CNTNAP2) Reveals the Presence of Two Caspr2 Isoforms with Overlapping Interactomes☆
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Likely Pathogenic Variants in One Third of Non-Syndromic Discontinuous Cleft Lip and Palate Patients
G C A T T A C G G C A T genes Article Likely Pathogenic Variants in One Third of Non-Syndromic Discontinuous Cleft Lip and Palate Patients Bénédicte Demeer 1,2,3,4 , Nicole Revencu 1,5, Raphael Helaers 1 , Cica Gbaguidi 6, Stéphanie Dakpe 3,4,6 , Geneviève François 7, Bernard Devauchelle 3,4,6,Bénédicte Bayet 8 and Miikka Vikkula 1,* 1 Human Molecular Genetics, de Duve Institute, University of Louvain, 1200 Brussels, Belgium; [email protected] (B.D.); [email protected] (N.R.); [email protected] (R.H.) 2 Center for Human Genetics, CLAD Nord de France, CHU Amiens-Picardie, 80054 Amiens, France 3 Université Picardie Jules Verne, EA CHIMERE, EA 7516, 80054 Amiens, France; [email protected] (S.D.); [email protected] (B.D.) 4 Facing Faces Institute, 80054 Amiens, France 5 Center for Human Genetics, Cliniques universitaires Saint-Luc, University of Louvain, 1200 Brussels, Belgium 6 Department of Maxillofacial Surgery and Stomatology, Centre de Compétence Fentes et Malformations Faciales (MAFACE), CHU Amiens-Picardie, 80054 Amiens, France; [email protected] 7 Department of Pediatrics, Cliniques Universitaires Saint-Luc, University of Louvain, 1200 Brussels, Belgium; [email protected] 8 Centre Labiopalatin, Division of Plastic Surgery, Cliniques Universitaires Saint-Luc, University of Louvain, 1200 Brussels, Belgium; [email protected] * Correspondence: [email protected]; Tel.: +32-2-764-7490 Received: 6 September 2019; Accepted: 19 October 2019; Published: 22 October 2019 Abstract: Oral clefts are composed of cleft of the lip, cleft of the lip and palate, or cleft of the palate, and they are associated with a wide range of expression and severity. -

Chr21 Protein-Protein Interactions: Enrichment in Products Involved in Intellectual Disabilities, Autism and Late Onset Alzheimer Disease
bioRxiv preprint doi: https://doi.org/10.1101/2019.12.11.872606; this version posted December 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Chr21 protein-protein interactions: enrichment in products involved in intellectual disabilities, autism and Late Onset Alzheimer Disease Julia Viard1,2*, Yann Loe-Mie1*, Rachel Daudin1, Malik Khelfaoui1, Christine Plancon2, Anne Boland2, Francisco Tejedor3, Richard L. Huganir4, Eunjoon Kim5, Makoto Kinoshita6, Guofa Liu7, Volker Haucke8, Thomas Moncion9, Eugene Yu10, Valérie Hindie9, Henri Bléhaut11, Clotilde Mircher12, Yann Herault13,14,15,16,17, Jean-François Deleuze2, Jean- Christophe Rain9, Michel Simonneau1, 18, 19, 20** and Aude-Marie Lepagnol- Bestel1** 1 Centre Psychiatrie & Neurosciences, INSERM U894, 75014 Paris, France 2 Laboratoire de génomique fonctionnelle, CNG, CEA, Evry 3 Instituto de Neurociencias CSIC-UMH, Universidad Miguel Hernandez-Campus de San Juan 03550 San Juan (Alicante), Spain 4 Department of Neuroscience, The Johns Hopkins University School of Medicine, Baltimore, MD 21205 USA 5 Center for Synaptic Brain Dysfunctions, Institute for Basic Science, Daejeon 34141, Republic of Korea 6 Department of Molecular Biology, Division of Biological Science, Nagoya University Graduate School of Science, Furo, Chikusa, Nagoya, Japan 7 Department of Biological Sciences, University of Toledo, Toledo, OH, 43606, USA 8 Leibniz Forschungsinstitut für Molekulare Pharmakologie -

Neural Stem Cell-Derived Exosomes Revert HFD-Dependent Memory Impairment Via CREB-BDNF Signalling
International Journal of Molecular Sciences Article Neural Stem Cell-Derived Exosomes Revert HFD-Dependent Memory Impairment via CREB-BDNF Signalling Matteo Spinelli 1, Francesca Natale 1,2, Marco Rinaudo 1, Lucia Leone 1,2, Daniele Mezzogori 1, Salvatore Fusco 1,2,* and Claudio Grassi 1,2 1 Department of Neuroscience, Università Cattolica del Sacro Cuore, 00168 Rome, Italy; [email protected] (M.S.); [email protected] (F.N.); [email protected] (M.R.); [email protected] (L.L.); [email protected] (D.M.); [email protected] (C.G.) 2 Fondazione Policlinico Universitario Agostino Gemelli IRCCS, 00168 Rome, Italy * Correspondence: [email protected] Received: 6 October 2020; Accepted: 25 November 2020; Published: 26 November 2020 Abstract: Overnutrition and metabolic disorders impair cognitive functions through molecular mechanisms still poorly understood. In mice fed with a high fat diet (HFD) we analysed the expression of synaptic plasticity-related genes and the activation of cAMP response element-binding protein (CREB)-brain-derived neurotrophic factor (BDNF)-tropomyosin receptor kinase B (TrkB) signalling. We found that a HFD inhibited both CREB phosphorylation and the expression of a set of CREB target genes in the hippocampus. The intranasal administration of neural stem cell (NSC)-derived exosomes (exo-NSC) epigenetically restored the transcription of Bdnf, nNOS, Sirt1, Egr3, and RelA genes by inducing the recruitment of CREB on their regulatory sequences. Finally, exo-NSC administration rescued both BDNF signalling and memory in HFD mice. Collectively, our findings highlight novel mechanisms underlying HFD-related memory impairment and provide evidence of the potential therapeutic effect of exo-NSC against metabolic disease-related cognitive decline. -

Clinical Application of Chromosomal Microarray Analysis for Fetuses With
Xu et al. Molecular Cytogenetics (2020) 13:38 https://doi.org/10.1186/s13039-020-00502-5 RESEARCH Open Access Clinical application of chromosomal microarray analysis for fetuses with craniofacial malformations Chenyang Xu1†, Yanbao Xiang1†, Xueqin Xu1, Lili Zhou1, Huanzheng Li1, Xueqin Dong1 and Shaohua Tang1,2* Abstract Background: The potential correlations between chromosomal abnormalities and craniofacial malformations (CFMs) remain a challenge in prenatal diagnosis. This study aimed to evaluate 118 fetuses with CFMs by applying chromosomal microarray analysis (CMA) and G-banded chromosome analysis. Results: Of the 118 cases in this study, 39.8% were isolated CFMs (47/118) whereas 60.2% were non-isolated CFMs (71/118). The detection rate of chromosomal abnormalities in non-isolated CFM fetuses was significantly higher than that in isolated CFM fetuses (26/71 vs. 7/47, p = 0.01). Compared to the 16 fetuses (16/104; 15.4%) with pathogenic chromosomal abnormalities detected by karyotype analysis, CMA identified a total of 33 fetuses (33/118; 28.0%) with clinically significant findings. These 33 fetuses included cases with aneuploidy abnormalities (14/118; 11.9%), microdeletion/microduplication syndromes (9/118; 7.6%), and other pathogenic copy number variations (CNVs) only (10/118; 8.5%).We further explored the CNV/phenotype correlation and found a series of clear or suspected dosage-sensitive CFM genes including TBX1, MAPK1, PCYT1A, DLG1, LHX1, SHH, SF3B4, FOXC1, ZIC2, CREBBP, SNRPB, and CSNK2A1. Conclusion: These findings enrich our understanding of the potential causative CNVs and genes in CFMs. Identification of the genetic basis of CFMs contributes to our understanding of their pathogenesis and allows detailed genetic counselling. -

Supplemental Information
Supplemental information Dissection of the genomic structure of the miR-183/96/182 gene. Previously, we showed that the miR-183/96/182 cluster is an intergenic miRNA cluster, located in a ~60-kb interval between the genes encoding nuclear respiratory factor-1 (Nrf1) and ubiquitin-conjugating enzyme E2H (Ube2h) on mouse chr6qA3.3 (1). To start to uncover the genomic structure of the miR- 183/96/182 gene, we first studied genomic features around miR-183/96/182 in the UCSC genome browser (http://genome.UCSC.edu/), and identified two CpG islands 3.4-6.5 kb 5’ of pre-miR-183, the most 5’ miRNA of the cluster (Fig. 1A; Fig. S1 and Seq. S1). A cDNA clone, AK044220, located at 3.2-4.6 kb 5’ to pre-miR-183, encompasses the second CpG island (Fig. 1A; Fig. S1). We hypothesized that this cDNA clone was derived from 5’ exon(s) of the primary transcript of the miR-183/96/182 gene, as CpG islands are often associated with promoters (2). Supporting this hypothesis, multiple expressed sequences detected by gene-trap clones, including clone D016D06 (3, 4), were co-localized with the cDNA clone AK044220 (Fig. 1A; Fig. S1). Clone D016D06, deposited by the German GeneTrap Consortium (GGTC) (http://tikus.gsf.de) (3, 4), was derived from insertion of a retroviral construct, rFlpROSAβgeo in 129S2 ES cells (Fig. 1A and C). The rFlpROSAβgeo construct carries a promoterless reporter gene, the β−geo cassette - an in-frame fusion of the β-galactosidase and neomycin resistance (Neor) gene (5), with a splicing acceptor (SA) immediately upstream, and a polyA signal downstream of the β−geo cassette (Fig. -

Downloads/ (Accessed on 17 January 2020)
cells Review Novel Approaches for Identifying the Molecular Background of Schizophrenia Arkadiy K. Golov 1,2,*, Nikolay V. Kondratyev 1 , George P. Kostyuk 3 and Vera E. Golimbet 1 1 Mental Health Research Center, 34 Kashirskoye shosse, 115522 Moscow, Russian; [email protected] (N.V.K.); [email protected] (V.E.G.) 2 Institute of Gene Biology, Russian Academy of Sciences, 34/5 Vavilova Street, 119334 Moscow, Russian 3 Alekseev Psychiatric Clinical Hospital No. 1, 2 Zagorodnoye shosse, 115191 Moscow, Russian; [email protected] * Correspondence: [email protected] Received: 5 November 2019; Accepted: 16 January 2020; Published: 18 January 2020 Abstract: Recent advances in psychiatric genetics have led to the discovery of dozens of genomic loci associated with schizophrenia. However, a gap exists between the detection of genetic associations and understanding the underlying molecular mechanisms. This review describes the basic approaches used in the so-called post-GWAS studies to generate biological interpretation of the existing population genetic data, including both molecular (creation and analysis of knockout animals, exploration of the transcriptional effects of common variants in human brain cells) and computational (fine-mapping of causal variability, gene set enrichment analysis, partitioned heritability analysis) methods. The results of the crucial studies, in which these approaches were used to uncover the molecular and neurobiological basis of the disease, are also reported. Keywords: schizophrenia; GWAS; causal genetic variants; enhancers; brain epigenomics; genome/epigenome editing 1. Introduction Schizophrenia is a severe mental illness that affects between 0.5% and 0.7% of the human population [1]. Both environmental and genetic factors are thought to be involved in its pathogenesis, with genetic factors playing a key role in disease risk, as the heritability of schizophrenia is estimated to be 70–85% [2,3]. -

MYRF Is a Membrane-Associated Transcription Factor That Autoproteolytically Cleaves to Directly Activate Myelin Genes
MYRF Is a Membrane-Associated Transcription Factor That Autoproteolytically Cleaves to Directly Activate Myelin Genes Helena Bujalka1., Matthias Koenning1., Stacey Jackson1, Victoria M. Perreau1, Bernard Pope2, Curtis M. Hay1, Stanlislaw Mitew1, Andrew F. Hill3, Q. Richard Lu4, Michael Wegner5, Rajini Srinivasan6, John Svaren6, Melanie Willingham1, Ben A. Barres7, Ben Emery1,8* 1 Department of Anatomy and Neuroscience and the Centre for Neuroscience Research, University of Melbourne, Parkville, Australia, 2 Department of Computing and Information Systems and Victorian Life Sciences Computation Initiative (VLSCI), University of Melbourne, Parkville, Australia, 3 Department of Biochemistry and Molecular Biology, Bio21 Molecular Science and Biotechnology Institute, University of Melbourne, Parkville, Australia, 4 Department of Developmental Biology and Kent Waldrep Foundation Center for Basic Neuroscience Research on Nerve Growth and Regeneration, University of Texas at Austin, Austin, Texas, United States of America, 5 Institute for Biochemistry, University Erlangen-Nuernberg, Germany, 6 Waisman Center, University of Wisconsin–Madison, Madison, Wisconsin, United States of America, 7 Department of Neurobiology, Stanford University, Stanford, California, United States of America, 8 Florey Institute of Neuroscience and Mental Health, Parkville, Australia Abstract The myelination of axons is a crucial step during vertebrate central nervous system (CNS) development, allowing for rapid and energy efficient saltatory conduction of nerve impulses. Accordingly, the differentiation of oligodendrocytes, the myelinating cells of the CNS, and their expression of myelin genes are under tight transcriptional control. We previously identified a putative transcription factor, Myelin Regulatory Factor (Myrf), as being vital for CNS myelination. Myrf is required for the generation of CNS myelination during development and also for its maintenance in the adult. -

BDNF Integrity in Ageing and Stress
MOJ Gerontology & Geriatrics Editorial Open Access BDNF integrity in ageing and stress Editorial Volume 1 Issue 6 - 2017 Alzheimer’s disease (AD) presents a progressive, stage-dependent Trevor Archer age-related neurodegenerative disorder that is characterized by Department of Psychology, University of Gothenburg, Sweden aggregation of toxic forms of amyloid β peptide (Aβ). There is much support for the notion that that brain-derived neurotrophic factor Correspondence: Trevor Archer, Department of Psychology, (BDNF) mediates beneficial effects of exercise on neuroplasticity and University of Gothenburg, Sweden, cellular stress resistance.1‒3 BDNF is associated with neuroplasticity Email [email protected] changes promoting health and well-being whereas these influences Received: July 16, 2017 | Published: August 01, 2017 may be opposed by pro-inflammatory cytokines, key factors in neurodegenerative processes.4 Lower serum levels are linked to greater symptom, cognitive, impairment.5 Thus, interventions, as for example physical exercise, whether acute or chronic, endurance or resistance, promote the mobilization of BDNF and other neurotrophic factors6,7 increase hippocampal and other brain regional resilience8 and therewith progression under conditions of stress when confronted critical life protect against cell death by inducing cell proliferation and maturation episodes or periods, appears to be modulated by the availability and with enhanced neuronal reparation, neurogenesis and the growth and accessibility of BDNF.20‒22 Over a wide range of neurologic and functioning of neurons in neurodegenerative disorders.9,10 Sampaio et psychiatric disease states it is found that the integrity and concentrations al.11 in a review of the therapeutic applications of neurotrophic factors, of brain, serum and plasma BDNF is compromised, particularly under particularly BDNF, postulate that they are essential for survival, stressful conditions. -

Identification of Differentially Expressed Genes in Urinary Bladder Cancer by Meta-Analysis by Using a Bioinformatics Tool Sathya Mylsamy1* and K Devaki2
Research Article iMedPub Journals Journal of Clinical & Experimental Nephrology 2019 http://www.imedpub.com/ Vol 5.No.5:95 ISSN 2472-5056 Identification of Differentially Expressed Genes in Urinary Bladder Cancer by Meta-Analysis by Using a Bioinformatics Tool Sathya Mylsamy1* and K Devaki2 1Department of Biochemistry, CMS College of Science and Commerce, Coimbatore, Tamil Nadu, India 2Department of Biochemistry, Karpagam Academy of Higher Education, Coimbatore, Tamil Nadu, India *Corresponding author: Sathya Mylsamy, CMS College of Science and Commerce, Tamil Nadu, India, Tel: (+91)7904349498; E-mail: [email protected] Received date: November 18, 2020 ; Accepted date: December 02, 2020 ; Published date: December 09, 2020 Citation: Mylsamy S, Devaki K (2020) Identification of Differentially Expressed Genes in Urinary Bladder Cancer by Meta-Analysis by using a Bioinformatics Tool. J Clin Exp Nephrol Vol.5 No.5:95. Copyright: © 2020 Mylsamy S, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. understand the mechanism behind the tumor progression and its diagnosis. Abstract Background: Bladder cancer is the ninth most prevalent Keywords: Bladder cancer; GEO; CCNB2; NUSAP1; malignant disease globally, which ranges from mild with low ADAM22; KEGG; Gene ontology mortality rate to extremely high grade tumors associated with high mortality rate. The present study was aimed to identify the key genes associated with bladder cancer Introduction progression and later it may also be used as marker in the diagnosis and prognosis. The bladder is probably one of the few sites in the body where the environmental factors play an undeniable role in Materials and methods: The GSE3167, GSE7476, GSE68928 genesis of cancer [1]. -
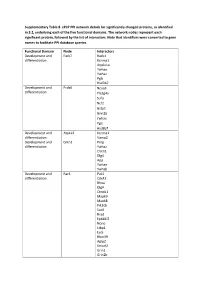
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4 -

Lgi1 (NM 145769) Rat Tagged ORF Clone – RR210565 | Origene
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RR210565 Lgi1 (NM_145769) Rat Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: Lgi1 (NM_145769) Rat Tagged ORF Clone Tag: Myc-DDK Symbol: Lgi1 Synonyms: MGC105310 Vector: pCMV6-Entry (PS100001) E. coli Selection: Kanamycin (25 ug/mL) Cell Selection: Neomycin This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 4 Lgi1 (NM_145769) Rat Tagged ORF Clone – RR210565 ORF Nucleotide >RR210565 representing NM_145769 Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGGAATCAGAAAGCATCAGAAGGATGGGAAATGCCTGCATTCCCCTGAAAAGAATTGCCTATTTCCTAT GCCTCTTTTCTGTGGTTTTGCTGACTGAGGGGAAGAAACCAGCGAAGCCAAAATGCCCTGCTGTGTGTAC TTGTAGCAAAGATAACGCTTTATGTGAGAATGCGAGATCCATTCCACGCACCGTTCCTCCTGATGTTATC TCACTATCCTTTGTGAGATCTGGTTTTACTGAAATCTCAGAAGGGAGTTTCTTGTTCACACCATCGCTGC AGCTCTTGTTATTCACGTCGAACTCCTTTGATGTGATCAGTGACGATGCTTTTATTGGTCTTCCACACCT AGAATATTTATTCATAGAAAACAACAATATCAAGTCCATTTCAAGACATACTTTCCGGGGACTCAAGTCT CTGATTCACTTGAGTCTTGCAAACAACAATCTCCAGACACTCCCAAAAGACATTTTCAAAGGCCTGGATT CTTTAACAAATGTGGACCTAAGAGGGAACTCATTTAACTGTGACTGTAAACTGAAGTGGCTGGTGGAATG GCTCGGCCACACCAACGCAACCGTGGAAGACATCTACTGCGAAGGACCACCGGAGTATAAGAAACGTAAA -

Product Description P408-A2 ADLTE-LGI1-V02
MRC-Holland ® Product Description version A2-02; Issued 02 December 2020 MLPA Product Description SALSA® MLPA® Probemix P408-A2 ADLTE-LGI1 To be used with the MLPA General Protocol. Version A2. For complete product history see page 7. Catalogue numbers: • P408-025R: SALSA MLPA Probemix P408 ADLTE-LGI1, 25 reactions. • P408-050R: SALSA MLPA Probemix P408 ADLTE-LGI1, 50 reactions. • P408-100R: SALSA MLPA Probemix P408 ADLTE-LGI1, 100 reactions. To be used in combination with a SALSA MLPA reagent kit and Coffalyser.Net data analysis software. MLPA reagent kits are either provided with FAM or Cy5.0 dye-labelled PCR primer, suitable for Applied Biosystems and Beckman/SCIEX capillary sequencers, respectively (see www.mlpa.com). Certificate of Analysis: Information regarding storage conditions, quality tests, and a sample electropherogram from the current sales lot is available at www.mlpa.com. Precautions and warnings: For professional use only. Always consult the most recent product description AND the MLPA General Protocol before use: www.mlpa.com. It is the responsibility of the user to be aware of the latest scientific knowledge of the application before drawing any conclusions from findings generated with this product. General information: The SALSA MLPA Probemix P408 ADLTE-LGI1 is a research use only (RUO) assay for the detection of deletions or duplications in ADAM22, ADGRV1 (was GPR98), KCNA1, KCNA4, KCNAB1, LGI1, and PDYN genes, which are associated with autosomal dominant lateral temporal epilepsy (ADLTE). ADLTE, also known as autosomal dominant partial epilepsy with auditory features (ADPEAF), is a rare familial form of epilepsy likely arising from the lateral temporal lobe of the brain.