Rabbit Anti-CYFIP2/FITC Conjugated Antibody-SL14140R-FITC
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

S41436-020-01011-X.Pdf
Zurich Open Repository and Archive University of Zurich Main Library Strickhofstrasse 39 CH-8057 Zurich www.zora.uzh.ch Year: 2020 New insights into the clinical and molecular spectrum of the novel CYFIP2-related neurodevelopmental disorder and impairment of the WRC-mediated actin dynamics Begemann, Anaïs ; Sticht, Heinrich ; Begtrup, Amber ; Vitobello, Antonio ; Faivre, Laurence ; Asadollahi, Reza ; Zweier, Markus ; Steindl, Katharina ; Rauch, Anita Abstract: PURPOSE A few de novo missense variants in the cytoplasmic FMRP-interacting protein 2 (CYFIP2) gene have recently been described as a novel cause of severe intellectual disability, seizures, and hypotonia in 18 individuals, with p.Arg87 substitutions in the majority. METHODS We assembled data from 19 newly identified and all 18 previously published individuals with CYFIP2 variants. By structural modeling and investigation of WAVE-regulatory complex (WRC)-mediated actin polymerization in six patient fibroblast lines we assessed the impact of CYFIP2 variants on the WRC. RESULTS Sixteenof 19 individuals harbor two previously described and 11 novel (likely) disease-associated missense variants. We report p.Asp724 as second mutational hotspot (4/19 cases). Genotype-phenotype correlation con- firms a consistently severe phenotype in p.Arg87 patients but a more variable phenotype inp.Asp724 and other substitutions. Three individuals with milder phenotypes carry putative loss-of-function vari- ants, which remain of unclear pathogenicity. Structural modeling predicted missense variants to disturb interactions within the WRC or impair CYFIP2 stability. Consistent with its role in WRC-mediated actin polymerization we substantiate aberrant regulation of the actin cytoskeleton in patient fibroblasts. CONCLUSION Our study expands the clinical and molecular spectrum of CYFIP2-related neurodevel- opmental disorder and provides evidence for aberrant WRC-mediated actin dynamics as contributing cellular pathomechanism. -

Supplemental Information
Supplemental information Dissection of the genomic structure of the miR-183/96/182 gene. Previously, we showed that the miR-183/96/182 cluster is an intergenic miRNA cluster, located in a ~60-kb interval between the genes encoding nuclear respiratory factor-1 (Nrf1) and ubiquitin-conjugating enzyme E2H (Ube2h) on mouse chr6qA3.3 (1). To start to uncover the genomic structure of the miR- 183/96/182 gene, we first studied genomic features around miR-183/96/182 in the UCSC genome browser (http://genome.UCSC.edu/), and identified two CpG islands 3.4-6.5 kb 5’ of pre-miR-183, the most 5’ miRNA of the cluster (Fig. 1A; Fig. S1 and Seq. S1). A cDNA clone, AK044220, located at 3.2-4.6 kb 5’ to pre-miR-183, encompasses the second CpG island (Fig. 1A; Fig. S1). We hypothesized that this cDNA clone was derived from 5’ exon(s) of the primary transcript of the miR-183/96/182 gene, as CpG islands are often associated with promoters (2). Supporting this hypothesis, multiple expressed sequences detected by gene-trap clones, including clone D016D06 (3, 4), were co-localized with the cDNA clone AK044220 (Fig. 1A; Fig. S1). Clone D016D06, deposited by the German GeneTrap Consortium (GGTC) (http://tikus.gsf.de) (3, 4), was derived from insertion of a retroviral construct, rFlpROSAβgeo in 129S2 ES cells (Fig. 1A and C). The rFlpROSAβgeo construct carries a promoterless reporter gene, the β−geo cassette - an in-frame fusion of the β-galactosidase and neomycin resistance (Neor) gene (5), with a splicing acceptor (SA) immediately upstream, and a polyA signal downstream of the β−geo cassette (Fig. -

Characterization of a 7.6-Mb Germline Deletion Encompassing the NF1 Locus and About a Hundred Genes in an NF1 Contiguous Gene Syndrome Patient
European Journal of Human Genetics (2008) 16, 1459–1466 & 2008 Macmillan Publishers Limited All rights reserved 1018-4813/08 $32.00 www.nature.com/ejhg ARTICLE Characterization of a 7.6-Mb germline deletion encompassing the NF1 locus and about a hundred genes in an NF1 contiguous gene syndrome patient Eric Pasmant*,1,2, Aure´lie de Saint-Trivier2, Ingrid Laurendeau1, Anne Dieux-Coeslier3, Be´atrice Parfait1,2, Michel Vidaud1,2, Dominique Vidaud1,2 and Ivan Bie`che1,2 1UMR745 INSERM, Universite´ Paris Descartes, Faculte´ des Sciences Pharmaceutiques et Biologiques, Paris, France; 2Service de Biochimie et de Ge´ne´tique Mole´culaire, Hoˆpital Beaujon AP-HP, Clichy, France; 3Service de Ge´ne´tique Clinique, Hoˆpital Jeanne de Flandre, Lille, France We describe a large germline deletion removing the NF1 locus, identified by heterozygosity mapping based on microsatellite markers, in an 8-year-old French girl with a particularly severe NF1 contiguous gene syndrome. We used gene-dose mapping with sequence-tagged site real-time PCR to locate the deletion end points, which were precisely characterized by means of long-range PCR and nucleotide sequencing. The deletion is located on chromosome arm 17q and is exactly 7 586 986 bp long. It encompasses the entire NF1 locus and about 100 other genes, including numerous chemokine genes, an attractive in silico-selected cerebrally expressed candidate gene (designated NUFIP2, for nuclear fragile X mental retardation protein interacting protein 2; NM_020772) and four microRNA genes. Interestingly, the centromeric breakpoint is located in intron 4 of the PIPOX gene (pipecolic acid oxidase; NM_016518) and the telomeric breakpoint in intron 5 of the GGNBP2 gene (gametogenetin binding protein 2; NM_024835) coding a transcription factor. -

Download Article (PDF)
Article in press - uncorrected proof BioMol Concepts, Vol. 2 (2011), pp. 343–352 • Copyright ᮊ by Walter de Gruyter • Berlin • Boston. DOI 10.1515/BMC.2011.033 Review Fragile X family members have important and non-overlapping functions Claudia Winograd2 and Stephanie Ceman1,2,* member, FMR1, was isolated by positional cloning of the X 1 Department of Cell and Developmental Biology, chromosomal region containing the inducible fragile site in University of Illinois, 601 S. Goodwin Avenue, individuals with fragile X syndrome (1). Cloning of the gene Urbana–Champaign, IL 61801, USA revealed the molecular defect to be a trinucleotide (CGG) 2 Neuroscience Program and College of Medicine, repeat expansion in exon 1 (2). Normally, individuals have University of Illinois, 601 S. Goodwin Avenue, less than 45 repeats with an average around 30 repeats; how- Urbana–Champaign, IL 61801, USA ever, expansion to greater than 200 repeats leads to aberrant methylation of the cytosines, leading to recruitment of * Corresponding author histone deacetylases with consequent transcriptional silenc- e-mail: [email protected] ing of the FMR1 locus (3). Thus, individuals with fragile X syndrome do not express transcript from the FMR1 locus. Abstract To identify the Xenopus laevis ortholog of FMR1 for further use in developmental studies, the human FMR1 gene was The fragile X family of genes encodes a small family of used to screen a cDNA library prepared from Xenopus laevis RNA binding proteins including FMRP, FXR1P and FXR2P ovary. In addition to identifying the Xenopus laevis ortholog that were identified in the 1990s. All three members are of FMR1, the first autosomal paralog FXR1 was discovered encoded by 17 exons and show alternative splicing at the 39 because of its sequence similarity to FMR1 (4). -

A-To-I RNA Editing Does Not Change with Age in the Healthy Male Rat Brain
Biogerontology (2013) 14:395–400 DOI 10.1007/s10522-013-9433-8 RESEARCH ARTICLE A-to-I RNA editing does not change with age in the healthy male rat brain Andrew P. Holmes • Shona H. Wood • Brian J. Merry • Joa˜o Pedro de Magalha˜es Received: 18 January 2013 / Accepted: 15 May 2013 / Published online: 26 May 2013 Ó The Author(s) 2013. This article is published with open access at Springerlink.com Abstract RNA editing is a post-transcriptional pro- Introduction cess, which results in base substitution modifications to RNA. It is an important process in generating Adenosine to inosine (A-to-I) RNA editing is a post- protein diversity through amino acid substitution and transcriptional process that alters the sequences of the modulation of splicing events. Previous studies RNA molecules. The adenosine deaminases ADAR have suggested a link between gene-specific reduc- and ADARB1 convert specific adenosine residues on tions in adenosine to inosine RNA editing and aging in RNA to inosine bases. During translation, sequencing, the human brain. Here we demonstrate that changes in and splicing, inosine is recognized as guanosine. RNA editing observed in humans with age are not Therefore, A-to-I RNA editing has important impli- observed during aging in healthy rats. Furthermore, we cations in altering specific amino acids, miRNA identify a conserved editing site in rats, in Cog3.We targeting, and in the modulation of alternative splicing propose that either age-related changes in RNA (Nishikura 2010). editing are specific to primates or humans, or that Targets of A-to-I RNA editing are often genes they are the manifestation of disease pathology. -

Spatially Clustering De Novo Variants in CYFIP2, Encoding the Cytoplasmic FMRP Interacting Protein 2, Cause Intellectual Disability and Seizures
European Journal of Human Genetics (2019) 27:747–759 https://doi.org/10.1038/s41431-018-0331-z ARTICLE Spatially clustering de novo variants in CYFIP2, encoding the cytoplasmic FMRP interacting protein 2, cause intellectual disability and seizures 1 1,2 3 3 2,4 5,6 Markus Zweier ● Anaïs Begemann ● Kirsty McWalter ● Megan T. Cho ● Lucia Abela ● Siddharth Banka ● 7 8 9,10 11 12 Bettina Behring ● Andrea Berger ● Chester W. Brown ● Maryline Carneiro ● Jiani Chen ● 13 14 13 Gregory M. Cooper ● Deciphering Developmental Disorders (DDD) Study ● Candice R. Finnila ● 3 15 16 1 5,6 17,18 Maria J. Guillen Sacoto ● Alex Henderson ● Ulrike Hüffmeier ● Pascal Joset ● Bronwyn Kerr ● Gaetan Lesca ● 19 5 20 3 9 Gloria S. Leszinski ● John Henry McDermott ● Meira R. Meltzer ● Kristin G. Monaghan ● Roya Mostafavi ● 21,22 2,4,23 24 12 21,22,25 Katrin Õunap ● Barbara Plecko ● Zöe Powis ● Gabriela Purcarin ● Tiia Reimand ● 19,26 20 27 12,28 29 Korbinian M. Riedhammer ● John M. Schreiber ● Deepa Sirsi ● Klaas J. Wierenga ● Monica H. Wojcik ● 1,30 1 31 1,2,32,33 Sorina M. Papuc ● Katharina Steindl ● Heinrich Sticht ● Anita Rauch Received: 30 May 2018 / Revised: 31 October 2018 / Accepted: 7 November 2018 / Published online: 21 January 2019 © European Society of Human Genetics 2019 1234567890();,: 1234567890();,: Abstract CYFIP2, encoding the evolutionary highly conserved cytoplasmic FMRP interacting protein 2, has previously been proposed as a candidate gene for intellectual disability and autism because of its important role linking FMRP-dependent transcription regulation and actin polymerization via the WAVE regulatory complex (WRC). Recently, de novo variants affecting the amino acid p.Arg87 of CYFIP2 were reported in four individuals with epileptic encephalopathy. -

A Fragile X Mental Retardation-Like Gene in a Cnidarian
Gene 343 (2004) 231–238 www.elsevier.com/locate/gene A fragile X mental retardation-like gene in a cnidarian Jasenka Guduric-Fuchsa, Frank Mfhrlena, Marcus Frohmeb, Uri Franka,* aInstitute of Zoology, University of Heidelberg, INF 230, Heidelberg 69120, Germany bDepartment of Functional Genome Analysis, German Cancer Research Center, INF 580, 69120 Heidelberg, Germany Received 6 August 2004; received in revised form 9 September 2004; accepted 5 October 2004 Available online 10 November 2004 Received by D.A. Tagle Abstract The fragile X mental retardation syndrome in humans is caused by a mutational loss of function of the fragile X mental retardation gene 1 (FMR1). FMR1 is an RNA-binding protein, involved in the development and function of the nervous system. Despite of its medical significance, the evolutionary origin of FMR1 has been unclear. Here, we report the molecular characterization of HyFMR1, an FMR1 orthologue, from the cnidarian hydroid Hydractinia echinata. Cnidarians are the most basal metazoans possessing neurons. HyFMR1is expressed throughout the life cycle of Hydractinia. Its expression pattern correlates to the position of neurons and their precursor stem cells in the animal. Our data indicate that the origin of the fraxile X related (FXR) protein family dates back at least to the common ancestor of cnidarians and bilaterians. The lack of FXR proteins in other invertebrates may have been due to gene loss in particular lineages. D 2004 Elsevier B.V. All rights reserved. Keywords: FMR1; FMRP; FXR; Hydractinia; Evolution; Hydrozoa 1. Introduction three types of RNA binding motifs: a ribonucleoprotein K homology domain (KH domain; FMR1 has two such The fragile X syndrome is the most common form of domains), an arginine and glycine rich domain (RGG box) inherited mental retardation in humans. -
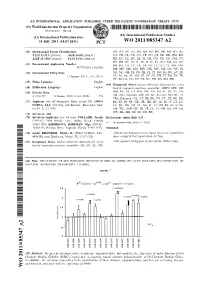
WO2011085347A2.Pdf
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date 14 July 2011 (14.07.2011) WO 2011/085347 A2 (51) International Patent Classification: AO, AT, AU, AZ, BA, BB, BG, BH, BR, BW, BY, BZ, C12N 15/113 (2010.01) A61K 48/00 (2006.01) CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, DO, A61K 31/7088 (2006.01) C12N 15/63 (2006.01) DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KM, KN, KP, (21) International Application Number: KR, KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, PCT/US201 1/020768 ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, (22) International Filing Date: NO, NZ, OM, PE, PG, PH, PL, PT, RO, RS, RU, SC, SD, 11 January 201 1 ( 11.01 .201 1) SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (25) Filing Language: English (84) Designated States (unless otherwise indicated, for every (26) Publication Language: English kind of regional protection available): ARIPO (BW, GH, (30) Priority Data: GM, KE, LR, LS, MW, MZ, NA, SD, SL, SZ, TZ, UG, 61/293,739 11 January 2010 ( 11.01 .2010) US ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, MD, RU, TJ, TM), European (AL, AT, BE, BG, CH, CY, CZ, DE, DK, (71) Applicant (for all designated States except US): OPKO EE, ES, FI, FR, GB, GR, HR, HU, IE, IS, ΓΓ, LT, LU, CURNA, LLC [US/US]; 440 Biscayne Boulevard, Mia LV, MC, MK, MT, NL, NO, PL, PT, RO, RS, SE, SI, SK, mi, FL 33 137 (US). -

Comparative RNA Editing in Autistic and Neurotypical Cerebella
Molecular Psychiatry (2013) 18, 1041–1048 & 2013 Macmillan Publishers Limited All rights reserved 1359-4184/13 www.nature.com/mp ORIGINAL ARTICLE Comparative RNA editing in autistic and neurotypical cerebella A Eran1,2,JBLi3, K Vatalaro2, J McCarthy2, F Rahimov2,4, C Collins2,4, K Markianos2,5, DM Margulies5,6,7, EN Brown1,8,9, SE Calvo10, IS Kohane1,5,7 and LM Kunkel2,4,5,11 Adenosine-to-inosine (A-to-I) RNA editing is a neurodevelopmentally regulated epigenetic modification shown to modulate complex behavior in animals. Little is known about human A-to-I editing, but it is thought to constitute one of many molecular mechanisms connecting environmental stimuli and behavioral outputs. Thus, comprehensive exploration of A-to-I RNA editing in human brains may shed light on gene–environment interactions underlying complex behavior in health and disease. Synaptic function is a main target of A-to-I editing, which can selectively recode key amino acids in synaptic genes, directly altering synaptic strength and duration in response to environmental signals. Here, we performed a high-resolution survey of synaptic A-to-I RNA editing in a human population, and examined how it varies in autism, a neurodevelopmental disorder in which synaptic abnormalities are a common finding. Using ultra-deep (41000 Â ) sequencing, we quantified the levels of A-to-I editing of 10 synaptic genes in postmortem cerebella from 14 neurotypical and 11 autistic individuals. A high dynamic range of editing levels was detected across individuals and editing sites, from 99.6% to below detection limits. In most sites, the extreme ends of the population editing distributions were individuals with autism. -

Altered Adenosine-To-Inosine RNA Editing in Human Cancer
Downloaded from genome.cshlp.org on September 26, 2021 - Published by Cold Spring Harbor Laboratory Press Letter Altered adenosine-to-inosine RNA editing in human cancer Nurit Paz,1,2 Erez Y. Levanon,3,12 Ninette Amariglio,1,2 Amy B. Heimberger,4 Zvi Ram,5 Shlomi Constantini,6 Zohar S. Barbash,1,2 Konstantin Adamsky,1 Michal Safran,1,2 Avi Hirschberg,1,2 Meir Krupsky,2,7 Issachar Ben-Dov,2,8 Simona Cazacu,9 Tom Mikkelsen,9 Chaya Brodie,9,10 Eli Eisenberg,11 and Gideon Rechavi1,2,13 1Cancer Research Center, Chaim Sheba Medical Center, Tel Hashomer 52621, Israel; 2Sackler School of Medicine, Tel Aviv University, Tel Aviv 69978, Israel; 3Compugen Ltd., Tel Aviv 69512, Israel; 4Department of Neurosurgery, Brain Tumor Center, University of Texas M.D. Anderson Cancer Center, Houston 77030, Texas, USA; 5Department of Neurosurgery, Sourasky Medical Center, Tel Aviv 64239, Israel; 6Department of Pediatric Neurosurgery, Dana Children’s Hospital, Sourasky Medical Center, Tel Aviv 64239, Israel; 7Department of Internal Medicine, Chaim Sheba Medical Center, Tel Hashomer 52621, Israel; 8Pulmonary Institute, Chaim Sheba Medical Center, Tel Hashomer 52621, Israel; 9Hermelin Brain Tumor Center, Department of Neurosurgery, Henry Ford Hospital, Detroit, Michigan 48202, USA; 10Neuro-Oncology Branch, NCI/NINDS, NIH, Bethesda 20892, Maryland, USA; 11School of Physics and Astronomy, Raymond and Beverly Sackler Faculty of Exact Sciences, Tel Aviv University 69978 Israel Adenosine-to-inosine (A-to-I) RNA editing was recently shown to be abundant in the human transcriptome, affecting thousands of genes. Employing a bioinformatic approach, we identified significant global hypoediting of Alu repetitive elements in brain, prostate, lung, kidney, and testis tumors. -

Anti-DDX21 Pab
For Research Use Only. RN090PW Page 1 Not for use in diagnostic procedures. Anti-DDX21 pAb CODE No. RN090PW CLONALITY Polyclonal ISOTYPE Rabbit Ig, affinity purified QUANTITY 100 L, 1 mg/mL SOURCE Purified Ig from rabbit serum FORMURATION PBS containing 50% Glycerol (pH 7.2). No preservative is contained. STORAGE This antibody solution is stable for one year from the date of purchase when stored at -20°C. APPLICATIONS-CONFIRMED Western blotting 1:1,000 for chemiluminescence detection system Immunoprecipitation 5 L/500 L of cell extract from 2 x 107 cells APPLICATIONS-UNDER EVALUATION Immunocytochemistry 1:400 SPECIES CROSS REACTIVITY on WB Species Human Mouse Rat Hamster Cell HeLa, A431, 293T Not tested Not tested Not tested Reactivity Entrez Gene ID 9188 (Human) For more information, please visit our web site https://ruo.mbl.co.jp/je/rip-assay/ LICENSING OPPORTUNITY: The RIP-Assay uses patented technology (US patent No. 6,635,422, US patent No. 7,504,210) of Ribonomics, Inc. MBL manufactures and distributes this product under license from Ribonomics, Inc. Researchers may use this product for their own research. Researchers are not allowed to use this product or RIP-Assay technology for commercial purpose without a license. For commercial use, please contact us for licensing opportunities at [email protected] MEDICAL & BIOLOGICAL LABORATORIES CO., LTD. URL https://ruo.mbl.co.jp/je/rip-assay/ e-mail [email protected], TEL 052-238-1904 RN090PW Page 2 RELATED PRODUCTS RN047PW Anti-PTBP2 (polyclonal) RN048PW Anti-G3BP1 (polyclonal) RIP-Assay -

Comparative Behavioral Phenotypes of Fmr1 KO, Fxr2 Het, and Fmr1 KO/Fxr2 Het Mice
brain sciences Article Comparative Behavioral Phenotypes of Fmr1 KO, Fxr2 Het, and Fmr1 KO/Fxr2 Het Mice Rachel Michelle Saré, Christopher Figueroa, Abigail Lemons, Inna Loutaev and Carolyn Beebe Smith * Section on Neuroadaptation and Protein Metabolism, National Institute of Mental Health, National Institutes of Health, Department of Health and Human Services, Bethesda, MD 20814, USA; [email protected] (R.M.S.); [email protected] (C.F.); [email protected] (A.L.); [email protected] (I.L.) * Correspondence: [email protected]; Tel.: +1-301-402-3120 Received: 6 November 2018; Accepted: 10 January 2019; Published: 16 January 2019 Abstract: Fragile X syndrome (FXS) is caused by silencing of the FMR1 gene leading to loss of the protein product fragile X mental retardation protein (FMRP). FXS is the most common monogenic cause of intellectual disability. There are two known mammalian paralogs of FMRP,FXR1P,and FXR2P. The functions of FXR1P and FXR2P and their possible roles in producing or modulating the phenotype observed in FXS are yet to be identified. Previous studies have revealed that mice lacking Fxr2 display similar behavioral abnormalities as Fmr1 knockout (KO) mice. In this study, we expand upon the +/ behavioral phenotypes of Fmr1 KO and Fxr2 − (Het) mice and compare them with Fmr1 KO/Fxr2 Het mice. We find that Fmr1 KO and Fmr1 KO/Fxr2 Het mice are similarly hyperactive compared to WT and Fxr2 Het mice. Fmr1 KO/Fxr2 Het mice have more severe learning and memory impairments than Fmr1 KO mice. Fmr1 KO mice display significantly impaired social behaviors compared to WT mice, which are paradoxically reversed in Fmr1 KO/Fxr2 Het mice.