Temporal Epigenomic Profiling Identifies AHR and GLIS1 As Super
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Transcriptomic Analysis of Pluripotent Stem Cells: Insights Into Health and Disease Jia-Chi Yeo1,2 and Huck-Hui Ng1,2,3,4,5,*
Yeo and Ng Genome Medicine 2011, 3:68 http://genomemedicine.com/content/3/10/68 REVIEW Transcriptomic analysis of pluripotent stem cells: insights into health and disease Jia-Chi Yeo1,2 and Huck-Hui Ng1,2,3,4,5,* Abstract types, termed ‘pluripotency’, allows researchers to study early mammalian development in an artificial setting and Embryonic stem cells (ESCs) and induced pluripotent offers opportunities for regenerative medicine, whereby stem cells (iPSCs) hold tremendous clinical potential ESCs could generate clinically relevant cell types for because of their ability to self-renew, and to tissue repair. However, this same malleability of ESCs dierentiate into all cell types of the body. This unique also renders it a challenge to obtain in vitro differentiation capacity of ESCs and iPSCs to form all cell lineages of ESCs to specific cell types at high efficacy. erefore, is termed pluripotency. While ESCs and iPSCs are harnessing the full potential of ESCs requires an in-depth pluripotent and remarkably similar in appearance, understanding of the factors and mechanisms regulating whether iPSCs truly resemble ESCs at the molecular ESC pluripotency and cell lineage decisions. level is still being debated. Further research is therefore Early studies on ESCs led to the discovery of the core needed to resolve this issue before iPSCs may be safely pluripotency factors Oct4, Sox2 and Nanog [1], and, applied in humans for cell therapy or regenerative increasingly, the use of genome-level screening assays has medicine. Nevertheless, the use of iPSCs as an in vitro revealed new insights by uncovering additional trans- human genetic disease model has been useful in cription factors, transcriptional cofactors and chromatin studying the molecular pathology of complex genetic remodeling complexes involved in the maintenance of diseases, as well as facilitating genetic or drug screens. -

KLF2 Induced
UvA-DARE (Digital Academic Repository) The transcription factor KLF2 in vascular biology Boon, R.A. Publication date 2008 Link to publication Citation for published version (APA): Boon, R. A. (2008). The transcription factor KLF2 in vascular biology. General rights It is not permitted to download or to forward/distribute the text or part of it without the consent of the author(s) and/or copyright holder(s), other than for strictly personal, individual use, unless the work is under an open content license (like Creative Commons). Disclaimer/Complaints regulations If you believe that digital publication of certain material infringes any of your rights or (privacy) interests, please let the Library know, stating your reasons. In case of a legitimate complaint, the Library will make the material inaccessible and/or remove it from the website. Please Ask the Library: https://uba.uva.nl/en/contact, or a letter to: Library of the University of Amsterdam, Secretariat, Singel 425, 1012 WP Amsterdam, The Netherlands. You will be contacted as soon as possible. UvA-DARE is a service provided by the library of the University of Amsterdam (https://dare.uva.nl) Download date:23 Sep 2021 Supplementary data: Genes induced by KLF2 Dekker et al. LocusLink Accession Gene Sequence Description Fold p-value ID number symbol change (FDR) 6654 AK022099 SOS1 cDNA FLJ12037 fis, clone HEMBB1001921. 100.00 5.9E-09 56999 AF086069 ADAMTS9 full length insert cDNA clone YZ35C05. 100.00 1.2E-09 6672 AF085934 SP100 full length insert cDNA clone YR57D07. 100.00 6.7E-13 9031 AF132602 BAZ1B Williams Syndrome critical region WS25 mRNA, partial sequence. -

Homeobox Gene Expression Profile in Human Hematopoietic Multipotent
Leukemia (2003) 17, 1157–1163 & 2003 Nature Publishing Group All rights reserved 0887-6924/03 $25.00 www.nature.com/leu Homeobox gene expression profile in human hematopoietic multipotent stem cells and T-cell progenitors: implications for human T-cell development T Taghon1, K Thys1, M De Smedt1, F Weerkamp2, FJT Staal2, J Plum1 and G Leclercq1 1Department of Clinical Chemistry, Microbiology and Immunology, Ghent University Hospital, Ghent, Belgium; and 2Department of Immunology, Erasmus Medical Center, Rotterdam, The Netherlands Class I homeobox (HOX) genes comprise a large family of implicated in this transformation proces.14 The HOX-C locus transcription factors that have been implicated in normal and has been primarily implicated in lymphomas.15 malignant hematopoiesis. However, data on their expression or function during T-cell development is limited. Using degener- Hematopoietic cells are derived from stem cells that reside in ated RT-PCR and Affymetrix microarray analysis, we analyzed fetal liver (FL) in the embryo and in the adult bone marrow the expression pattern of this gene family in human multipotent (ABM), which have the unique ability to self-renew and thereby stem cells from fetal liver (FL) and adult bone marrow (ABM), provide a life-long supply of blood cells. T lymphocytes are a and in T-cell progenitors from child thymus. We show that FL specific type of hematopoietic cells that play a major role in the and ABM stem cells are similar in terms of HOX gene immune system. They develop through a well-defined order of expression, but significant differences were observed between differentiation steps in the thymus.16 Several transcription these two cell types and child thymocytes. -

Angiogenic Patterning by STEEL, an Endothelial-Enriched Long
Angiogenic patterning by STEEL, an endothelial- enriched long noncoding RNA H. S. Jeffrey Mana,b, Aravin N. Sukumara,b, Gabrielle C. Lamc,d, Paul J. Turgeonb,e, Matthew S. Yanb,f, Kyung Ha Kub,e, Michelle K. Dubinskya,b, J. J. David Hob,f, Jenny Jing Wangb,e, Sunit Dasg,h, Nora Mitchelli, Peter Oettgeni, Michael V. Seftonc,d,j, and Philip A. Marsdena,b,e,f,1 aInstitute of Medical Science, University of Toronto, Toronto, ON M5S 1A8, Canada; bKeenan Research Centre for Biomedical Science in the Li Ka Shing Knowledge Institute, St. Michael’s Hospital, University of Toronto, Toronto, ON M5B 1T8, Canada; cDonnelly Centre for Cellular and Biomolecular Research, University of Toronto, Toronto, ON M5S 3E2, Canada; dInstitute of Biomaterials and Biomedical Engineering, University of Toronto, Toronto, ON M5S 3G9, Canada; eDepartment of Laboratory Medicine and Pathobiology, University of Toronto, Toronto, ON M5S 1A8, Canada; fDepartment of Medical Biophysics, University of Toronto, Toronto, ON M5G 1L7, Canada; gArthur and Sonia Labatt Brain Tumour Research Institute, Hospital for SickKids, University of Toronto, Toronto, ON M5G 1X8, Canada; hDivision of Neurosurgery and Keenan Research Centre for Biomedical Science, St. Michael’s Hospital, University of Toronto, Toronto, ON M5B 1W8, Canada; iDepartment of Medicine, Beth Israel Deaconess Medical Center, Harvard Medical School, Boston, MA 02115; and jDepartment of Chemical Engineering and Applied Chemistry, University of Toronto, Toronto, ON M5S 3E5, Canada Edited by Napoleone Ferrara, University of California, San Diego, La Jolla, CA, and approved January 24, 2018 (received for review August 28, 2017) Endothelial cell (EC)-enriched protein coding genes, such as endothelial formation in vitro and blood vessel formation in vivo. -

The Gli2 Transcription Factor Is Required for Normal Mouse Mammary Gland Development
Developmental Biology 238, 133–144 (2001) doi:10.1006/dbio.2001.0410, available online at http://www.idealibrary.com on The Gli2 Transcription Factor Is Required for Normal Mouse Mammary Gland Development Michael T. Lewis,*,1,2 Sarajane Ross,*,3 Phyllis A. Strickland,* Charles W. Sugnet,* Elsa Jimenez,* Chi-chung Hui,† and Charles W. Daniel* *Department of Biology, Sinsheimer Laboratories, University of California, Santa Cruz, California 95064; and †The Hospital for Sick Children, 555 University Avenue, Toronto, Ontario M5G 1X8, Canada The hedgehog signal transduction network performs critical roles in mediating cell–cell interactions during embryogenesis and organogenesis. Loss-of-function or misexpression mutation of hedgehog network components can cause birth defects, skin cancer, and other tumors. The Gli gene family (Gli1, Gli2, and Gli3) encodes zinc finger transcription factors that act as mediators of hedgehog signal transduction. In this study, we investigate the role of Gli2 in mammary gland development. Mammary expression of Gli2 is developmentally regulated in a tissue compartment-specific manner. Expression is exclusively stromal during virgin stages of development but becomes both epithelial and stromal during pregnancy and lactation. The null phenotype with respect to both ductal and alveolar development was examined by transplantation rescue of embryonic mammary glands into physiologically normal host females. Glands derived from both wild type and null embryo donors showed ductal outgrowths that developed to equivalent extents in virgin hosts. However, in null transplants, ducts were frequently distended or irregularly shaped and showed a range of histological alterations similar to micropap- illary ductal hyperplasias in the human breast. Alveolar development during pregnancy was not overtly affected by loss of Gli2 function. -

Supplemental Figure 1 Supplemental Figure 1. Flow Cytometry
Supplemental Figure 1 Initial neutrophil isolation sample Single cells Live 83.4% 99.6% 99.1% W - SC SC S S FSC FSC FSC-H Live/ dead CD45 positive 100% SC S CD16 PE cy7 CD16 CD45+ CD16+ CD66b+ neutrophils 98.1% CD45 V500 CD16 PerCP cy5.5 Supplemental Figure 1. Flow cytometry with gating strategey depicted confirms 98.1% purity of CD66b/CD16 double positive neutrophils. Supplemental Figure 1. Flow cytometry with gating strategy depicted confirms 98.1% purity of CD66b/CD16 double positive neutrophils. (a) (b) Supplemental Figure 2 A. B. (c) (d) -2000 bp 0 2000bp Position relative to TSS Supplemental Fig 2. a) No response to CpG in qRTPCR. b) Healthy volunteer neutrophils do not produce NETs via sytox green assay in response to pathogen ligands at 1 hour or immediately following live organism challenge supporting this time point for ATAC-seq. (PMA is a positive control)Supplemental(ligand donors Fign=4ure, live 2organism. A. Representativedonors n=2) c) Representative QC plots demonstratingQC plots demonstrating librarylibrary prepprep resultsresults inin expected insert size expecteddistribution and insertd) reads sizeare distributionenriched around andtranscription B. readsstart aresites enriched(TSS). around transcription start sites (TSS). Supplemental Figure 3. LTA BGP LPS HMGB1 FLAG E. coli R848 S. aureus Supplemental Fig 3. Quality control for DiffBind method of identification of differentially accessible regions of chromatin. Correlation heat maps and principal component analysis (PCA) of differentially accessible chromatin. We found that for any given challenge across donors, stimulated samples cluster together, control samples cluster Supplementaltogether, and the stimulated Figandurecontrol 3.cluster Qualityaway from controleach other, forsuggesting DiffBindhigh quality methoddata and accessibleof identificationchromatin region ofidentification differentiallythat allows for analysis accessibleof four healthy donor regionsdata. -

Effects of Downregulating GLIS1 Transcript On
Journal of Reproduction and Development, Vol. 61, No 5, 2015 —Original Article— Effects of downregulating GLIS1 transcript on preimplantation development and gene expression of bovine embryos Kazuki TAKAHASHI1), Nobuyuki SAKURAI2), Natsuko EMURA1), Tsutomu HASHIZUME1, 2) and Ken SAWAI1, 2) 1)Faculty of Agriculture, Iwate University, Iwate 020-8550, Japan 2)The United Graduate School of Agricultural Sciences, Iwate University, Iwate 020-8550, Japan Abstract. Krüppel-like protein Gli-similar 1 (GLIS1) is known as a direct reprogramming factor for the generation of induced pluripotent stem cells. The objective of this study was to investigate the role of GLIS1 in the preimplantation development of bovine embryos. GLIS1 transcripts in in vitro-matured oocytes and 1-cell to 4-cell stage embryos were detected, but they were either absent or at trace levels at the 8-cell to blastocyst stages. We attempted GLIS1 downregulation of bovine early embryos by RNA interference and evaluated developmental competency and gene transcripts, which are involved in zygotic gene activation (ZGA) in GLIS1-downregulated embryos. Injection of specific siRNA resulted in a distinct decrease inGLIS1 transcript in bovine embryos at the 4-cell stage. Although the bovine embryos injected with GLIS1-siRNA could develop to the 16-cell stage, these embryos had difficulty in developing beyond the 32-cell stage. Gene transcripts of PDHA1 and HSPA8, which are transcribed after ZGA, showed lower level in GLIS1 downregulated embryos. It is possible that GLIS1- downregulated embryos fail to initiate ZGA. Our results indicated that GLIS1 is an important factor for the preimplantation development of bovine embryos. Key words: Bovine embryo, Early development, Gene expression, GLIS1, RNA interference (J. -

Drosophila Pax6 Promotes Development of the Entire Eye-Antennal Disc, Thereby Ensuring Proper Adult Head Formation
PAPER Drosophila Pax6 promotes development of the entire COLLOQUIUM eye-antennal disc, thereby ensuring proper adult head formation Jinjin Zhua, Sneha Palliyila, Chen Ranb, and Justin P. Kumara,1 aDepartment of Biology, Indiana University, Bloomington, IN 47405; and bDepartment of Biology, Stanford University, Stanford, CA 94305 Edited by Ellen V. Rothenberg, California Institute of Technology, Pasadena, CA, and accepted by Editorial Board Member Neil H. Shubin February 17, 2017 (received for review July 26, 2016) Paired box 6 (Pax6) is considered to be the master control gene for molecular battle among GRNs allows for the subdivision of the eye development in all seeing animals studied so far. In vertebrates, eye-antennal disc to be maintained within a single continuous it is required not only for lens/retina formation but also for the cellular field (13–16). Of the GRNs that are known to operate development of the CNS, olfactory system, and pancreas. Although within the eye-antennal disc, the retinal determination (RD) Pax6 plays important roles in cell differentiation, proliferation, and network, which controls eye development, is the best studied (17). patterning during the development of these systems, the underlying At the core of the RD network lie the Paired box 6 (Pax6) genes mechanism remains poorly understood. In the fruit fly, Drosophila eyeless (ey)andtwin of eyeless (toy), the SIX family member sine melanogaster, Pax6 also functions in a range of tissues, including oculis (so), the transcriptional coactivator eyes absent (eya), and the the eye and brain. In this report, we describe the function of Pax6 in Ski/Sno family member dachshund (dac)(17). -
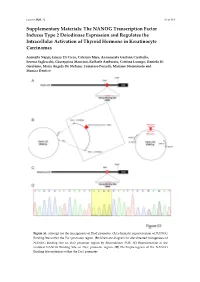
The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas
Cancers 2020, 12 S1 of S18 Supplementary Materials: The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas Annarita Nappi, Emery Di Cicco, Caterina Miro, Annunziata Gaetana Cicatiello, Serena Sagliocchi, Giuseppina Mancino, Raffaele Ambrosio, Cristina Luongo, Daniela Di Girolamo, Maria Angela De Stefano, Tommaso Porcelli, Mariano Stornaiuolo and Monica Dentice Figure S1. Strategy for the mutagenesis of Dio2 promoter. (A) Schematic representation of NANOG Binding Site within the Dio2 promoter region. (B) Schematic diagram for site‐directed mutagenesis of NANOG Binding Site on Dio2 promoter region by Recombinant PCR. (C) Representation of the mutated NANOG Binding Site on Dio2 promoter region. (D) Electropherogram of the NANOG Binding Site mutation within the Dio2 promoter. Cancers 2020, 12 S2 of S18 Figure S2. Strategy for the silencing of NANOG expression. (A) Cloning strategies for the generation of NANOG shRNA expression vectors. (B) Electropherograms of the NANOG shRNA sequences cloned into pcDNA3.1 vector. (C) Validation of effective NANOG down-modulation by two different NANOG shRNA vectors was assessed by Western Blot analysis of NANOG expression in BCC cells. (D) Quantification of NANOG protein levels versus Tubulin levels in the same experiment as in C is represented by histograms. Cancers 2020, 12 S3 of S18 Figure S3. The CD34+ cells are characterized by the expression of typical epithelial stemness genes. The mRNA levels of a panel of indicated stemness markers of epidermis were measured by Real Time PCR in the same experiment indicated in figure 3F and G. Cancers 2020, 12 S4 of S18 Figure S4. -

TP53 9606.ENSP00000269305 Tumor Protein P53; Acts As a Tumor
TP53 9606.ENSP00000269305 tumor protein p53; Acts as a tumor suppressor in many tumor types; induces growth arrest or apoptosis depending on the physiological circumstances and cell type. Involved in cell cycle regulation as a trans-activator that acts to negatively regulate cell division by controlling a set of genes required for this process. One of the activated genes is an inhibitor of cyclin-dependent kinases. Apoptosis induction seems to be mediated either by stimulation of BAX and FAS antigen expression, or by repression of Bcl-2 expression. Implicated in Notch signaling cross-over TAF1 9606.ENSP00000276072 TAF1 RNA polymerase II, TATA box binding protein (TBP)-associated factor, 250kDa; Largest component and core scaffold of the TFIID basal transcription factor complex. Contains novel N- and C-terminal Ser/Thr kinase domains which can autophosphorylate or transphosphorylate other transcription factors. Phosphorylates TP53 on 'Thr-55' which leads to MDM2-mediated degradation of TP53. Phosphorylates GTF2A1 and GTF2F1 on Ser residues. Possesses DNA- binding activity. Essential for progression of the G1 phase of the cell cycle SUMO2 9606.ENSP00000405965 SMT3 suppressor of mif two 3 homolog 2 (S. cerevisiae); Ubiquitin-like protein which can be covalently attached to target lysines either as a monomer or as a lysine-linked polymer. Does not seem to be involved in protein degradation and may function as an antagonist of ubiquitin in the degradation process. Plays a role in a number of cellular processes such as nuclear transport, DNA -

The Impact of Transcription Factor Prospero Homeobox 1 on the Regulation of Thyroid Cancer Malignancy
International Journal of Molecular Sciences Review The Impact of Transcription Factor Prospero Homeobox 1 on the Regulation of Thyroid Cancer Malignancy Magdalena Rudzi ´nska 1,2 and Barbara Czarnocka 1,* 1 Department of Biochemistry and Molecular Biology, Centre of Postgraduate Medical Education, 01-813 Warsaw, Poland; [email protected] 2 Institute of Molecular Medicine, Sechenov First Moscow State Medical University, 119991 Moscow, Russia * Correspondence: [email protected]; Tel.: +48-225693812; Fax: +48-225693712 Received: 7 April 2020; Accepted: 30 April 2020; Published: 2 May 2020 Abstract: Transcription factor Prospero homeobox 1 (PROX1) is continuously expressed in the lymphatic endothelial cells, playing an essential role in their differentiation. Many reports have shown that PROX1 is implicated in cancer development and acts as an oncoprotein or suppressor in a tissue-dependent manner. Additionally, the PROX1 expression in many types of tumors has prognostic significance and is associated with patient outcomes. In our previous experimental studies, we showed that PROX1 is present in the thyroid cancer (THC) cells of different origins and has a high impact on follicular thyroid cancer (FTC) phenotypes, regulating migration, invasion, focal adhesion, cytoskeleton reorganization, and angiogenesis. Herein, we discuss the PROX1 transcript and protein structures, the expression pattern of PROX1 in THC specimens, and its epigenetic regulation. Next, we emphasize the biological processes and genes regulated by PROX1 in CGTH-W-1 cells, derived from squamous cell carcinoma of the thyroid gland. Finally, we discuss the interaction of PROX1 with other lymphatic factors. In our review, we aimed to highlight the importance of vascular molecules in cancer development and provide an update on the functionality of PROX1 in THC biology regulation. -

DLX Genes: Roles in Development and Cancer
cancers Review DLX Genes: Roles in Development and Cancer Yinfei Tan 1,* and Joseph R. Testa 1,2,* 1 Genomics Facility, Fox Chase Cancer Center, Philadelphia, PA 19111, USA 2 Cancer Signaling and Epigenetics Program, Fox Chase Cancer Center, Philadelphia, PA 19111, USA * Correspondence: [email protected] (Y.T.); [email protected] (J.R.T.) Simple Summary: DLX homeobox family genes encode transcription factors that have indispensable roles in embryonic and postnatal development. These genes are critically linked to the morphogene- sis of craniofacial structures, branchial arches, forebrain, and sensory organs. DLX genes are also involved in postnatal homeostasis, particularly hematopoiesis and, when dysregulated, oncogen- esis. DLX1/2, DLX3/4, and DLX5/6 exist as bigenes on different chromosomes, sharing intergenic enhancers between gene pairs, which allows orchestrated spatiotemporal expression. Genomic alterations of human DLX gene enhancers or coding sequences result in congenital disorders such as split-hand/foot malformation. Aberrant postnatal expression of DLX genes is associated with hematological malignancies, including leukemias and lymphomas. In several mouse models of T-cell lymphoma, Dlx5 has been shown to act as an oncogene by cooperating with activated Akt, Notch1/3, and/or Wnt to drive tumor formation. In humans, DLX5 is aberrantly expressed in lung and ovarian carcinomas and holds promise as a therapeutic target. Abstract: Homeobox genes control body patterning and cell-fate decisions during development. The homeobox genes consist of many families, only some of which have been investigated regarding a possible role in tumorigenesis. Dysregulation of HOX family genes have been widely implicated in cancer etiology.