The Origin of Animals
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Predicted Glycosyltransferases Promote Development and Prevent Spurious Cell Clumping in the Choanoflagellate S
RESEARCH ADVANCE Predicted glycosyltransferases promote development and prevent spurious cell clumping in the choanoflagellate S. rosetta Laura A Wetzel1,2, Tera C Levin1,2, Ryan E Hulett1,2, Daniel Chan1,2, Grant A King1,2, Reef Aldayafleh1,2, David S Booth1,2, Monika Abedin Sigg1,2, Nicole King1,2* 1Department of Molecular and Cell Biology, University of California, Berkeley, Berkeley, United States; 2Howard Hughes Medical Institute, University of California, Berkeley, Berkeley, United States Abstract In a previous study we established forward genetics in the choanoflagellate Salpingoeca rosetta and found that a C-type lectin gene is required for rosette development (Levin et al., 2014). Here we report on critical improvements to genetic screens in S. rosetta while also investigating the genetic basis for rosette defect mutants in which single cells fail to develop into orderly rosettes and instead aggregate promiscuously into amorphous clumps of cells. Two of the mutants, Jumble and Couscous, mapped to lesions in genes encoding two different predicted glycosyltransferases and displayed aberrant glycosylation patterns in the basal extracellular matrix (ECM). In animals, glycosyltransferases sculpt the polysaccharide-rich ECM, regulate integrin and cadherin activity, and, when disrupted, contribute to tumorigenesis. The finding that predicted glycosyltransferases promote proper rosette development and prevent cell aggregation in S. rosetta suggests a pre-metazoan role for glycosyltransferases in regulating development and preventing abnormal tumor-like multicellularity. *For correspondence: DOI: https://doi.org/10.7554/eLife.41482.001 [email protected] Competing interests: The authors declare that no competing interests exist. Introduction Funding: See page 23 The transition to multicellularity was essential for the evolution of animals from their single celled Received: 05 September 2018 ancestors (Szathma´ry and Smith, 1995). -

The Architecture of Cell Differentiation in Choanoflagellates And
bioRxiv preprint doi: https://doi.org/10.1101/452185; this version posted October 29, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 The architecture of cell differentiation in choanoflagellates 2 and sponge choanocytes 3 Davis Laundon1,2,6, Ben Larson3, Kent McDonald3, Nicole King3,4 and Pawel 4 Burkhardt1,5* 5 1 Marine Biological Association of the United Kingdom, The Laboratory, Citadel Hill, 6 Plymouth, PL1 2PB, United Kingdom 7 2 Plymouth University, Drake Circus, Plymouth, PL4 8AA, United Kingdom 8 3 Department of Molecular and Cell Biology, University of California, Berkeley, USA 9 4 Howard Hughes Medical Institute 10 5 Sars International Centre for Molecular Marine Biology, University of Bergen, 11 Thormohlensgate 55, 5020 Bergen, Norway 12 6 Current Affiliation: University of East Anglia, Norwich, NR4 7TJ, United Kingdom 13 14 *Correspondence [email protected] 15 16 17 18 19 20 21 22 23 24 25 bioRxiv preprint doi: https://doi.org/10.1101/452185; this version posted October 29, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 26 SUMMARY 27 Collar cells are ancient animal cell types which are conserved across the animal 28 kingdom [1] and their closest relatives, the choanoflagellates [2]. -

Modularity As a Concept Modern Ideas About Mental Modularity Typically Use Fodor (1983) As a Key Touchstone
COGNITIVE PROCESSING International Quarterly of Cognitive Science Mental modularity, metaphors, and the marriage of evolutionary and cognitive sciences GARY L. BRASE University of Missouri – Columbia Abstract - As evolutionary approaches in the behavioral sciences become increasingly prominent, issues arising from the proposition that the mind is a collection of modular adaptations (the multi-modular mind thesis) become even more pressing. One purpose of this paper is to help clarify some valid issues raised by this thesis and clarify why other issues are not as critical. An aspect of the cognitive sciences that appears to both promote and impair progress on this issue (in different ways) is the use of metaphors for understand- ing the mind. Utilizing different metaphors can yield different perspectives and advancement in our understanding of the nature of the human mind. A second purpose of this paper is to outline the kindred natures of cognitive science and evolutionary psychology, both of which cut across traditional academic divisions and engage in functional analyses of problems. Key words: Evolutionary Theory, Cognitive Science, Modularity, Metaphors Evolutionary approaches in the behavioral sciences have begun to influence a wide range of fields, from cognitive neuroscience (e.g., Gazzaniga, 1998), to clini- cal psychology (e.g., Baron-Cohen, 1997; McGuire and Troisi, 1998), to literary theory (e.g., Carroll, 1999). At the same time, however, there are ongoing debates about the details of what exactly an evolutionary approach – often called evolu- tionary psychology— entails (Holcomb, 2001). Some of these debates are based on confusions of terminology, implicit arguments, or misunderstandings – things that can in principle be resolved by clarifying current ideas. -

A Six-Gene Phylogeny Provides New Insights Into Choanoflagellate Evolution Martin Carr, Daniel J
A six-gene phylogeny provides new insights into choanoflagellate evolution Martin Carr, Daniel J. Richter, Parinaz Fozouni, Timothy J. Smith, Alexandra Jeuck, Barry S.C. Leadbeater, Frank Nitsche To cite this version: Martin Carr, Daniel J. Richter, Parinaz Fozouni, Timothy J. Smith, Alexandra Jeuck, et al.. A six- gene phylogeny provides new insights into choanoflagellate evolution. Molecular Phylogenetics and Evolution, Elsevier, 2017, 107, pp.166 - 178. 10.1016/j.ympev.2016.10.011. hal-01393449 HAL Id: hal-01393449 https://hal.archives-ouvertes.fr/hal-01393449 Submitted on 7 Nov 2016 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution| 4.0 International License Molecular Phylogenetics and Evolution 107 (2017) 166–178 Contents lists available at ScienceDirect Molecular Phylogenetics and Evolution journal homepage: www.elsevier.com/locate/ympev A six-gene phylogeny provides new insights into choanoflagellate evolution ⇑ Martin Carr a, ,1, Daniel J. Richter b,1,2, Parinaz Fozouni b,3, Timothy J. Smith a, Alexandra Jeuck c, Barry S.C. Leadbeater d, Frank Nitsche c a School of Applied Sciences, University of Huddersfield, Huddersfield HD1 3DH, UK b Department of Molecular and Cell Biology, University of California, Berkeley, CA 94720-3200, USA c University of Cologne, Biocentre, General Ecology, Zuelpicher Str. -

The Closest Unicellular Relatives of Animals
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Current Biology, Vol. 12, 1773–1778, October 15, 2002, 2002 Elsevier Science Ltd. All rights reserved. PII S0960-9822(02)01187-9 The Closest Unicellular Relatives of Animals B.F. Lang,1,2 C. O’Kelly,1,3 T. Nerad,4 M.W. Gray,1,5 Results and Discussion and G. Burger1,2,6 1The Canadian Institute for Advanced Research The evolution of the Metazoa from single-celled protists Program in Evolutionary Biology is an issue that has intrigued biologists for more than a 2 De´ partement de Biochimie century. Early morphological and more recent ultra- Universite´ de Montre´ al structural and molecular studies have converged in sup- Succursale Centre-Ville porting the now widely accepted view that animals are Montre´ al, Que´ bec H3C 3J7 related to Fungi, choanoflagellates, and ichthyosporean Canada protists. However, controversy persists as to the spe- 3 Bigelow Laboratory for Ocean Sciences cific evolutionary relationships among these major P.O. Box 475 groups. This uncertainty is reflected in the plethora of 180 McKown Point Road published molecular phylogenies that propose virtually West Boothbay Harbor, Maine 04575 all of the possible alternative tree topologies involving 4 American Type Culture Collection Choanoflagellata, Fungi, Ichthyosporea, and Metazoa. 10801 University Boulevard For example, a monophyletic MetazoaϩChoanoflagel- Manassas, Virginia 20110 lata group has been suggested on the basis of small 5 Department of Biochemistry subunit (SSU) rDNA sequences [1, 6, 7]. Other studies and Molecular Biology using the same sequences have allied Choanoflagellata Dalhousie University with the Fungi [8], placed Choanoflagellata prior to the Halifax, Nova Scotia B3H 4H7 divergence of animals and Fungi [9], or even placed Canada them prior to the divergence of green algae and land plants [10]. -

Transformations of Lamarckism Vienna Series in Theoretical Biology Gerd B
Transformations of Lamarckism Vienna Series in Theoretical Biology Gerd B. M ü ller, G ü nter P. Wagner, and Werner Callebaut, editors The Evolution of Cognition , edited by Cecilia Heyes and Ludwig Huber, 2000 Origination of Organismal Form: Beyond the Gene in Development and Evolutionary Biology , edited by Gerd B. M ü ller and Stuart A. Newman, 2003 Environment, Development, and Evolution: Toward a Synthesis , edited by Brian K. Hall, Roy D. Pearson, and Gerd B. M ü ller, 2004 Evolution of Communication Systems: A Comparative Approach , edited by D. Kimbrough Oller and Ulrike Griebel, 2004 Modularity: Understanding the Development and Evolution of Natural Complex Systems , edited by Werner Callebaut and Diego Rasskin-Gutman, 2005 Compositional Evolution: The Impact of Sex, Symbiosis, and Modularity on the Gradualist Framework of Evolution , by Richard A. Watson, 2006 Biological Emergences: Evolution by Natural Experiment , by Robert G. B. Reid, 2007 Modeling Biology: Structure, Behaviors, Evolution , edited by Manfred D. Laubichler and Gerd B. M ü ller, 2007 Evolution of Communicative Flexibility: Complexity, Creativity, and Adaptability in Human and Animal Communication , edited by Kimbrough D. Oller and Ulrike Griebel, 2008 Functions in Biological and Artifi cial Worlds: Comparative Philosophical Perspectives , edited by Ulrich Krohs and Peter Kroes, 2009 Cognitive Biology: Evolutionary and Developmental Perspectives on Mind, Brain, and Behavior , edited by Luca Tommasi, Mary A. Peterson, and Lynn Nadel, 2009 Innovation in Cultural Systems: Contributions from Evolutionary Anthropology , edited by Michael J. O ’ Brien and Stephen J. Shennan, 2010 The Major Transitions in Evolution Revisited , edited by Brett Calcott and Kim Sterelny, 2011 Transformations of Lamarckism: From Subtle Fluids to Molecular Biology , edited by Snait B. -

Multigene Eukaryote Phylogeny Reveals the Likely Protozoan Ancestors of Opis- Thokonts (Animals, Fungi, Choanozoans) and Amoebozoa
Accepted Manuscript Multigene eukaryote phylogeny reveals the likely protozoan ancestors of opis- thokonts (animals, fungi, choanozoans) and Amoebozoa Thomas Cavalier-Smith, Ema E. Chao, Elizabeth A. Snell, Cédric Berney, Anna Maria Fiore-Donno, Rhodri Lewis PII: S1055-7903(14)00279-6 DOI: http://dx.doi.org/10.1016/j.ympev.2014.08.012 Reference: YMPEV 4996 To appear in: Molecular Phylogenetics and Evolution Received Date: 24 January 2014 Revised Date: 2 August 2014 Accepted Date: 11 August 2014 Please cite this article as: Cavalier-Smith, T., Chao, E.E., Snell, E.A., Berney, C., Fiore-Donno, A.M., Lewis, R., Multigene eukaryote phylogeny reveals the likely protozoan ancestors of opisthokonts (animals, fungi, choanozoans) and Amoebozoa, Molecular Phylogenetics and Evolution (2014), doi: http://dx.doi.org/10.1016/ j.ympev.2014.08.012 This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain. 1 1 Multigene eukaryote phylogeny reveals the likely protozoan ancestors of opisthokonts 2 (animals, fungi, choanozoans) and Amoebozoa 3 4 Thomas Cavalier-Smith1, Ema E. Chao1, Elizabeth A. Snell1, Cédric Berney1,2, Anna Maria 5 Fiore-Donno1,3, and Rhodri Lewis1 6 7 1Department of Zoology, University of Oxford, South Parks Road, Oxford OX1 3PS, UK. -
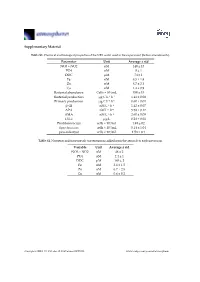
Supplementary Material Parameter Unit Average ± Std NO3 + NO2 Nm
Supplementary Material Table S1. Chemical and biological properties of the NRS water used in the experiment (before amendments). Parameter Unit Average ± std NO3 + NO2 nM 140 ± 13 PO4 nM 8 ± 1 DOC μM 74 ± 1 Fe nM 8.5 ± 1.8 Zn nM 8.7 ± 2.1 Cu nM 1.4 ± 0.9 Bacterial abundance Cells × 104/mL 350 ± 15 Bacterial production μg C L−1 h−1 1.41 ± 0.08 Primary production μg C L−1 h−1 0.60 ± 0.01 β-Gl nM L−1 h−1 1.42 ± 0.07 APA nM L−1 h−1 5.58 ± 0.17 AMA nM L−1·h−1 2.60 ± 0.09 Chl-a μg/L 0.28 ± 0.01 Prochlorococcus cells × 104/mL 1.49 ± 02 Synechococcus cells × 104/mL 5.14 ± 1.04 pico-eukaryot cells × 103/mL 1.58 × 0.1 Table S2. Nutrients and trace metals concentrations added from the aerosols to each mesocosm. Variable Unit Average ± std NO3 + NO2 nM 48 ± 2 PO4 nM 2.4 ± 1 DOC μM 165 ± 2 Fe nM 2.6 ± 1.5 Zn nM 6.7 ± 2.5 Cu nM 0.6 ± 0.2 Atmosphere 2019, 10, 358; doi:10.3390/atmos10070358 www.mdpi.com/journal/atmosphere Atmosphere 2019, 10, 358 2 of 6 Table S3. ANOVA test results between control, ‘UV-treated’ and ‘live-dust’ treatments at 20 h or 44 h, with significantly different values shown in bold. ANOVA df Sum Sq Mean Sq F Value p-value Chl-a 20 H 2, 6 0.03, 0.02 0.02, 0 4.52 0.0634 44 H 2, 6 0.02, 0 0.01, 0 23.13 0.002 Synechococcus Abundance 20 H 2, 7 8.23 × 107, 4.11 × 107 4.11 × 107, 4.51 × 107 0.91 0.4509 44 H 2, 7 5.31 × 108, 6.97 × 107 2.65 × 108, 1.16 × 107 22.84 0.0016 Prochlorococcus Abundance 20 H 2, 8 4.22 × 107, 2.11 × 107 2.11 × 107, 2.71 × 106 7.77 0.0216 44 H 2, 8 9.02 × 107, 1.47 × 107 4.51 × 107, 2.45 × 106 18.38 0.0028 Pico-eukaryote -

Predatory Flagellates – the New Recently Discovered Deep Branches of the Eukaryotic Tree and Their Evolutionary and Ecological Significance
Protistology 14 (1), 15–22 (2020) Protistology Predatory flagellates – the new recently discovered deep branches of the eukaryotic tree and their evolutionary and ecological significance Denis V. Tikhonenkov Papanin Institute for Biology of Inland Waters, Russian Academy of Sciences, Borok, 152742, Russia | Submitted March 20, 2020 | Accepted April 6, 2020 | Summary Predatory protists are poorly studied, although they are often representing important deep-branching evolutionary lineages and new eukaryotic supergroups. This short review/opinion paper is inspired by the recent discoveries of various predatory flagellates, which form sister groups of the giant eukaryotic clusters on phylogenetic trees, and illustrate an ancestral state of one or another supergroup of eukaryotes. Here we discuss their evolutionary and ecological relevance and show that the study of such protists may be essential in addressing previously puzzling evolutionary problems, such as the origin of multicellular animals, the plastid spread trajectory, origins of photosynthesis and parasitism, evolution of mitochondrial genomes. Key words: evolution of eukaryotes, heterotrophic flagellates, mitochondrial genome, origin of animals, photosynthesis, predatory protists, tree of life Predatory flagellates and diversity of eu- of the hidden diversity of protists (Moon-van der karyotes Staay et al., 2000; López-García et al., 2001; Edg- comb et al., 2002; Massana et al., 2004; Richards The well-studied multicellular animals, plants and Bass, 2005; Tarbe et al., 2011; de Vargas et al., and fungi immediately come to mind when we hear 2015). In particular, several prevailing and very abun- the term “eukaryotes”. However, these groups of dant ribogroups such as MALV, MAST, MAOP, organisms represent a minority in the real diversity MAFO (marine alveolates, stramenopiles, opistho- of evolutionary lineages of eukaryotes. -

Sak/Plk4 and Mitotic Fidelity
Oncogene (2005) 24, 306–312 & 2005 Nature Publishing Group All rights reserved 0950-9232/05 $30.00 www.nature.com/onc Sak/Plk4 and mitotic fidelity Carol J Swallow1,2, Michael A Ko1,2, Najeeb U Siddiqui1,3,4, John W Hudson5 and James W Dennis*,1,3,4 1Samuel Lunenfeld Research Institute, Mount Sinai Hospital, 600 University Ave. R988, Toronto, Ontario, Canada M5G 1X5; 2Department of Surgery, University of Toronto, Ontario, Canada; 3Department of Microbiology and Medical Genetics, University of Toronto, Ontario, Canada; 4Department of Laboratory Medicine and Pathobiology, University of Toronto, Ontario, Canada; 5Department of Biological Sciences, University of Windsor, Ontario, Canada Sak/Plk4 differs from other polo-like kinases in having (Fernebro et al., 2002). Mutation of classical tumor only a single polo box, which assumes a novel dimer fold suppressor genes follows the Knudsen 2-hit model, that localizes to the nucleolus, centrosomes and the whereby loss of the wild-type allele as a ‘second hit’ cleavage furrow.Sak expression increases gradually in S frequently results in relaxed cellular growth controls through M phase, and Sak is destroyed by APC/C (Knudson, 1971). Inherited cancer syndromes such as dependent proteolysis.Sak-deficient mouse embryos Li-Fraumeni (p53, Chk2) and ataxia telangiectasia arrest at E7.5 and display an increased incidence of (ATM) are examples of autosomal recessive mutations apoptosis and anaphase arrest.Sak þ /À mice are haploin- in checkpoint proteins that normally delay the cell cycle sufficient for tumor suppression, with spontaneous tumors in response to DNA damage and environmental stresses developing primarily in the liver with advanced age. -

A Unicellular Relative of Animals Generates a Layer of Polarized Cells
RESEARCH ARTICLE A unicellular relative of animals generates a layer of polarized cells by actomyosin- dependent cellularization Omaya Dudin1†*, Andrej Ondracka1†, Xavier Grau-Bove´ 1,2, Arthur AB Haraldsen3, Atsushi Toyoda4, Hiroshi Suga5, Jon Bra˚ te3, In˜ aki Ruiz-Trillo1,6,7* 1Institut de Biologia Evolutiva (CSIC-Universitat Pompeu Fabra), Barcelona, Spain; 2Department of Vector Biology, Liverpool School of Tropical Medicine, Liverpool, United Kingdom; 3Section for Genetics and Evolutionary Biology (EVOGENE), Department of Biosciences, University of Oslo, Oslo, Norway; 4Department of Genomics and Evolutionary Biology, National Institute of Genetics, Mishima, Japan; 5Faculty of Life and Environmental Sciences, Prefectural University of Hiroshima, Hiroshima, Japan; 6Departament de Gene`tica, Microbiologia i Estadı´stica, Universitat de Barcelona, Barcelona, Spain; 7ICREA, Barcelona, Spain Abstract In animals, cellularization of a coenocyte is a specialized form of cytokinesis that results in the formation of a polarized epithelium during early embryonic development. It is characterized by coordinated assembly of an actomyosin network, which drives inward membrane invaginations. However, whether coordinated cellularization driven by membrane invagination exists outside animals is not known. To that end, we investigate cellularization in the ichthyosporean Sphaeroforma arctica, a close unicellular relative of animals. We show that the process of cellularization involves coordinated inward plasma membrane invaginations dependent on an *For correspondence: actomyosin network and reveal the temporal order of its assembly. This leads to the formation of a [email protected] (OD); polarized layer of cells resembling an epithelium. We show that this stage is associated with tightly [email protected] (IR-T) regulated transcriptional activation of genes involved in cell adhesion. -

S41467-021-25308-W.Pdf
ARTICLE https://doi.org/10.1038/s41467-021-25308-w OPEN Phylogenomics of a new fungal phylum reveals multiple waves of reductive evolution across Holomycota ✉ ✉ Luis Javier Galindo 1 , Purificación López-García 1, Guifré Torruella1, Sergey Karpov2,3 & David Moreira 1 Compared to multicellular fungi and unicellular yeasts, unicellular fungi with free-living fla- gellated stages (zoospores) remain poorly known and their phylogenetic position is often 1234567890():,; unresolved. Recently, rRNA gene phylogenetic analyses of two atypical parasitic fungi with amoeboid zoospores and long kinetosomes, the sanchytrids Amoeboradix gromovi and San- chytrium tribonematis, showed that they formed a monophyletic group without close affinity with known fungal clades. Here, we sequence single-cell genomes for both species to assess their phylogenetic position and evolution. Phylogenomic analyses using different protein datasets and a comprehensive taxon sampling result in an almost fully-resolved fungal tree, with Chytridiomycota as sister to all other fungi, and sanchytrids forming a well-supported, fast-evolving clade sister to Blastocladiomycota. Comparative genomic analyses across fungi and their allies (Holomycota) reveal an atypically reduced metabolic repertoire for sanchy- trids. We infer three main independent flagellum losses from the distribution of over 60 flagellum-specific proteins across Holomycota. Based on sanchytrids’ phylogenetic position and unique traits, we propose the designation of a novel phylum, Sanchytriomycota. In addition, our results indicate that most of the hyphal morphogenesis gene repertoire of multicellular fungi had already evolved in early holomycotan lineages. 1 Ecologie Systématique Evolution, CNRS, Université Paris-Saclay, AgroParisTech, Orsay, France. 2 Zoological Institute, Russian Academy of Sciences, St. ✉ Petersburg, Russia. 3 St.