Microcell-Mediated Chromosome Transfer Identifies EPB41L3 As a Functional Suppressor of Epithelial Ovarian Cancers
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Case of Prenatal Diagnosis of 18P Deletion Syndrome Following
Zhao et al. Molecular Cytogenetics (2019) 12:53 https://doi.org/10.1186/s13039-019-0464-y CASE REPORT Open Access A case of prenatal diagnosis of 18p deletion syndrome following noninvasive prenatal testing Ganye Zhao, Peng Dai, Shanshan Gao, Xuechao Zhao, Conghui Wang, Lina Liu and Xiangdong Kong* Abstract Background: Chromosome 18p deletion syndrome is a disease caused by the complete or partial deletion of the short arm of chromosome 18, there were few cases reported about the prenatal diagnosis of 18p deletion syndrome. Noninvasive prenatal testing (NIPT) is widely used in the screening of common fetal chromosome aneuploidy. However, the segmental deletions and duplications should also be concerned. Except that some cases had increased nuchal translucency or holoprosencephaly, most of the fetal phenotype of 18p deletion syndrome may not be evident during the pregnancy, 18p deletion syndrome was always accidentally discovered during the prenatal examination. Case presentations: In our case, we found a pure partial monosomy 18p deletion during the confirmation of the result of NIPT by copy number variation sequencing (CNV-Seq). The result of NIPT suggested that there was a partial or complete deletion of X chromosome. The amniotic fluid karyotype was normal, but result of CNV-Seq indicated a 7.56 Mb deletion on the short arm of chromosome 18 but not in the couple, which means the deletion was de novo deletion. Finally, the parents chose to terminate the pregnancy. Conclusions: To our knowledge, this is the first case of prenatal diagnosis of 18p deletion syndrome following NIPT.NIPT combined with ultrasound may be a relatively efficient method to screen chromosome microdeletions especially for the 18p deletion syndrome. -

PARSANA-DISSERTATION-2020.Pdf
DECIPHERING TRANSCRIPTIONAL PATTERNS OF GENE REGULATION: A COMPUTATIONAL APPROACH by Princy Parsana A dissertation submitted to The Johns Hopkins University in conformity with the requirements for the degree of Doctor of Philosophy Baltimore, Maryland July, 2020 © 2020 Princy Parsana All rights reserved Abstract With rapid advancements in sequencing technology, we now have the ability to sequence the entire human genome, and to quantify expression of tens of thousands of genes from hundreds of individuals. This provides an extraordinary opportunity to learn phenotype relevant genomic patterns that can improve our understanding of molecular and cellular processes underlying a trait. The high dimensional nature of genomic data presents a range of computational and statistical challenges. This dissertation presents a compilation of projects that were driven by the motivation to efficiently capture gene regulatory patterns in the human transcriptome, while addressing statistical and computational challenges that accompany this data. We attempt to address two major difficulties in this domain: a) artifacts and noise in transcriptomic data, andb) limited statistical power. First, we present our work on investigating the effect of artifactual variation in gene expression data and its impact on trans-eQTL discovery. Here we performed an in-depth analysis of diverse pre-recorded covariates and latent confounders to understand their contribution to heterogeneity in gene expression measurements. Next, we discovered 673 trans-eQTLs across 16 human tissues using v6 data from the Genotype Tissue Expression (GTEx) project. Finally, we characterized two trait-associated trans-eQTLs; one in Skeletal Muscle and another in Thyroid. Second, we present a principal component based residualization method to correct gene expression measurements prior to reconstruction of co-expression networks. -

Identification of Crucial Aberrantly Methylated and Differentially
Bioscience Reports (2020) 40 BSR20194365 https://doi.org/10.1042/BSR20194365 Research Article Identification of crucial aberrantly methylated and differentially expressed genes related to cervical cancer using an integrated bioinformatics analysis 1,2,3,* 1,* 1,2,3 1 1 2,3 1 Xiaoling Ma , Jinhui Liu , Hui Wang ,YiJiang,YicongWan, Yankai Xia and Wenjun Cheng Downloaded from http://portlandpress.com/bioscirep/article-pdf/40/5/BSR20194365/879046/bsr-2019-4365.pdf by guest on 28 September 2021 1Department of Gynecology, The First Affiliated Hospital of Nanjing Medical University, Nanjing, China; 2State Key Laboratory of Reproductive Medicine, Center for Global Health, School of Public Health, Nanjing Medical University, Nanjing, China; 3State Key Laboratory of Modern Toxicology of Ministry of Education, School of Public Health, Nanjing Medical University, Nanjing, China Correspondence: Wenjun Cheng ([email protected]) or Yankai Xia ([email protected]) Methylation functions in the pathogenesis of cervical cancer. In the present study, we ap- plied an integrated bioinformatics analysis to identify the aberrantly methylated and dif- ferentially expressed genes (DEGS), and their related pathways in cervical cancer. Data of gene expression microarrays (GSE9750) and gene methylation microarrays (GSE46306) were gained from Gene Expression Omnibus (GEO) databases. Hub genes were identi- fied by ‘limma’ packages and Venn diagram tool. Functional analysis was conducted by FunRich. Search Tool for the Retrieval of Interacting Genes Database (STRING) was used to analyze protein–protein interaction (PPI) information. Gene Expression Profiling Interac- tive Analysis (GEPIA), immunohistochemistry staining, and ROC curve analysis were con- ducted for validation. Gene Set Enrichment Analysis (GSEA) was also performed to iden- tify potential functions.We retrieved two upregulated-hypomethylated oncogenes and eight downregulated-hypermethylated tumor suppressor genes (TSGs) for functional analysis. -

Identification of Potential Key Genes and Pathway Linked with Sporadic Creutzfeldt-Jakob Disease Based on Integrated Bioinformatics Analyses
medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Identification of potential key genes and pathway linked with sporadic Creutzfeldt-Jakob disease based on integrated bioinformatics analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 , Iranna Kotturshetti 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. 3. Department of Ayurveda, Rajiv Gandhi Education Society`s Ayurvedic Medical College, Ron, Karnataka 562209, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India NOTE: This preprint reports new research that has not been certified by peer review and should not be used to guide clinical practice. medRxiv preprint doi: https://doi.org/10.1101/2020.12.21.20248688; this version posted December 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Abstract Sporadic Creutzfeldt-Jakob disease (sCJD) is neurodegenerative disease also called prion disease linked with poor prognosis. The aim of the current study was to illuminate the underlying molecular mechanisms of sCJD. The mRNA microarray dataset GSE124571 was downloaded from the Gene Expression Omnibus database. Differentially expressed genes (DEGs) were screened. -

PROTEOME of the HUMAN CHROMOSOME 18: GENE-CENTRIC IDENTIFICATION of TRANSCRIPTS, PROTEINS and PEPTIDES Addendum to the Roadmap
PROTEOME OF THE HUMAN CHROMOSOME 18: GENE-CENTRIC IDENTIFICATION OF TRANSCRIPTS, PROTEINS AND PEPTIDES Addendum to the Roadmap: HEALTH ASPECTS 1. PROTEOMICS MEETS MEDICINE At its very beginning, one of the goals of human proteomics became a disease biomarker discovery. Many works compared diseased and normal tissues and liquids to get diagnostic profiles by many proteomics methods. Of them, some cancer proteome profiling studies were considered too optimistic in terms of clinical applicability due to incorrect experimental design [Petricoin], thereby conferring the negative expectations from proteomics in translational medicine [Diamantidis, Nature]. The interlaboratory reproducibility of proteomics pipelines also was considered as a shortage in some papers, e.g. in the works of Bell et al [2009] who tested the proteome MS methods with 20-protein standard sample. These difficulties at the early stage of proteomics were partly caused by the fact that many attempts were mostly directed to the technique adjustment rather than to the clinically relevant result. However, the recent advances in mass-spectrometry including the use of MRM to quantify peptides of proteome [Anderson Hunter 2006] made the community to have a view of cautious optimism on the problem of translation to medicine [Nilsson 2010]. A reproducibility problem stated in [Bell 2009] was shown to be mainly caused by the bioinformatics misinterpretation whereas the MS itself worked properly. The readiness of MRM-based platforms to the clinical use is illustrated by the attempt to pass FDA with the mock application which describes MS-based quantitation test for 10 proteins [Regnier FE 2010]. In its current state, the test has not got a clearance. -

Views for Entrez
BASIC RESEARCH www.jasn.org Phosphoproteomic Analysis Reveals Regulatory Mechanisms at the Kidney Filtration Barrier †‡ †| Markus M. Rinschen,* Xiongwu Wu,§ Tim König, Trairak Pisitkun,¶ Henning Hagmann,* † † † Caroline Pahmeyer,* Tobias Lamkemeyer, Priyanka Kohli,* Nicole Schnell, †‡ †† ‡‡ Bernhard Schermer,* Stuart Dryer,** Bernard R. Brooks,§ Pedro Beltrao, †‡ Marcus Krueger,§§ Paul T. Brinkkoetter,* and Thomas Benzing* *Department of Internal Medicine II, Center for Molecular Medicine, †Cologne Excellence Cluster on Cellular Stress | Responses in Aging-Associated Diseases, ‡Systems Biology of Ageing Cologne, Institute for Genetics, University of Cologne, Cologne, Germany; §Laboratory of Computational Biology, National Heart, Lung, and Blood Institute, National Institutes of Health, Bethesda, Maryland; ¶Faculty of Medicine, Chulalongkorn University, Bangkok, Thailand; **Department of Biology and Biochemistry, University of Houston, Houston, Texas; ††Division of Nephrology, Baylor College of Medicine, Houston, Texas; ‡‡European Molecular Biology Laboratory–European Bioinformatics Institute, Hinxton, Cambridge, United Kingdom; and §§Max Planck Institute for Heart and Lung Research, Bad Nauheim, Germany ABSTRACT Diseases of the kidney filtration barrier are a leading cause of ESRD. Most disorders affect the podocytes, polarized cells with a limited capacity for self-renewal that require tightly controlled signaling to maintain their integrity, viability, and function. Here, we provide an atlas of in vivo phosphorylated, glomerulus- expressed -

Transdifferentiation of Human Mesenchymal Stem Cells
Transdifferentiation of Human Mesenchymal Stem Cells Dissertation zur Erlangung des naturwissenschaftlichen Doktorgrades der Julius-Maximilians-Universität Würzburg vorgelegt von Tatjana Schilling aus San Miguel de Tucuman, Argentinien Würzburg, 2007 Eingereicht am: Mitglieder der Promotionskommission: Vorsitzender: Prof. Dr. Martin J. Müller Gutachter: PD Dr. Norbert Schütze Gutachter: Prof. Dr. Georg Krohne Tag des Promotionskolloquiums: Doktorurkunde ausgehändigt am: Hiermit erkläre ich ehrenwörtlich, dass ich die vorliegende Dissertation selbstständig angefertigt und keine anderen als die von mir angegebenen Hilfsmittel und Quellen verwendet habe. Des Weiteren erkläre ich, dass diese Arbeit weder in gleicher noch in ähnlicher Form in einem Prüfungsverfahren vorgelegen hat und ich noch keinen Promotionsversuch unternommen habe. Gerbrunn, 4. Mai 2007 Tatjana Schilling Table of contents i Table of contents 1 Summary ........................................................................................................................ 1 1.1 Summary.................................................................................................................... 1 1.2 Zusammenfassung..................................................................................................... 2 2 Introduction.................................................................................................................... 4 2.1 Osteoporosis and the fatty degeneration of the bone marrow..................................... 4 2.2 Adipose and bone -

ZFP226 Is a Novel Artificial Transcription Factor for Selective
www.nature.com/scientificreports OPEN ZFP226 is a novel artifcial transcription factor for selective activation of tumor suppressor Received: 5 October 2017 Accepted: 26 February 2018 KIBRA Published: xx xx xxxx Katrin Schelleckes1, Boris Schmitz2, Malte Lenders1, Mirja Mewes1, Stefan-Martin Brand2 & Eva Brand1 KIBRA has been suggested as a key regulator of the hippo pathway, regulating organ size, cell contact inhibition as well as tissue regeneration and tumorigenesis. Recently, alterations of KIBRA expression caused by promotor methylation have been reported for several types of cancer. Our current study aimed to design an artifcial transcription factor capable of re-activating expression of the tumor suppressor KIBRA and the hippo pathway. We engineered a new gene named ‘ZFP226′ encoding for a ~23 kDa fusion protein. ZFP226 belongs to the Cys2-His2 zinc fnger type and recognizes a nine base-pair DNA sequence 5′-GGC-GGC-GGC-3′ in the KIBRA core promoter P1a. ZFP226 showed nuclear localization in human immortalized kidney epithelial cells and activated the KIBRA core promoter (p < 0.001) resulting in signifcantly increased KIBRA mRNA and protein levels (p < 0.001). Furthermore, ZFP226 led to activation of hippo signaling marked by elevated YAP and LATS phosphorylation. In Annexin V fow cytometry assays ZFP226 overexpression showed strong pro-apoptotic capacity on MCF-7 breast cancer cells (p < 0.01 early-, p < 0.001 late-apoptotic cells). We conclude that the artifcial transcription factor ZFP226 can be used for target KIBRA and hippo pathway activation. This novel molecule may represent a molecular tool for the development of future applications in cancer treatment. -

Protein 4.1, a Component of the Erythrocyte Membrane Skeleton and Its Related Homologue Proteins Forming the Protein 4.1/FERM Superfamily
FOLIA HISTOCHEMICA Review article ET CYTOBIOLOGICA Vol. 44, No. 4, 2006 pp. 231-248 Protein 4.1, a component of the erythrocyte membrane skeleton and its related homologue proteins forming the protein 4.1/FERM superfamily Witold Diakowski, Micha³ Grzybek and Aleksander F. Sikorski Faculty of Biotechnology, University of Wroc³aw, Wroc³aw, Poland Abstract: The review is focused on the domain structure and function of protein 4.1, one of the proteins belonging to the mem- brane skeleton. The protein 4.1 of the red blood cells (4.1R) is a multifunctional protein that localizes to the membrane skele- ton and stabilizes erythrocyte shape and membrane mechanical properties, such as deformability and stability, via lateral inter- actions with spectrin, actin, glycophorin C and protein p55. Protein 4.1 binding is modulated through the action of kinases and/or calmodulin-Ca2+. Non-erythroid cells express the 4.1R homologues: 4.1G (general type), 4.1B (brain type), and 4.1N (neuron type), and the whole group belongs to the protein 4.1 superfamily, which is characterized by the presence of a highly conserved FERM domain at the N-terminus of the molecule. Proteins 4.1R, 4.1 G, 4.1 N and 4.1 B are encoded by different genes. Most of the 4.1 superfamily proteins also contain an actin-binding domain. To date, more than 40 members have been identified. They can be divided into five groups: protein 4.1 molecules, ERM proteins, talin-related molecules, protein tyrosine phosphatase (PTPH) proteins and NBL4 proteins. We have focused our attention on the main, well known representatives of 4.1 superfamily and tried to choose the proteins which are close to 4.1R or which have distinct functions. -
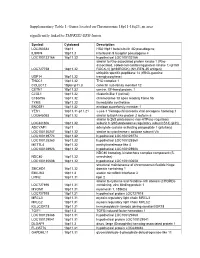
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

The Proteomic Analysis of Breast Cell Line Exosomes Reveals Disease Patterns and Potential Biomarkers Yousef Risha1, Zoran Minic2, Shahrokh M
www.nature.com/scientificreports OPEN The proteomic analysis of breast cell line exosomes reveals disease patterns and potential biomarkers Yousef Risha1, Zoran Minic2, Shahrokh M. Ghobadloo3 & Maxim V. Berezovski 1,2,3* Cancer cells release small extracellular vesicles, exosomes, that have been shown to contribute to various aspects of cancer development and progression. Diferential analysis of exosomal proteomes from cancerous and non-tumorigenic breast cell lines can provide valuable information related to breast cancer progression and metastasis. Moreover, such a comparison can be explored to fnd potentially new protein biomarkers for early disease detection. In this study, exosomal proteomes of MDA-MB-231, a metastatic breast cancer cell line, and MCF-10A, a non-cancerous epithelial breast cell line, were identifed by nano-liquid chromatography coupled to tandem mass spectrometry. We also tested three exosomes isolation methods (ExoQuick, Ultracentrifugation (UC), and Ultrafltration– Ultracentrifugation) and detergents (n-dodecyl β-d-maltoside, Triton X-100, and Digitonin) for solubilization of exosomal proteins and enhanced detection by mass spectrometry. A total of 1,107 exosomal proteins were identifed in both cell lines, 726 of which were unique to the MDA-MB-231 breast cancer cell line. Among them, 87 proteins were predicted to be relevant to breast cancer and 16 proteins to cancer metastasis. Three exosomal membrane/surface proteins, glucose transporter 1 (GLUT-1), glypican 1 (GPC-1), and disintegrin and metalloproteinase domain-containing protein 10 (ADAM10), were identifed as potential breast cancer biomarkers and validated with Western blotting and high-resolution fow cytometry. We demonstrated that exosomes are a rich source of breast cancer-related proteins and surface biomarkers that may be used for disease diagnosis and prognosis. -

Mouse Epb41l3 Conditional Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Epb41l3 Conditional Knockout Project (CRISPR/Cas9) Objective: To create a Epb41l3 conditional knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Epb41l3 gene (NCBI Reference Sequence: NM_013813 ; Ensembl: ENSMUSG00000024044 ) is located on Mouse chromosome 17. 22 exons are identified, with the ATG start codon in exon 2 and the TGA stop codon in exon 21 (Transcript: ENSMUST00000080208). Exon 2 will be selected as conditional knockout region (cKO region). Deletion of this region should result in the loss of function of the Mouse Epb41l3 gene. To engineer the targeting vector, homologous arms and cKO region will be generated by PCR using BAC clone RP23-391O3 as template. Cas9, gRNA and targeting vector will be co-injected into fertilized eggs for cKO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Mice homozygous for one knock-out allele display a normal phenotype. Mice homozygous for a different knock-out allele exhibit ataxia, gait abnormalities, clasping, hypermyelination, abnormal axon morphology, and decreased internode length. Exon 2 starts from about 100% of the coding region. The knockout of Exon 2 will result in frameshift of the gene. The size of intron 1 for 5'-loxP site insertion: 50948 bp, and the size of intron 2 for 3'-loxP site insertion: 2109 bp. The size of effective cKO region: ~707 bp. The cKO region does not have any other known gene. Page 1 of 8 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele gRNA region 5' gRNA region 3' 1 2 3 22 Targeting vector Targeted allele Constitutive KO allele (After Cre recombination) Legends Exon of mouse Epb41l3 Homology arm cKO region loxP site Page 2 of 8 https://www.alphaknockout.com Overview of the Dot Plot Window size: 10 bp Forward Reverse Complement Sequence 12 Note: The sequence of homologous arms and cKO region is aligned with itself to determine if there are tandem repeats.