ (2425) ENSG00000000938 FGR FGR proto-oncogene, Src family tyrosine kinase ENSG00000001036 FUCA2 alpha-L-fucosidase 2 ENSG00000001084 GCLC glutamate-cysteine ligase catalytic subunit ENSG00000001631 KRIT1 KRIT1 ankyrin repeat containing ENSG00000002079 MYH16 myosin heavy chain 16 pseudogene ENSG00000002587 HS3ST1 heparan sulfate-glucosamine 3-sulfotransferase 1 ENSG00000003056 M6PR mannose-6-phosphate receptor, cation dependent ENSG00000004059 ARF5 ADP ribosylation factor 5 ENSG00000004777 ARHGAP33 Rho GTPase activating protein 33 ENSG00000004799 PDK4 pyruvate dehydrogenase kinase 4 ENSG00000004848 ARX aristaless related homeobox ENSG00000005022 SLC25A5 solute carrier family 25 member 5 ENSG00000005108 THSD7A thrombospondin type 1 domain containing 7A ENSG00000005194 CIAPIN1 cytokine induced apoptosis inhibitor 1 ENSG00000005381 MPO myeloperoxidase ENSG00000005486 RHBDD2 rhomboid domain containing 2 ENSG00000005884 ITGA3 integrin subunit alpha 3 ENSG00000006016 CRLF1 cytokine receptor like -

Association of Plasma Proteins with Rate of Cognitive Decline and Dementia: 20-Year Follow-Up of the Whitehall II and ARIC Cohort Studies
medRxiv preprint doi: https://doi.org/10.1101/2020.11.18.20234070; this version posted November 20, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. 1 Association of plasma proteins with rate of cognitive decline and dementia: 20-year follow-up of the Whitehall II and ARIC cohort studies Joni V. Lindbohm, MD, PhD;1,2 Nina Mars, MD, PhD;3 Keenan A. Walker, PhD;4 Prof. Archana Singh-Manoux, PhD;1,5 Prof. Gill Livingston, MD, PhD;6,7 Prof. Eric J. Brunner, PhD;1 Pyry N. Sipilä, MD, PhD;2 Prof. Kalle Saksela, MD, PhD;8 Jane E. Ferrie, PhD;1,9 Prof. Ruth Lovering, PhD;10 Stephen A. Williams, MD, PhD;11 Prof. Aroon D. Hingorani, MD, PhD;12,13,14 Prof. Rebecca F. Gottesman, MD, PhD;4 Prof. Henrik Zetterberg, MD, PhD;15,16,17 Prof. Mika Kivimäki, PhD, FMedSci1,2 1 Department of Epidemiology and Public Health, University College London, 1-19 Torrington Place, London, UK 2 Clinicum, Department of Public Health, University of Helsinki, P.O. Box 41, FI-00014 Helsinki, Finland 3 Institute for Molecular Medicine Finland (FIMM), HiLIFE, University of Helsinki, Helsinki, Finland 4 Department of Neurology, The Johns Hopkins University, Baltimore, MD, USA. 5 Université de Paris, Inserm U1153, Epidemiology of Ageing and Neurodegenerative diseases, Paris, France 6 Division of Psychiatry, University College London, London, UK 7 Camden and Islington Foundation Trust, London, UK 8 Department of Virology, University of Helsinki and HUSLAB, Helsinki University Hospital, Helsinki, Finland 9 Bristol Medical School (PHS), University of Bristol, Bristol, UK 10 Functional Gene Annotation, Institute of Cardiovascular Science, University College London, London, UK 11 SomaLogic, Inc. -

Robust Sampling of Defective Pathways in Alzheimer's Disease. Implications in Drug Repositioning
International Journal of Molecular Sciences Article Robust Sampling of Defective Pathways in Alzheimer’s Disease. Implications in Drug Repositioning Juan Luis Fernández-Martínez 1,2,* , Óscar Álvarez-Machancoses 1,2 , Enrique J. deAndrés-Galiana 1,3 , Guillermina Bea 1 and Andrzej Kloczkowski 4,5 1 Group of Inverse Problems, Optimization and Machine Learning, Department of Mathematics, University of Oviedo, C/Federico García Lorca, 18, 33007 Oviedo, Spain; [email protected] (Ó.Á.-M.); [email protected] (E.J.d.-G.); [email protected] (G.B.) 2 DeepBioInsights, C/Federico García Lorca, 18, 33007 Oviedo, Spain 3 Department of Informatics and Computer Science, University of Oviedo, C/Federico García Lorca, 18, 33007 Oviedo, Spain 4 Battelle Center for Mathematical Medicine, Nationwide Children’s Hospital, Columbus, OH 43205, USA; [email protected] 5 Department of Pediatrics, The Ohio State University, Columbus, OH 43205, USA * Correspondence: [email protected] Received: 27 April 2020; Accepted: 13 May 2020; Published: 19 May 2020 Abstract: We present the analysis of the defective genetic pathways of the Late-Onset Alzheimer’s Disease (LOAD) compared to the Mild Cognitive Impairment (MCI) and Healthy Controls (HC) using different sampling methodologies. These algorithms sample the uncertainty space that is intrinsic to any kind of highly underdetermined phenotype prediction problem, by looking for the minimum-scale signatures (header genes) corresponding to different random holdouts. The biological pathways can be identified performing posterior analysis of these signatures established via cross-validation holdouts and plugging the set of most frequently sampled genes into different ontological platforms. That way, the effect of helper genes, whose presence might be due to the high degree of under determinacy of these experiments and data noise, is reduced.