Escan: Scan Regulatory Regions for Aggregate Association Testing
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

In Vivo Studies Using the Classical Mouse Diversity Panel
The Mouse Diversity Panel Predicts Clinical Drug Toxicity Risk Where Classical Models Fail Alison Harrill, Ph.D The Hamner-UNC Institute for Drug Safety Sciences 0 The Importance of Predicting Clinical Adverse Drug Reactions (ADR) Figure: Cath O’Driscoll Nature Publishing 2004 Risk ID PGx Testing 1 People Respond Differently to Drugs Pharmacogenetic Markers Identified by Genome-Wide Association Drug Adverse Drug Risk Allele Reaction (ADR) Abacavir Hypersensitivity HLA-B*5701 Flucloxacillin Hepatotoxicity Allopurinol Cutaneous ADR HLA-B*5801 Carbamazepine Stevens-Johnson HLA-B*1502 Syndrome Augmentin Hepatotoxicity DRB1*1501 Ximelagatran Hepatotoxicity DRB1*0701 Ticlopidine Hepatotoxicity HLA-A*3303 Average preclinical populations and human hepatocytes lack the diversity to detect incidence of adverse events that occur only in 1/10,000 people. Current Rodent Models of Risk Assessment The Challenge “At a time of extraordinary scientific progress, methods have hardly changed in several decades ([FDA] 2004)… Toxicologists face a major challenge in the twenty-first century. They need to embrace the new “omics” techniques and ensure that they are using the most appropriate animals if their discipline is to become a more effective tool in drug development.” -Dr. Michael Festing Quantitative geneticist Toxicol Pathol. 2010;38(5):681-90 Rodent Models as a Strategy for Hazard Characterization and Pharmacogenetics Genetically defined rodent models may provide ability to: 1. Improve preclinical prediction of drugs that carry a human safety risk 2. -

TFEB Regulates Murine Liver Cell Fate During Development and Regeneration ✉ Nunzia Pastore 1,2 , Tuong Huynh1,2, Niculin J
ARTICLE https://doi.org/10.1038/s41467-020-16300-x OPEN TFEB regulates murine liver cell fate during development and regeneration ✉ Nunzia Pastore 1,2 , Tuong Huynh1,2, Niculin J. Herz1,2, Alessia Calcagni’ 1,2, Tiemo J. Klisch1,2, Lorenzo Brunetti 3,4, Kangho Ho Kim5, Marco De Giorgi6, Ayrea Hurley6, Annamaria Carissimo7, Margherita Mutarelli7, Niya Aleksieva 8, Luca D’Orsi7, William R. Lagor6, David D. Moore 5, ✉ Carmine Settembre7,9, Milton J. Finegold10, Stuart J. Forbes8 & Andrea Ballabio 1,2,7,9 1234567890():,; It is well established that pluripotent stem cells in fetal and postnatal liver (LPCs) can differentiate into both hepatocytes and cholangiocytes. However, the signaling pathways implicated in the differentiation of LPCs are still incompletely understood. Transcription Factor EB (TFEB), a master regulator of lysosomal biogenesis and autophagy, is known to be involved in osteoblast and myeloid differentiation, but its role in lineage commitment in the liver has not been investigated. Here we show that during development and upon regen- eration TFEB drives the differentiation status of murine LPCs into the progenitor/cho- langiocyte lineage while inhibiting hepatocyte differentiation. Genetic interaction studies show that Sox9, a marker of precursor and biliary cells, is a direct transcriptional target of TFEB and a primary mediator of its effects on liver cell fate. In summary, our findings identify an unexplored pathway that controls liver cell lineage commitment and whose dysregulation may play a role in biliary cancer. 1 Jan and Dan Duncan Neurological Research Institute, Texas Children Hospital, Houston, TX 77030, USA. 2 Department of Molecular and Human Genetics, Baylor College of Medicine, Houston, TX 77030, USA. -

Full-Length Cdna, Expression Pattern and Association Analysis of the Porcine FHL3 Gene
1473 Asian-Aust. J. Anim. Sci. Vol. 20, No. 10 : 1473 - 1477 October 2007 www.ajas.info Full-length cDNA, Expression Pattern and Association Analysis of the Porcine FHL3 Gene Bo Zuo*, YuanZhu Xiong, Hua Yang and Jun Wang Key Laboratory of Swine Genetics and Breeding, Ministry of Agriculture & Key Lab of Agricultural Animal Genetics, Breeding and Reproduction, Ministry of Education, College of Animal Science and Veterinary Medicine, Huazhong Agricultural University, Wuhan, 430070, P. R. China ABSTRACT : Four-and-a-half LIM-only protein 3 (FHL3) is a member of the LIM protein superfamily and can participate in mediating protein-protein interaction by binding one another through their LIM domains. In this study, the 5'- and 3'- cDNA ends were characterized by RACE (Rapid Amplification of the cDNA Ends) methodology in combination with in silica cloning based on the partial cDNA sequence obtained. Bioinformatics analysis showed FHL3 protein contained four LIM domains and four LIM zinc-binding domains. In silica mapping assigned this gene to the gene cluster MTF1-INPP5B-SF3A3-FHL3-CGI-94 on pig chromosome 6 where several QTL affecting intramuscular fat and eye muscle area had previously been identified. Transcription of the FHL3 gene was detected in spleen, liver, kidney, small intestine, skeletal muscle, fat and stomach, with the greatest expression in skeletal muscle. The A/G polymorphism in exon II was significantly associated with birth weight, average daily gain before weaning, drip loss rate, water holding capacity and intramuscular fat in a Landrace-derived pig population. Together, the present study provided the useful information for further studies to determine the roles of FHL3 gene in the regulation of skeletal muscle cell growth and differentiation in pigs. -

Dynamic Transcriptomic Profiles of Zebrafish Gills in Response to Zinc
Zheng et al. BMC Genomics 2010, 11:548 http://www.biomedcentral.com/1471-2164/11/548 RESEARCH ARTICLE Open Access Dynamic transcriptomic profiles of zebrafish gills in response to zinc depletion Dongling Zheng1,4, Peter Kille2, Graham P Feeney2, Phil Cunningham1, Richard D Handy3, Christer Hogstrand1* Abstract Background: Zinc deficiency is detrimental to organisms, highlighting its role as an essential micronutrient contributing to numerous biological processes. To investigate the underlying molecular events invoked by zinc depletion we performed a temporal analysis of transcriptome changes observed within the zebrafish gill. This tissue represents a model system for studying ion absorption across polarised epithelial cells as it provides a major pathway for fish to acquire zinc directly from water whilst sharing a conserved zinc transporting system with mammals. Results: Zebrafish were treated with either zinc-depleted (water = 2.61 μgL-1; diet = 26 mg kg-1) or zinc-adequate (water = 16.3 μgL-1; diet = 233 mg kg-1) conditions for two weeks. Gill samples were collected at five time points and transcriptome changes analysed in quintuplicate using a 16K oligonucleotide array. Of the genes represented the expression of a total of 333 transcripts showed differential regulation by zinc depletion (having a fold-change greater than 1.8 and an adjusted P-value less than 0.1, controlling for a 10% False Discovery Rate). Down-regulation was dominant at most time points and distinct sets of genes were regulated at different stages. Annotation enrichment analysis revealed that ‘Developmental Process’ was the most significantly overrepresented Biological Process GO term (P = 0.0006), involving 26% of all regulated genes. -

Mouse Population-Guided Resequencing Reveals That Variants in CD44 Contribute to Acetaminophen-Induced Liver Injury in Humans
Downloaded from genome.cshlp.org on October 2, 2021 - Published by Cold Spring Harbor Laboratory Press Letter Mouse population-guided resequencing reveals that variants in CD44 contribute to acetaminophen-induced liver injury in humans Alison H. Harrill,1,2,12 Paul B. Watkins,3,12 Stephen Su,6 Pamela K. Ross,2 David E. Harbourt,5 Ioannis M. Stylianou,7 Gary A. Boorman,8 Mark W. Russo,3 Richard S. Sackler,9 Stephen C. Harris,11 Philip C. Smith,5 Raymond Tennant,8 Molly Bogue,7 Kenneth Paigen,7 Christopher Harris,9,10 Tanupriya Contractor,9 Timothy Wiltshire,5 Ivan Rusyn,1,2,14 and David W. Threadgill1,4,13,14,15 1Curriculum in Toxicology, University of North Carolina, Chapel Hill, North Carolina 27599, USA; 2Department of Environmental Sciences and Engineering, University of North Carolina, Chapel Hill, North Carolina 27599, USA; 3Division of Gastroenterology and Hepatology, University of North Carolina, Chapel Hill, North Carolina 27599, USA; 4Department of Genetics, University of North Carolina, Chapel Hill, North Carolina 27599, USA; 5School of Pharmacy, University of North Carolina, Chapel Hill, North Carolina 27599, USA; 6Department of Mouse Genetics, Genomics Institute of the Novartis Research Foundation, San Diego, California 92121, USA; 7The Jackson Laboratory, Bar Harbor, Maine 04609, USA; 8National Institute of Environmental Health Sciences, Research Triangle Park, North Carolina 27709, USA; 9Verto Institute Research Laboratories, New Brunswick, New Jersey 08903, USA; 10Cancer Institute of New Jersey, New Brunswick, New Jersey 08903, USA; 11Purdue Pharma L.P., Stamford, Connecticut 06901, USA; 12Hamner-UNC Center for Drug Safety Sciences, The Hamner Institutes for Health Sciences, Research Triangle Park, North Carolina 27709, USA; 13Department of Genetics, North Carolina State University, Raleigh, North Carolina 27695, USA Interindividual variability in response to chemicals and drugs is a common regulatory concern. -

Functional Genomics Atlas of Synovial Fibroblasts Defining Rheumatoid Arthritis
medRxiv preprint doi: https://doi.org/10.1101/2020.12.16.20248230; this version posted December 18, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. All rights reserved. No reuse allowed without permission. Functional genomics atlas of synovial fibroblasts defining rheumatoid arthritis heritability Xiangyu Ge1*, Mojca Frank-Bertoncelj2*, Kerstin Klein2, Amanda Mcgovern1, Tadeja Kuret2,3, Miranda Houtman2, Blaž Burja2,3, Raphael Micheroli2, Miriam Marks4, Andrew Filer5,6, Christopher D. Buckley5,6,7, Gisela Orozco1, Oliver Distler2, Andrew P Morris1, Paul Martin1, Stephen Eyre1* & Caroline Ospelt2*,# 1Versus Arthritis Centre for Genetics and Genomics, School of Biological Sciences, Faculty of Biology, Medicine and Health, The University of Manchester, Manchester, UK 2Department of Rheumatology, Center of Experimental Rheumatology, University Hospital Zurich, University of Zurich, Zurich, Switzerland 3Department of Rheumatology, University Medical Centre, Ljubljana, Slovenia 4Schulthess Klinik, Zurich, Switzerland 5Institute of Inflammation and Ageing, University of Birmingham, Birmingham, UK 6NIHR Birmingham Biomedical Research Centre, University Hospitals Birmingham NHS Foundation Trust, University of Birmingham, Birmingham, UK 7Kennedy Institute of Rheumatology, University of Oxford Roosevelt Drive Headington Oxford UK *These authors contributed equally #corresponding author: [email protected] NOTE: This preprint reports new research that has not been certified by peer review and should not be used to guide clinical practice. 1 medRxiv preprint doi: https://doi.org/10.1101/2020.12.16.20248230; this version posted December 18, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. -

Gynecologic and Obstetric Pathology
LABORATORY INVESTIGATION THE BASIC AND TRANSLATIONAL PATHOLOGY RESEARCH JOURNAL LI VOLUME 99 | SUPPLEMENT 1 | MARCH 2019 2019 ABSTRACTS GYNECOLOGIC AND OBSTETRIC PATHOLOGY (993-1161) MARCH 16-21, 2019 PLATF OR M & 2 01 9 ABSTRACTS P OSTER PRESENTATI ONS EDUCATI ON C O M MITTEE Jason L. Hornick , C h air Ja mes R. Cook R h o n d a K. Y a nti s s, Chair, Abstract Revie w Board S ar a h M. Dr y and Assign ment Co m mittee Willi a m C. F a q ui n Laura W. La mps , Chair, C ME Subco m mittee C ar ol F. F ar v er St e v e n D. Billi n g s , Interactive Microscopy Subco m mittee Y uri F e d ori w Shree G. Shar ma , Infor matics Subco m mittee Meera R. Ha meed R aj a R. S e et h al a , Short Course Coordinator Mi c h ell e S. Hir s c h Il a n W ei nr e b , Subco m mittee for Unique Live Course Offerings Laksh mi Priya Kunju D a vi d B. K a mi n s k y ( Ex- Of ici o) A n n a M ari e M ulli g a n Aleodor ( Doru) Andea Ri s h P ai Zubair Baloch Vi nita Parkas h Olca Bast urk A nil P ar w a ni Gregory R. Bean , Pat h ol o gist-i n- Trai ni n g D e e p a P atil D a ni el J. -

Single-Cell RNA-Seq Analysis Reveals the Progression of Human
Osteoarthritis Ann Rheum Dis: first published as 10.1136/annrheumdis-2017-212863 on 19 July 2018. Downloaded from TRANSLATIONAL SCIENCE Single-cell RNA-seq analysis reveals the progression of human osteoarthritis Quanbo Ji,1,2 Yuxuan Zheng,2,3,4 Guoqiang Zhang,1 Yuqiong Hu,2,3,4 Xiaoying Fan,2,3 Yu Hou,2,3 Lu Wen,2,3 Li Li,2,3 Yameng Xu,5 Yan Wang,1 Fuchou Tang2,3,4 Handling editor Josef S ABSTRact of critical importance. Nevertheless, intense efforts Smolen Objectives Understanding the molecular mechanisms have focused on the transplantation of tissue-en- underlying human cartilage degeneration and gineered cartilage derived from stem cells and ► Additional material is published online only. To view regeneration is helpful for improving therapeutic chondrocytes for cartilage regeneration in treating please visit the journal online strategies for treating osteoarthritis (OA). Here, we report OA.6–8 However, OA cartilage cell-type compo- (http:// dx. doi. org/ 10. 1136/ the molecular programmes and lineage progression sition, biochemical markers that can effectively annrheumdis- 2017- 212863). patterns controlling human OA pathogenesis using predict OA and the cellular heterogeneity leading For numbered affiliations see single-cell RNA sequencing (scRNA-seq). to OA progression remain largely unknown. end of article. Methods We performed unbiased transcriptome- Articular cartilage is derived from condensed wide scRNA-seq analysis, computational analysis and mesenchymal stem cells (MSCs), which subsequently Correspondence to histological assays on 1464 chondrocytes from 10 differentiate into chondrocytes.9 10 Following this Prof. Yan Wang, Department of patients with OA undergoing knee arthroplasty surgery. chondrogenesis, cells produce a collagenous extra- Orthopaedics, General Hospital We investigated the relationship between transcriptional of Chinese People’s Liberation cellular matrix, a ground substance containing Army, Beijing 100853, China; programmes of the OA landscape and clinical outcome abundant collagen and proteoglycans. -
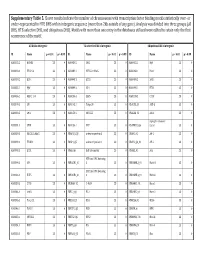
Supplementary Table 5. Clover Results Indicate the Number Of
Supplementary Table 5. Clover results indicate the number of chromosomes with transcription factor binding motifs statistically over‐ or under‐represented in HTE DHS within intergenic sequence (more than 2kb outside of any gene). Analysis was divided into three groups (all DHS, HTE‐selective DHS, and ubiquitous DHS). Motifs with more than one entry in the databases utilized were edited to retain only the first occurrence of the motif. All DHS x Intergenic TEselective DHS x Intergenic Ubiquitous DHS x Intergenic ID Name p < 0.01 p > 0.99 ID Name p < 0.01 p > 0.99 ID Name p < 0.01 p > 0.99 MA0002.2 RUNX1 23 0 MA0080.2 SPI1 23 0 MA0055.1 Myf 23 0 MA0003.1 TFAP2A 23 0 MA0089.1 NFE2L1::MafG 23 0 MA0068.1 Pax4 23 0 MA0039.2 Klf4 23 0 MA0098.1 ETS1 23 0 MA0080.2 SPI1 23 0 MA0055.1 Myf 23 0 MA0099.2 AP1 23 0 MA0098.1 ETS1 23 0 MA0056.1 MZF1_1‐4 23 0 MA0136.1 ELF5 23 0 MA0139.1 CTCF 23 0 MA0079.2 SP1 23 0 MA0145.1 Tcfcp2l1 23 0 V$ALX3_01 ALX‐3 23 0 MA0080.2 SPI1 23 0 MA0150.1 NFE2L2 23 0 V$ALX4_02 Alx‐4 23 0 myocyte enhancer MA0081.1 SPIB 23 0 MA0156.1 FEV 23 0 V$AMEF2_Q6 factor 23 0 MA0089.1 NFE2L1::MafG 23 0 V$AP1FJ_Q2 activator protein 1 23 0 V$AP1_01 AP‐1 23 0 MA0090.1 TEAD1 23 0 V$AP4_Q5 activator protein 4 23 0 V$AP2_Q6_01 AP‐2 23 0 MA0098.1 ETS1 23 0 V$AR_Q6 half‐site matrix 23 0 V$ARX_01 Arx 23 0 BTB and CNC homolog MA0099.2 AP1 23 0 V$BACH1_01 1 23 0 V$BARHL1_01 Barhl‐1 23 0 BTB and CNC homolog MA0136.1 ELF5 23 0 V$BACH2_01 2 23 0 V$BARHL2_01 Barhl2 23 0 MA0139.1 CTCF 23 0 V$CMAF_02 C‐MAF 23 0 V$BARX1_01 Barx1 23 0 MA0144.1 Stat3 23 0 -

What Your Genome Doesn't Tell
UC San Diego UC San Diego Electronic Theses and Dissertations Title Multi-layered epigenetic control of T cell fate decisions Permalink https://escholarship.org/uc/item/8rs7c7b3 Author Yu, Bingfei Publication Date 2018 Peer reviewed|Thesis/dissertation eScholarship.org Powered by the California Digital Library University of California UNIVERSITY OF CALIFORNIA SAN DIEGO Multi-layered epigenetic control of T cell fate decisions A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Biology by Bingfei Yu Committee in charge: Professor Ananda Goldrath, Chair Professor John Chang Professor Stephen Hedrick Professor Cornelis Murre Professor Wei Wang 2018 Copyright Bingfei Yu, 2018 All rights reserved. The dissertation of Bingfei Yu is approved, and it is ac- ceptable in quality and form for publication on microfilm and electronically: Chair University of California San Diego 2018 iii DEDICATION To my parents who have been giving me countless love, trust and support to make me who I am. iv EPIGRAPH Stay hungary. Stay foolish. | Steve Jobs quoted from the back cover of the 1974 edition of the Whole Earth Catalog v TABLE OF CONTENTS Signature Page.................................. iii Dedication..................................... iv Epigraph.....................................v Table of Contents................................. vi List of Figures.................................. ix Acknowledgements................................x Vita........................................ xii Abstract of -
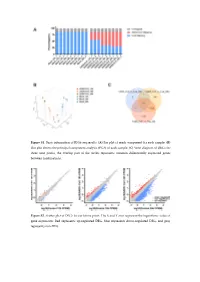
Figure S1. Basic Information of RNA-Seq Results. (A) Bar Plot of Reads Component for Each Sample
Figure S1. Basic information of RNA-seq results. (A) Bar plot of reads component for each sample. (B) Dot plot shows the principal component analysis (PCA) of each sample. (C) Venn diagram of DEGs for three time points, the overlap part of the circles represents common differentially expressed genes between combinations. Figure S2. Scatter plot of DEGs for each time point. The X and Y axes represent the logarithmic value of gene expression. Red represents up-regulated DEG, blue represents down-regulated DEG, and gray represents non-DEG. Table S1. Primers used for quantitative real-time PCR analysis of DEGs. Gene Primer Sequence Forward 5’-CTACGAGTGGATGGTCAAGAGC-3’ FOXO1 Reverse 5’-CCAGTTCCTTCATTCTGCACACG-3’ Forward 5’-GACGTCCGGCATCAGAGAAA-3’ IRS2 Reverse 5’-TCCACGGCTAATCGTCACAG-3’ Forward 5’-CACAACCAGGACCTCACACC-3’ IRS1 Reverse 5’-CTTGGCACGATAGAGAGCGT-3’ Forward 5’-AGGATACCACTCCCAACAGACCT-3’ IL6 Reverse 5’-CAAGTGCATCATCGTTGTTCATAC-3’ Forward 5’-TCACGTTGTACGCAGCTACC-3’ CCL5 Reverse 5’-CAGTCCTCTTACAGCCTTTGG-3’ Forward 5’-CTGTGCAGCCGCAGTGCCTACC-3’ BMP7 Reverse 5’-ATCCCTCCCCACCCCACCATCT-3’ Forward 5’-CTCTCCCCCTCGACTTCTGA-3’ BCL2 Reverse 5’-AGTCACGCGGAACACTTGAT-3’ Forward 5’-CTGTCGAACACAGTGGTACCTG-3’ FGF7 Reverse 5’-CCAACTGCCACTGTCCTGATTTC-3’ Forward 5’-GGGAGCCAAAAGGGTCATCA-3’ GAPDH Reverse 5’-CGTGGACTGTGGTCATGAGT-3’ Supplementary material: Differentially expressed genes log2(SADS-CoV_12h/ Qvalue (SADS-CoV _12h/ Gene Symbol Control_12h) Control_12h) PTGER4 -1.03693 6.79E-04 TMEM72 -3.08132 3.66E-04 IFIT2 -1.02918 2.11E-07 FRAT2 -1.09282 4.66E-05 -

Overview Gene List Target Scan Vs DIANA Group a Group B Group A
Overview Gene list Target scan vs DIANA Group A Group B Group A hsa-miR-181a hsa-miR-323 hsa-miR-326 Target scan Diana microT Overlap Target scan Diana microT Overlap Target scan SEPT3 SEPT3 SEPT3 SEPT7 ADARB1 HPCAL4 ABHD2 ABL2 ABHD13 ACVR2A ADCYAP1R1 AKAP13 PDPK1 ACRBP ACAN ABI1 ADAMTS1 ALAD APOBR ACVRL1 ACCN2 ABLIM1 ADAMTSL1 ANKRD52 ATXN1 ADAM19 ACER3 ACSL1 AKAP7 ARID2 C18orf23,RNF165 ADAM33 ACVR2A ACTN2 ANKRD43 ARL3 C20orf29 ADAMTS2 ADAMTS1 ACVR2A AP1S3 ARRB1 CACNG4 AHCYL2 ADAMTS18 ACVR2B ARID2 BBC3 CCNJL ALOX15B ADAMTS5 ADAM11 ATP11A BTG1 CYP2E1 ANK1 ADAMTSL1 ADAM22 ATXN1 C18orf62 GNB1L ANKS6 ADARB1 ADAMTS1 B4GALT1 C1orf21 GPR61 APBA1 AFAP1 ADAMTS6 BAG4 CADM4 GTSE1 ARCN1 AFTPH ADAMTSL1 BAI3 CALML4 HPCAL4 ARHGEF37 AK3 ADCY9 BNC2 CAPN6 KIAA0152 ARID3B AKAP7 ADRBK1 BRD1 CBFA2T2 KIF1A ARL8A ANAPC16 AFF2 BRWD1 CEBPA MACF1 ATP2B2 ANK1 AHCTF1,AHCTF1PBTBD3 CHD1 MYO1D ATP6V1G2 ANKRD12 AKAP2,PALM2 C13orf23 CIT PCNT AUP1 ANKRD33B AKAP6 C14orf43 CLASP2 PDPK1 BCL2L2 ANKRD43 AKAP7 CAPRIN1 CLCN5 PLEKHG4B BHLHE40 ANKRD44 AKAP9 CARM1 CLIP3 PPARA BTBD3 ANKRD52 AKT3 CBX4 COL5A2 PRB1,PRB2,PRB4 BTRC AP1S3 ALG9 CCDC117 CTNS PTPRT C10orf26 APBA1 ANKRD13C CCNJ DCTN4 PYCR1 C14orf1 APLP2 ANKRD20B CDH13 DCUN1D4 RAPGEF1 C16orf45 APOO ANKRD43 CDON DDB1 SRCAP C16orf54 ARID2 ANKRD50 CDYL DDX39B TMEM63C C1orf106 ARL3 AP1G1 CEP350 DIP2C C1orf27 ARRDC3 AP1S3 CHD7 DNAJB3 C22orf29 ATF7 API5 CHIC1 EEPD1 C9orf3 ATG2B ARFGEF2 CLIP1 EIF2C1 CACNA1E ATG7 ARHGAP12 CNOT6L ELFN2 CAPN12 ATP11A ARHGAP26 CNR1 ELK1 CASKIN1 ATP2B3 ARHGAP29 CNTN4 FAM172A CBFA2T3 ATP8B2 ARHGEF3 CNTNAP2