Suppl Figure 1
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Structure and Function of the Fgd Family of Divergent FYVE Domain Proteins
Biochemistry and Cell Biology Structure and Function of the Fgd Family of Divergent FYVE Domain Proteins Journal: Biochemistry and Cell Biology Manuscript ID bcb-2018-0185.R1 Manuscript Type: Mini Review Date Submitted by the 03-Aug-2018 Author: Complete List of Authors: Eitzen, Gary; University of Alberta Faculty of Medicine and Dentistry Smithers, Cameron C.; University of Alberta, Biochemistry Murray, Allan; University of Alberta Faculty of Medicine and Dentistry Overduin, Michael; University of Alberta Faculty of Medicine and Dentistry Draft Fgd, Pleckstrin Homology domain, FYVE domain, Dbl Homology Domain, Keyword: Rho GEF Is the invited manuscript for consideration in a Special CSMB Special Issue Issue? : https://mc06.manuscriptcentral.com/bcb-pubs Page 1 of 37 Biochemistry and Cell Biology Title: Structure and Function of the Fgd Family of Divergent FYVE Domain Proteins Authors: Gary Eitzen1, Cameron C. Smithers2, Allan G Murray3 and Michael Overduin2* Draft 1Department of Cell Biology, 2Department of Biochemistry, 3Department of Medicine, University of Alberta, Edmonton, Alberta, Canada *Corresponding author. Michael Overduin Telephone: +1 780 492 3518 Fax: +1 780 492-0886 E-mail: [email protected] https://mc06.manuscriptcentral.com/bcb-pubs Biochemistry and Cell Biology Page 2 of 37 Abstract FYVE domains are highly conserved protein modules that typically bind phosphatidylinositol 3-phosphate (PI3P) on the surface of early endosomes. Along with pleckstrin homology (PH) and phox homology (PX) domains, FYVE domains are the principal readers of the phosphoinositide (PI) code that mediate specific recognition of eukaryotic organelles. Of all the human FYVE domain-containing proteins, those within the Faciogenital dysplasia (Fgd) subfamily are particularly divergent, and couple with GTPases to exert unique cellular functions. -

JB Review Biochemical and Structural Properties of Heterochromatin Protein 1: Understanding Its Role in Chromatin Assembly
J. Biochem. 2014;1Journal10 doi:10.1093/jb/mvu032 of Biochemistry Advance Access published June 4, 2014 JB Review Biochemical and structural properties of heterochromatin protein 1: understanding its role in chromatin assembly Received March 20, 2014; accepted April 9, 2014; published online May 13, 2014 Gohei Nishibuchi and Jun-ichi Nakayama* homologues and isoforms have been identified in eu- karyotic organisms, ranging from fission yeast to Graduate School of Natural Sciences, Nagoya City University, mammals, flies, and nematodes (69). In mammals, Nagoya 467-8501, Japan the three HP1 isoforms HP1a, HP1b and HP1g *Jun-ichi Nakayama, Graduate School of Natural Sciences, Nagoya (Fig. 1A) have different localization patterns within City University, 1 Yamanohata, Mizuho-cho, Mizuho-ku, Nagoya cells (11, 12). Mice deficient for each HP1 isoform ex- 467-8501, Japan. Tel: þ81-52-872-5838, Fax: þ81-52-872-5866, email: [email protected] hibit different phenotypes (1315), indicating that the isoforms have distinct functions in forming higher- Heterochromatin protein 1 (HP1) is an evolutionarily order chromatin structures. conserved chromosomal protein that binds lysine HP1-family proteins share a basic structure consist- Downloaded from 9-methylated histone H3 (H3K9me), a hallmark of het- ing of an amino-terminal chromodomain (CD) and a erochromatin, and plays a crucial role in forming carboxyl-terminal chromo shadow domain (CSD) higher-order chromatin structures. HP1 has an N-ter- linked by an unstructured hinge region (16). The CD minal chromodomain and a C-terminal chromo shadow functions as a binding module that targets lysine domain, linked by an unstructured hinge region. -

Functional Roles of Bromodomain Proteins in Cancer
cancers Review Functional Roles of Bromodomain Proteins in Cancer Samuel P. Boyson 1,2, Cong Gao 3, Kathleen Quinn 2,3, Joseph Boyd 3, Hana Paculova 3 , Seth Frietze 3,4,* and Karen C. Glass 1,2,4,* 1 Department of Pharmaceutical Sciences, Albany College of Pharmacy and Health Sciences, Colchester, VT 05446, USA; [email protected] 2 Department of Pharmacology, Larner College of Medicine, University of Vermont, Burlington, VT 05405, USA; [email protected] 3 Department of Biomedical and Health Sciences, University of Vermont, Burlington, VT 05405, USA; [email protected] (C.G.); [email protected] (J.B.); [email protected] (H.P.) 4 University of Vermont Cancer Center, Burlington, VT 05405, USA * Correspondence: [email protected] (S.F.); [email protected] (K.C.G.) Simple Summary: This review provides an in depth analysis of the role of bromodomain-containing proteins in cancer development. As readers of acetylated lysine on nucleosomal histones, bromod- omain proteins are poised to activate gene expression, and often promote cancer progression. We examined changes in gene expression patterns that are observed in bromodomain-containing proteins and associated with specific cancer types. We also mapped the protein–protein interaction network for the human bromodomain-containing proteins, discuss the cellular roles of these epigenetic regu- lators as part of nine different functional groups, and identify bromodomain-specific mechanisms in cancer development. Lastly, we summarize emerging strategies to target bromodomain proteins in cancer therapy, including those that may be essential for overcoming resistance. Overall, this review provides a timely discussion of the different mechanisms of bromodomain-containing pro- Citation: Boyson, S.P.; Gao, C.; teins in cancer, and an updated assessment of their utility as a therapeutic target for a variety of Quinn, K.; Boyd, J.; Paculova, H.; cancer subtypes. -

Tsdhn-2, a Unique Dehydrin Protein from Thellungiella and Its Role in Salt Tolerance
TsDHN-2, A Unique Dehydrin Protein from Thellungiella and its Role in Salt Tolerance A Thesis Submitted to the College of Graduate Studies and Research in Partial Fulfillment of the Requirements for the Degree of Master of Science in the Department of Biochemistry University of Saskatchewan Saskatoon by Sarah Catherine Klatt © Copyright Sarah C. Klatt, July, 2011. All rights reserved. - i - PERMISSION TO USE In presenting this thesis in partial fulfillment of the requirements for a master’s degree from the University of Saskatchewan. I agree that the Libraries of this University may make it freely available for inspection. Moreover, I agree that permission for copying of this thesis in any manner, in whole or in part, for scholarly purposes may be granted by the department or the dean of the college in which this thesis work was done. It is understood that any copying or publication or use of this thesis or parts there of for financial gain without approval by the University of Saskatchewan and the author’s written permission is prohibited. It is also understood that due recognition shall be given to the author and to the University of Saskatchewan in any scholarly use which may be made of any material used in this thesis. Requests for permission to copy or to make either use of the material presented in this thesis in while or part should be addressed to: Head of the Department of Biochemistry University of Saskatchewan Saskatoon, Saskatchewan S7N 5A8 CANADA - iii - - ABSTRACT Salt stress, or salinity, is one of the most common environmental stresses affecting crop yield worldwide. -

Adaptive Responses by Transcriptional Regulators to Small Molecules in Prokaryotes
Adaptive Responses by Transcriptional Regulators to small molecules in Prokaryotes Structural studies of two bacterial one-component signal transduction systems DntR and HpNikR Cyril Dian Stockholm University Doctoral thesis © Cyril Dian, Stockholm 2007 ISBN 978-91-7155-500-7 Department of Biochemistry and Biophysics The Arrhenius Laboratories for Natural Sciences Stockholm University SE-106 91 Stockholm Sweden All previously published papers are reprinted With permission from the publishers Intellecta Docusys, Stockholm 2007 Abstract Prokaryotes are continually exposed to environmental changes in their physiological conditions. In order to survive such unstable conditions, or to compete with others species for the same environmental niche, prokaryotes must monitor signals about both their extracellular environment and intracellular physiological status and provide rapid and appropriate responses to variations in their surroundings. This adaptive response to environmental signals is triggered mainly by transcriptional regulators via two components, the one- and two-component signal transduction systems. These scan intra- and extracellular small-molecule mixtures and modulate gene expression to provide the appropriate physiological response to the prevailing conditions. Most prokaryotic one component regulators are simple transcription factors comprising of a small-molecule binding domain (SMBD) and a DNA binding domain (DBD). Although the effects of transcription factors on the transcription machinery are well understood, the exact location -

Regulation of Stringent Factor by Branched-Chain Amino Acids
Regulation of stringent factor by branched-chain amino acids Mingxu Fanga and Carl E. Bauera,1 aMolecular and Cellular Biochemistry Department, Indiana University, Bloomington, IN 47405 Edited by Caroline S. Harwood, University of Washington, Seattle, WA, and approved May 9, 2018 (received for review February 21, 2018) When faced with amino acid starvation, prokaryotic cells induce a Under normal growth conditions, the synthetase activity of Rel is stringent response that modulates their physiology. The stringent thought to be self-inhibited; however, during times of amino acid response is manifested by production of signaling molecules starvation, Rel interacts with stalled ribosomes, which activates guanosine 5′-diphosphate,3′-diphosphate (ppGpp) and guanosine synthetase activity to produce (p)ppGpp. The regulation of hy- 5′-triphosphate,3′-diphosphate (pppGpp) that are also called drolase activity is less understood but may involve one or more alarmones. In many species, alarmone levels are regulated by a downstream domains called the TGS and ACT domains. The TGS multidomain bifunctional alarmone synthetase/hydrolase called domain of SpoT has been shown to interact with an acyl carrier Rel. In this enzyme, there is an ACT domain at the carboxyl region protein, so it is presumed to sense the status of fatty acid metab- that has an unknown function; however, similar ACT domains are olism in E. coli (4). The function of the ACT domain is not as clear; present in other enzymes that have roles in controlling amino acid however, recent cryo-EM structures of E. coli RelA show that this metabolism. In many cases, these other ACT domains have been domain is involved in binding deacyl-tRNA as well as the ribosome shown to allosterically regulate enzyme activity through the bind- (5–7). -

Arabidopsis Thaliana
Downloaded from genome.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press RESEARCH A Physical Map of Chromosome 2 of Arabidopsis thaliana Eve Ann Zachgo, 2,4 Ming Li Wang, 1'2'4 Julia Dewdney, 1'2 David Bouchez, 3 Christine Carnilleri, 3 Stephen Belmonte, 2 Lu Huang, 2 Maureen Dolan, 2 and Howard M. Goodman 1'2'5 1Department of Genetics, Harvard Medical School and 2Department of Molecular Biology, Massachusetts General Hospital, Boston, Massachusetts 02114; 3Laboratoire de Biologie Cellulaire, Institut National de la Recherche Agronomique (INRA), 78026 Versailles CEDEX, France A yeast artificial chromosome (YAC] physical map of chromosome 2 of Arabidopsis thaliana has been constructed by hybridization of 69 DNA markers and 61 YAC end probes to gridded arrays of YAC clones. Thirty-four YACs in four contigs define the chromosome. Complete closure of the map was not attained because some regions of the chromosome were repetitive or were not represented in the YAC library. Based on the sizes of the YACs and their coverage of the chromosome, the length of chromosome 2 is estimated to be at least 18 Mb. These data provide the means for immediately identifying the YACs containing a genetic locus mapped on Arabidopsis chromosome 2. The small flowering plant Arabidopsis thaliana is ters (Maluszynska and Heslop-Harrison 1991; A1- an excellent model system for metabolic, genetic, bini 1994; Copenhaver et al. 1995). We present and developmental studies in plants. Its haploid here a YAC contig physical map of chromosome nuclear genome is small (-100 Mb), consisting of 2 of A. -

Expressed Sequence Tag Analysis of the Response of Apple
Physiologia Plantarum 133: 298–317. 2008 Copyright ª Physiologia Plantarum 2008, ISSN 0031-9317 Expressed sequence tag analysis of the response of apple (Malus x domestica ‘Royal Gala’) to low temperature and water deficit Michael Wisniewskia,*, Carole Bassetta,*, John Norellia, Dumitru Macarisina, Timothy Artlipa, Ksenija Gasicb and Schuyler Korbanb aUnited States Department of Agriculture – Agricultural Research Service (USDA-ARS), Appalachian Fruit Research Station, 2217 Wiltshire Road, Kearneysville, WV 25430, USA bDepartment of Natural Resources and Environmental Sciences, University of Illinois, 310 ERML, 1201 W. Gregory Drive, Urbana, IL 61801, USA Correspondence Leaf, bark, xylem and root tissues were used to make nine cDNA libraries from *Corresponding author, non-stressed (control) ‘Royal Gala’ apple trees, and from ‘Royal Gala’ trees e-mail: [email protected]; exposed to either low temperature (5°C for 24 h) or water deficit (45% of [email protected] saturated pot mass for 2 weeks). Over 22 600 clones from the nine libraries # Received 26 September 2007; revised 3 were subjected to 5 single-pass sequencing, clustered and annotated using January 2008 BLASTX. The number of clusters in the libraries ranged from 170 to 1430. Regarding annotation of the sequences, BLASTX analysis indicated that within doi: 10.1111/j.1399-3054.2008.01063.x the libraries 65–72% of the clones had a high similarity to known function genes, 6–15% had no functional assignment and 15–26% were completely novel. The expressed sequence tags were combined into three classes (control, low-temperature and water deficit) and the annotated genes in each class were placed into 1 of 10 different functional categories. -

To Find Information About Arabidopsis Genes Leonore Reiser1, Shabari
UNIT 1.11 Using The Arabidopsis Information Resource (TAIR) to Find Information About Arabidopsis Genes Leonore Reiser1, Shabari Subramaniam1, Donghui Li1, and Eva Huala1 1Phoenix Bioinformatics, Redwood City, CA USA ABSTRACT The Arabidopsis Information Resource (TAIR; http://arabidopsis.org) is a comprehensive Web resource of Arabidopsis biology for plant scientists. TAIR curates and integrates information about genes, proteins, gene function, orthologs gene expression, mutant phenotypes, biological materials such as clones and seed stocks, genetic markers, genetic and physical maps, genome organization, images of mutant plants, protein sub-cellular localizations, publications, and the research community. The various data types are extensively interconnected and can be accessed through a variety of Web-based search and display tools. This unit primarily focuses on some basic methods for searching, browsing, visualizing, and analyzing information about Arabidopsis genes and genome, Additionally we describe how members of the community can share data using TAIR’s Online Annotation Submission Tool (TOAST), in order to make their published research more accessible and visible. Keywords: Arabidopsis ● databases ● bioinformatics ● data mining ● genomics INTRODUCTION The Arabidopsis Information Resource (TAIR; http://arabidopsis.org) is a comprehensive Web resource for the biology of Arabidopsis thaliana (Huala et al., 2001; Garcia-Hernandez et al., 2002; Rhee et al., 2003; Weems et al., 2004; Swarbreck et al., 2008, Lamesch, et al., 2010, Berardini et al., 2016). The TAIR database contains information about genes, proteins, gene expression, mutant phenotypes, germplasms, clones, genetic markers, genetic and physical maps, genome organization, publications, and the research community. In addition, seed and DNA stocks from the Arabidopsis Biological Resource Center (ABRC; Scholl et al., 2003) are integrated with genomic data, and can be ordered through TAIR. -

The Capacity of Long-Term in Vitro Proliferation of Acute Myeloid
The Capacity of Long-Term in Vitro Proliferation of Acute Myeloid Leukemia Cells Supported Only by Exogenous Cytokines Is Associated with a Patient Subset with Adverse Outcome Annette K. Brenner, Elise Aasebø, Maria Hernandez-Valladares, Frode Selheim, Frode Berven, Ida-Sofie Grønningsæter, Sushma Bartaula-Brevik and Øystein Bruserud Supplementary Material S2 of S31 Table S1. Detailed information about the 68 AML patients included in the study. # of blasts Viability Proliferation Cytokine Viable cells Change in ID Gender Age Etiology FAB Cytogenetics Mutations CD34 Colonies (109/L) (%) 48 h (cpm) secretion (106) 5 weeks phenotype 1 M 42 de novo 241 M2 normal Flt3 pos 31.0 3848 low 0.24 7 yes 2 M 82 MF 12.4 M2 t(9;22) wt pos 81.6 74,686 low 1.43 969 yes 3 F 49 CML/relapse 149 M2 complex n.d. pos 26.2 3472 low 0.08 n.d. no 4 M 33 de novo 62.0 M2 normal wt pos 67.5 6206 low 0.08 6.5 no 5 M 71 relapse 91.0 M4 normal NPM1 pos 63.5 21,331 low 0.17 n.d. yes 6 M 83 de novo 109 M1 n.d. wt pos 19.1 8764 low 1.65 693 no 7 F 77 MDS 26.4 M1 normal wt pos 89.4 53,799 high 3.43 2746 no 8 M 46 de novo 26.9 M1 normal NPM1 n.d. n.d. 3472 low 1.56 n.d. no 9 M 68 MF 50.8 M4 normal D835 pos 69.4 1640 low 0.08 n.d. -
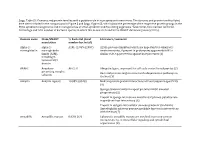
Supp. Table S2: Domains and Protein Families with a Putative Role in Host-Symbiont Interactions
Supp. Table S2: Domains and protein families with a putative role in host-symbiont interactions. The domains and protein families listed here were included in the comparisons in Figure 5 and Supp. Figure S5, which show the percentage of the respective protein groups in the Riftia symbiont metagenome and in metagenomes of other symbiotic and free-living organisms. % bacterial, total number bacterial: Percentage and total number of bacterial species in which this domain is found in the SMART database (January 2019). Domain name Pfam/SMART % bacterial (total Literature/comment annotation number bacterial) Alpha-2- alpha-2- A2M: 42.05% (2057) A2Ms: protease inhibitors which are important for eukaryotic macroglobulin macroglobulin innate immunity, if present in prokaryotes apparently fulfill a family (A2M), similar role, e.g. protection against host proteases (1) including N- terminal MG1 domain ANAPC Anaphase- APC2: 0 Ubiquitin ligase, important for cell cycle control in eukaryotes (2) promoting complex Bacterial proteins might interact with ubiquitination pathways in subunits the host (3) Ankyrin Ankyrin repeats 10.88% (8348) Mediate protein-protein interactions without sequence specificity (4) Sponge symbiont ankyrin-repeat proteins inhibit amoebal phagocytosis (5) Present in sponge microbiome metatranscriptomes, putative role in symbiont-host interactions (6) Present in obligate intracellular amoeba symbiont Candidatus Amoebophilus asiaticus genome, probable function in interactions with the host (7) Armadillo Armadillo repeats 0.83% (67) -

Flavonoid Glucodiversification with Engineered Sucrose-Active Enzymes Yannick Malbert
Flavonoid glucodiversification with engineered sucrose-active enzymes Yannick Malbert To cite this version: Yannick Malbert. Flavonoid glucodiversification with engineered sucrose-active enzymes. Biotechnol- ogy. INSA de Toulouse, 2014. English. NNT : 2014ISAT0038. tel-01219406 HAL Id: tel-01219406 https://tel.archives-ouvertes.fr/tel-01219406 Submitted on 22 Oct 2015 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Last name: MALBERT First name: Yannick Title: Flavonoid glucodiversification with engineered sucrose-active enzymes Speciality: Ecological, Veterinary, Agronomic Sciences and Bioengineering, Field: Enzymatic and microbial engineering. Year: 2014 Number of pages: 257 Flavonoid glycosides are natural plant secondary metabolites exhibiting many physicochemical and biological properties. Glycosylation usually improves flavonoid solubility but access to flavonoid glycosides is limited by their low production levels in plants. In this thesis work, the focus was placed on the development of new glucodiversification routes of natural flavonoids by taking advantage of protein engineering. Two biochemically and structurally characterized recombinant transglucosylases, the amylosucrase from Neisseria polysaccharea and the α-(1→2) branching sucrase, a truncated form of the dextransucrase from L. Mesenteroides NRRL B-1299, were selected to attempt glucosylation of different flavonoids, synthesize new α-glucoside derivatives with original patterns of glucosylation and hopefully improved their water-solubility.