High HSPA8 Expression Predicts Adverse Outcomes of Acute Myeloid Leukemia Jun Li1 and Zheng Ge1,2*
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

STUB1 / CHIP Antibody Rabbit Polyclonal Antibody Catalog # ALS16226
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 STUB1 / CHIP Antibody Rabbit Polyclonal Antibody Catalog # ALS16226 Specification STUB1 / CHIP Antibody - Product Information Application WB Primary Accession Q9UNE7 Reactivity Human Host Rabbit Clonality Polyclonal Calculated MW 35kDa KDa STUB1 / CHIP Antibody - Additional Information Sample (30 ug of whole cell lysate) A: 293T Gene ID 10273 10% SDS PAGE STUB1 antibody diluted at 1:10000 Other Names E3 ubiquitin-protein ligase CHIP, 6.3.2.-, Antigen NY-CO-7, CLL-associated antigen STUB1 / CHIP Antibody - Background KW-8, Carboxy terminus of Hsp70-interacting protein, STIP1 homology and U box-containing protein 1 E3 ubiquitin-protein ligase which targets {ECO:0000312|HGNC:HGNC:11427}, STUB1 misfolded chaperone substrates towards (<a href="http://www.genenames.org/cgi-bi proteasomal degradation. Collaborates with n/gene_symbol_report?hgnc_id=11427" ATXN3 in the degradation of misfolded target="_blank">HGNC:11427</a>) chaperone substrates: ATXN3 restricting the length of ubiquitin chain attached to Target/Specificity STUB1/CHIP substrates and preventing further Human STUB1 chain extension. Ubiquitinates NOS1 in concert with Hsp70 and Hsp40. Modulates the activity Reconstitution & Storage of several chaperone complexes, including Keep as concentrated solution. Aliquot and Hsp70, Hsc70 and Hsp90. Mediates transfer of store at -20°C or below. Avoid multiple non-canonical short ubiquitin chains to HSPA8 freeze-thaw cycles. that have no effect on HSPA8 degradation. Mediates polyubiquitination of DNA polymerase Precautions beta (POLB) at 'Lys-41', 'Lys-61' and 'Lys-81', STUB1 / CHIP Antibody is for research use thereby playing a role in base-excision repair: only and not for use in diagnostic or catalyzes polyubiquitination by amplifying the therapeutic procedures. -

The Functions of CHIP in Age Related Disease
Central JSM Enzymology and Protein Science Bringing Excellence in Open Access Review Article *Corresponding author Kathryn L Ball, The Institute of Genetics and Molecular Medicine, Edinburgh Cancer Research Centre, The Functions of CHIP in Age University of Edinburgh, Western General Hospital, Crewe Road South, EH4 2XR, UK, Tel: +44 0-131-651-8500; E-mail: Related Disease Submitted: 03 August 2016 Ball KL*, Ning J, Nita E, and Dias C Accepted: 29 September 2016 Institute of Genetics and Molecular Medicine, University of Edinburgh, UK Published: 01 October 2016 Copyright Abstract © 2016 Ball et al. CHIP is a key component of the protein homeostasis or ‘Proteostasis’ network OPEN ACCESS that maintains protein structure and function as a way to ensure the integrity of the proteome in individual cells and the health of the whole organism. Proteostasis Keywords influences the biogenesis, folding, trafficking and degradation of proteins. Originally • CHIP identified as a Hsc70 associated protein and a co-chaperone CHIP has E3-ubiquitin • E3-ligase ligase activity and also displays an intrinsic chaperoning ability. It has become clear • Chaperone that CHIP is a multi-functional protein with roles in cellular processes that go beyond • Structure function its co-chaperone activity. Not surprisingly, by unravelling the functions of CHIP, we are • Neurodegeneration beginning to appreciate that loss of CHIP’s integrity can lead to the development of • Cancer several serious pathological conditions. Here we will describe the key features of CHIPs structure and functions with an emphasis on the non-canonical activities of CHIP before concentrating on the role it plays in protecting against the age associated pathologies of neurodegeneration and cancer. -

The Role of RNA Editing in Cancer Development and Metabolic Disorders
Washington University School of Medicine Digital Commons@Becker Open Access Publications 2018 The oler of RNA editing in cancer development and metabolic disorders Che-Pei Kung Washington University School of Medicine in St. Louis Leonard B. Maggi Jr. Washington University School of Medicine in St. Louis Jason D. Weber Washington University School of Medicine in St. Louis Follow this and additional works at: https://digitalcommons.wustl.edu/open_access_pubs Recommended Citation Kung, Che-Pei; Maggi, Leonard B. Jr.; and Weber, Jason D., ,"The or le of RNA editing in cancer development and metabolic disorders." Frontiers in endocrinology.9,. 762. (2018). https://digitalcommons.wustl.edu/open_access_pubs/7400 This Open Access Publication is brought to you for free and open access by Digital Commons@Becker. It has been accepted for inclusion in Open Access Publications by an authorized administrator of Digital Commons@Becker. For more information, please contact [email protected]. REVIEW published: 18 December 2018 doi: 10.3389/fendo.2018.00762 The Role of RNA Editing in Cancer Development and Metabolic Disorders Che-Pei Kung 1,2*, Leonard B. Maggi Jr. 1,2 and Jason D. Weber 1,2,3* 1 ICCE Institute, Washington University School of Medicine, Saint Louis, MO, United States, 2 Division of Molecular Oncology, Department of Medicine, Washington University School of Medicine, Saint Louis, MO, United States, 3 Siteman Cancer Center, Department of Cell Biology and Physiology, Washington University School of Medicine, Saint Louis, MO, United States Numerous human diseases arise from alterations of genetic information, most notably DNA mutations. Thought to be merely the intermediate between DNA and protein, changes in RNA sequence were an afterthought until the discovery of RNA editing 30 years ago. -

Mohammad Karbaschi Thesis
STRUCTURAL, PHYSIOLOGICAL AND MOLECULAR CHARACTERISATION OF THE AUSTRALIAN NATIVE RESURRECTION GRASS TRIPOGON LOLIIFORMIS (F.MUELL.) C.E.HUBB. DURING DEHYDRATION AND REHYDRATION Mohammad Reza Karbaschi Submitted in fulfilment of the requirements for the degree of Doctor of Philosophy Centre for Tropical Crops and Biocommodities Science and Engineering Faculty Queensland University of Technology November 2015 Keywords Arabidopsis thaliana; Agrobacterium-mediated transformation; Anatomy; Anti-apoptotic proteins; BAG4; Escherichia coli; Bulliform cells; C4 photosynthesis; Cell wall folding; Cell membrane integrity; Chaperone-mediated autophagy; Chlorophyll fluorescence; Hsc70/Hsp70; Desiccation tolerance, Dehydration; Drought; Electrolyte leakage; Freehand sectioning; Homoiochlorophyllous; Leaf structure; Leaf folding; Reactive oxygen species (ROS); Resurrection plant; Morphology; Monocotyledon; Nicotiana benthamiana; Photosynthesis; Physiology; Plant tissue; Programed cell death (PCD); Propidium iodide staining; Protein microarray chip; Sclerenchymatous tissue; Stress; Structure; Tripogon loliiformis; Ubiquitin; Vacuole fragmentation; Kranz anatomy; XyMS+; Structural, physiological and molecular characterisation of the Australian native resurrection grass Tripogon loliiformis (F.Muell.) C.E.Hubb. during dehydration and rehydration i Abstract Plants, as sessile organisms must continually adapt to environmental changes. Water deficit is one of the major environmental stresses that affects plants. While most plants can tolerate moderate dehydration -
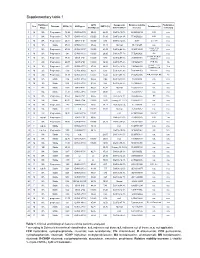
Supplementary Table 2 Supplementary Table 1
Supplementary table 1 Rai/ Binet IGHV Cytogenetic Relative viability Fludarabine- Sex Outcome CD38 (%) IGHV gene ZAP70 (%) Treatment (s) Stage identity (%) abnormalities* increase refractory 1 M 0/A Progressive 14,90 IGHV3-64*05 99,65 28,20 Del17p 18.0% 62,58322819 FCR n.a. 2 F 0/A Progressive 78,77 IGHV3-48*03 100,00 51,90 Del17p 24.8% 77,88052021 FCR n.a. 3 M 0/A Progressive 29,81 IGHV4-b*01 100,00 9,10 Del17p 12.0% 36,48 Len, Chl n.a. 4 M 1/A Stable 97,04 IGHV3-21*01 97,22 18,11 Normal 85,4191657 n.a. n.a. Chl+O, PCR, 5 F 0/A Progressive 87,00 IGHV4-39*07 100,00 43,20 Del13q 68.3% 35,23314039 n.a. HDMP+R 6 M 0/A Progressive 1,81 IGHV3-43*01 100,00 20,90 Del13q 77.7% 57,52490626 Chl n.a. Chl, FR, R-CHOP, 7 M 0/A Progressive 97,80 IGHV1-3*01 100,00 9,80 Del17p 88.5% 48,57389901 n.a. HDMP+R 8 F 2/B Progressive 69,07 IGHV5-a*03 100,00 16,50 Del17p 77.2% 107,9656878 FCR, BA No R-CHOP, FCR, 9 M 1/A Progressive 2,13 IGHV3-23*01 97,22 29,80 Del11q 16.3% 134,5866919 Yes Flavopiridol, BA 10 M 2/A Progressive 0,36 IGHV3-30*02 92,01 0,38 Del13q 81.9% 78,91844953 Unknown n.a. 11 M 2/B Progressive 15,17 IGHV3-20*01 100,00 13,20 Del11q 95.3% 75,52880995 FCR, R-CHOP, BR No 12 M 0/A Stable 0,14 IGHV3-30*02 90,62 7,40 Del13q 13.0% 13,0939004 n.a. -

Kinase SH3 NH2 Cdc42
A Dissertation The E3 Ligase CHIP Mediates Ubiquitination and Degradation of Mixed Lineage Kinase 3 and Mixed Lineage Kinase 4 Beta by Natalya A. Blessing Submitted to the Graduate Faculty as partial fulfillment of the requirements for the Doctor of Philosophy Degree in Biology _________________________________________ Dr. Deborah Chadee, Committee Chair _________________________________________ Dr. Richard Komuniecki, Committee Member _________________________________________ Dr. Malathi Krishnamurthy, Committee Member _________________________________________ Dr. Frank Pizza, Committee Member _________________________________________ Dr. Don Ronning, Committee Member _________________________________________ Dr. Patricia R. Komuniecki, Dean College of Graduate Studies The University of Toledo May 2015 Copyright 2015, Natalya A. Blessing This document is copyrighted material. Under copyright law, no parts of this document may be reproduced without the expressed permission of the author. An Abstract of The E3 Ligase CHIP Mediates Ubiquitination and Degradation of Mixed Lineage Kinase 3 and Mixed Lineage Kinase 4 Beta by Natalya Blessing Submitted to the Graduate Faculty as partial fulfillment of the requirements for the Doctor of Philosophy Degree in Biology The University of Toledo May 2015 The mixed lineage kinases (MLKs) are serine/threonine mitogen-activated protein kinase kinase kinases (MAP3Ks) that modulate the activities of extracellular signal- regulated kinase, c-Jun N-terminal kinase, and p38 signaling pathways. MLK3 plays a pivotal role in cell invasion, tumorigenesis and metastasis. Wild type MLK4 negatively regulates MAPK signaling by possibly inhibiting MLK3 activation, while mutant MLK4 plays in important role in driving colorectal cancer and glioblastomal tumorigenesis (Martini et al., 2013). The mechanisms by which MLK3 and MLK4 protein levels are regulated in cells are unknown. The carboxyl terminus of HSC-70 interacting protein (CHIP) is a U-box E3 ubiquitin ligase that regulates cytosolic protein degradation in response to stress. -

Senescence Inhibits the Chaperone Response to Thermal Stress
SUPPLEMENTAL INFORMATION Senescence inhibits the chaperone response to thermal stress Jack Llewellyn1, 2, Venkatesh Mallikarjun1, 2, 3, Ellen Appleton1, 2, Maria Osipova1, 2, Hamish TJ Gilbert1, 2, Stephen M Richardson2, Simon J Hubbard4, 5 and Joe Swift1, 2, 5 (1) Wellcome Centre for Cell-Matrix Research, Oxford Road, Manchester, M13 9PT, UK. (2) Division of Cell Matrix Biology and Regenerative Medicine, School of Biological Sciences, Faculty of Biology, Medicine and Health, Manchester Academic Health Science Centre, University of Manchester, Manchester, M13 9PL, UK. (3) Current address: Department of Biomedical Engineering, University of Virginia, Box 800759, Health System, Charlottesville, VA, 22903, USA. (4) Division of Evolution and Genomic Sciences, School of Biological Sciences, Faculty of Biology, Medicine and Health, Manchester Academic Health Science Centre, University of Manchester, Manchester, M13 9PL, UK. (5) Correspondence to SJH ([email protected]) or JS ([email protected]). Page 1 of 11 Supplemental Information: Llewellyn et al. Chaperone stress response in senescence CONTENTS Supplemental figures S1 – S5 … … … … … … … … 3 Supplemental table S6 … … … … … … … … 10 Supplemental references … … … … … … … … 11 Page 2 of 11 Supplemental Information: Llewellyn et al. Chaperone stress response in senescence SUPPLEMENTAL FIGURES Figure S1. A EP (passage 3) LP (passage 16) 200 µm 200 µm 1.5 3 B Mass spectrometry proteomics (n = 4) C mRNA (n = 4) D 100k EP 1.0 2 p < 0.0001 p < 0.0001 LP p < 0.0001 p < 0.0001 ) 0.5 1 2 p < 0.0001 p < 0.0001 10k 0.0 0 -0.5 -1 Cell area (µm Cell area fold change vs. EP fold change vs. -

Silencer of Death Domain (BAG4) (NM 004874) Human Recombinant Protein Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for TP306235 Silencer of Death Domain (BAG4) (NM_004874) Human Recombinant Protein Product data: Product Type: Recombinant Proteins Description: Recombinant protein of human BCL2-associated athanogene 4 (BAG4) Species: Human Expression Host: HEK293T Tag: C-Myc/DDK Predicted MW: 49.4 kDa Concentration: >50 ug/mL as determined by microplate BCA method Purity: > 80% as determined by SDS-PAGE and Coomassie blue staining Buffer: 25 mM Tris.HCl, pH 7.3, 100 mM glycine, 10% glycerol Preparation: Recombinant protein was captured through anti-DDK affinity column followed by conventional chromatography steps. Storage: Store at -80°C. Stability: Stable for 12 months from the date of receipt of the product under proper storage and handling conditions. Avoid repeated freeze-thaw cycles. RefSeq: NP_004865 Locus ID: 9530 UniProt ID: O95429 RefSeq Size: 4478 Cytogenetics: 8p11.23 RefSeq ORF: 1371 Synonyms: BAG-4; SODD This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 Silencer of Death Domain (BAG4) (NM_004874) Human Recombinant Protein – TP306235 Summary: The protein encoded by this gene is a member of the BAG1-related protein family. BAG1 is an anti-apoptotic protein that functions through interactions with a variety of cell apoptosis and growth related proteins including BCL-2, Raf-protein kinase, steroid hormone receptors, growth factor receptors and members of the heat shock protein 70 kDa family. -

STUB1 Mutations in Autosomal Recessive Ataxias
Heimdal et al. Orphanet Journal of Rare Diseases 2014, 9:146 http://www.ojrd.com/content/9/1/146 RESEARCH Open Access STUB1 mutations in autosomal recessive ataxias – evidence for mutation-specific clinical heterogeneity Ketil Heimdal1*, Monica Sanchez-Guixé2,3, Ingvild Aukrust2, Jens Bollerslev4,5, Ove Bruland2, Greg Eigner Jablonski6, Anne Kjersti Erichsen7, Einar Gude8, Jeanette A Koht9, Sigrid Erdal2, Torunn Fiskerstrand2,3, Bjørn Ivar Haukanes2, Helge Boman2, Lise Bjørkhaug10, Chantal ME Tallaksen11,12, Per M Knappskog2,13† and Stefan Johansson2,13† Abstract Background: A subset of hereditary cerebellar ataxias is inherited as autosomal recessive traits (ARCAs). Classification of recessive ataxias due to phenotypic differences in the cerebellum and cerebellar structures is constantly evolving due to new identified disease genes. Recently, reports have linked mutations in genes involved in ubiquitination (RNF216, OTUD4, STUB1) to ARCA with hypogonadism. Methods and results: With a combination of homozygozity mapping and exome sequencing, we identified three mutations in STUB1 in two families with ARCA and cognitive impairment; a homozygous missense variant (c.194A > G, p.Asn65Ser) that segregated in three affected siblings, and a missense change (c.82G > A, p.Glu28Lys) which was inherited in trans with a nonsense mutation (c.430A > T, p.Lys144Ter) in another patient. STUB1 encodes CHIP (C-terminus of Heat shock protein 70 – Interacting Protein), a dual function protein with a role in ubiquitination as a co-chaperone with heat shock proteins, and as an E3 ligase. We show that the p.Asn65Ser substitution impairs CHIP’s ability to ubiquitinate HSC70 in vitro, despite being able to self-ubiquitinate. -

Aggresomal Sequestration and STUB1-Mediated Ubiquitylation During Mammalian Proteaphagy of Inhibited Proteasomes
Aggresomal sequestration and STUB1-mediated ubiquitylation during mammalian proteaphagy of inhibited proteasomes Won Hoon Choia,b, Yejin Yuna,b, Seoyoung Parka,c, Jun Hyoung Jeona,b, Jeeyoung Leea,b, Jung Hoon Leea,c, Su-A Yangd, Nak-Kyoon Kime, Chan Hoon Jungb, Yong Tae Kwonb, Dohyun Hanf, Sang Min Lime, and Min Jae Leea,b,c,1 aDepartment of Biochemistry and Molecular Biology, Seoul National University College of Medicine, 03080 Seoul, Korea; bDepartment of Biomedical Sciences, Seoul National University Graduate School, 03080 Seoul, Korea; cNeuroscience Research Institute, Seoul National University College of Medicine, 03080 Seoul, Korea; dScience Division, Tomocube, 34109 Daejeon, Korea; eConvergence Research Center for Diagnosis, Korea Institute of Science and Technology, 02792 Seoul, Korea; and fProteomics Core Facility, Biomedical Research Institute, Seoul National University Hospital, 03080 Seoul, Korea Edited by Richard D. Vierstra, Washington University in St. Louis, St. Louis, MO, and approved July 1, 2020 (received for review November 18, 2019) The 26S proteasome, a self-compartmentalized protease complex, additional LC3-interacting region; the target cargoes can be plays a crucial role in protein quality control. Multiple levels of docked onto phosphatidylethanolamine-modified LC3 (LC3-II) regulatory systems modulate proteasomal activity for substrate on the expanding phagophore membrane, enveloped by an hydrolysis. However, the destruction mechanism of mammalian autophagosome, and eventually degraded in the autolysosomes. proteasomes is poorly understood. We found that inhibited pro- Notably, the enzymatic cascade attaching the lipid moiety at the teasomes are sequestered into the insoluble aggresome via C-terminal glycine of the cleaved LC3 protein in autophagy re- HDAC6- and dynein-mediated transport. -

Prognostic and Functional Significant of Heat Shock Proteins (Hsps)
biology Article Prognostic and Functional Significant of Heat Shock Proteins (HSPs) in Breast Cancer Unveiled by Multi-Omics Approaches Miriam Buttacavoli 1,†, Gianluca Di Cara 1,†, Cesare D’Amico 1, Fabiana Geraci 1 , Ida Pucci-Minafra 2, Salvatore Feo 1 and Patrizia Cancemi 1,2,* 1 Department of Biological Chemical and Pharmaceutical Sciences and Technologies (STEBICEF), University of Palermo, 90128 Palermo, Italy; [email protected] (M.B.); [email protected] (G.D.C.); [email protected] (C.D.); [email protected] (F.G.); [email protected] (S.F.) 2 Experimental Center of Onco Biology (COBS), 90145 Palermo, Italy; [email protected] * Correspondence: [email protected]; Tel.: +39-091-2389-7330 † These authors contributed equally to this work. Simple Summary: In this study, we investigated the expression pattern and prognostic significance of the heat shock proteins (HSPs) family members in breast cancer (BC) by using several bioinfor- matics tools and proteomics investigations. Our results demonstrated that, collectively, HSPs were deregulated in BC, acting as both oncogene and onco-suppressor genes. In particular, two different HSP-clusters were significantly associated with a poor or good prognosis. Interestingly, the HSPs deregulation impacted gene expression and miRNAs regulation that, in turn, affected important bio- logical pathways involved in cell cycle, DNA replication, and receptors-mediated signaling. Finally, the proteomic identification of several HSPs members and isoforms revealed much more complexity Citation: Buttacavoli, M.; Di Cara, of HSPs roles in BC and showed that their expression is quite variable among patients. In conclusion, G.; D’Amico, C.; Geraci, F.; we elaborated two panels of HSPs that could be further explored as potential biomarkers for BC Pucci-Minafra, I.; Feo, S.; Cancemi, P. -

Bcl‑2 Associated Athanogene 4 Promotes Proliferation, Migration and Invasion of Gastric Cancer Cells
MOLECULAR MEDICINE REPORTS 16: 3753-3760, 2017 Bcl‑2 associated athanogene 4 promotes proliferation, migration and invasion of gastric cancer cells LIZHI YI1*, ZHENBING LV2,3*, JIANMEI WANG4 and XIANFEI ZHONG1 1Department of Gastroenterology, Leshan People's Hospital, Leshan, Sichuan 614000; 2Department of General Surgery Two, Nanchong Central Hospital; 3The Second Clinical School of North Sichuan Medical College, Nanchong, Sichuan 637000; 4Department of Pathology, The Affiliated Hospital of Southwest Medical University, Luzhou, Sichuan 646000, P.R. China Received May 22, 2016; Accepted April 4, 2017 DOI: 10.3892/mmr.2017.7073 Abstract. Currently, with the increase of morbidity and the rate of GC mortality has been decreased (4). It is necessary mortality rate, gastric cancer (GC) is attracting increasing to identify the biomarkers that can distinguish between GC attention in China. Bcl-2-associated athanogene 4 (BAG4) patients with poor or good prognosis. has been identified as a tumor promoter in several tumors, Bcl-2 associated athanogene 4 [BAG4, also known as but its role in GC remains unknown. The present study Silencer of Death Domains (SODD)] is a member of the aimed to detect the expression of BAG4 and determine its BAG1‑related protein family (5). The BAG proteins are located function in the progression of GC. The results from reverse in both cytoplasmic and nuclear in cells (6), although the func- transcription‑quantitative polymerase chain reaction and tional importance of this remains unclear. BAG proteins have a western blotting revealed that BAG4 was markedly upregu- conserved BAG domain (BD) that binds to the ATPase domain lated in highly metastatic cell lines (SGC7901 and MGC803), of Hsp70/Hsc70, and regulates the activity of these molecular compared with the lower‑metastatic cell lines (AGS and chaperones (7-9).