Human Platelet Glycoprotein Ib Beta Chain(GP1BB
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Hemoglobin Interaction with Gp1ba Induces Platelet Activation And
ARTICLE Platelet Biology & its Disorders Hemoglobin interaction with GP1bα induces platelet activation and apoptosis: a novel mechanism associated with intravascular hemolysis Rashi Singhal,1,2,* Gowtham K. Annarapu,1,2,* Ankita Pandey,1 Sheetal Chawla,1 Amrita Ojha,1 Avinash Gupta,1 Miguel A. Cruz,3 Tulika Seth4 and Prasenjit Guchhait1 1Disease Biology Laboratory, Regional Centre for Biotechnology, National Capital Region, Biotech Science Cluster, Faridabad, India; 2Biotechnology Department, Manipal University, Manipal, Karnataka, India; 3Thrombosis Research Division, Baylor College of Medicine, Houston, TX, USA, and 4Hematology, All India Institute of Medical Sciences, New Delhi, India *RS and GKA contributed equally to this work. ABSTRACT Intravascular hemolysis increases the risk of hypercoagulation and thrombosis in hemolytic disorders. Our study shows a novel mechanism by which extracellular hemoglobin directly affects platelet activation. The binding of Hb to glycoprotein1bα activates platelets. Lower concentrations of Hb (0.37-3 mM) significantly increase the phos- phorylation of signaling adapter proteins, such as Lyn, PI3K, AKT, and ERK, and promote platelet aggregation in vitro. Higher concentrations of Hb (3-6 mM) activate the pro-apoptotic proteins Bak, Bax, cytochrome c, caspase-9 and caspase-3, and increase platelet clot formation. Increased plasma Hb activates platelets and promotes their apoptosis, and plays a crucial role in the pathogenesis of aggregation and development of the procoagulant state in hemolytic disorders. Furthermore, we show that in patients with paroxysmal nocturnal hemoglobinuria, a chronic hemolytic disease characterized by recurrent events of intravascular thrombosis and thromboembolism, it is the elevated plasma Hb or platelet surface bound Hb that positively correlates with platelet activation. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -

Genome-Wide DNA Methylation Analysis of KRAS Mutant Cell Lines Ben Yi Tew1,5, Joel K
www.nature.com/scientificreports OPEN Genome-wide DNA methylation analysis of KRAS mutant cell lines Ben Yi Tew1,5, Joel K. Durand2,5, Kirsten L. Bryant2, Tikvah K. Hayes2, Sen Peng3, Nhan L. Tran4, Gerald C. Gooden1, David N. Buckley1, Channing J. Der2, Albert S. Baldwin2 ✉ & Bodour Salhia1 ✉ Oncogenic RAS mutations are associated with DNA methylation changes that alter gene expression to drive cancer. Recent studies suggest that DNA methylation changes may be stochastic in nature, while other groups propose distinct signaling pathways responsible for aberrant methylation. Better understanding of DNA methylation events associated with oncogenic KRAS expression could enhance therapeutic approaches. Here we analyzed the basal CpG methylation of 11 KRAS-mutant and dependent pancreatic cancer cell lines and observed strikingly similar methylation patterns. KRAS knockdown resulted in unique methylation changes with limited overlap between each cell line. In KRAS-mutant Pa16C pancreatic cancer cells, while KRAS knockdown resulted in over 8,000 diferentially methylated (DM) CpGs, treatment with the ERK1/2-selective inhibitor SCH772984 showed less than 40 DM CpGs, suggesting that ERK is not a broadly active driver of KRAS-associated DNA methylation. KRAS G12V overexpression in an isogenic lung model reveals >50,600 DM CpGs compared to non-transformed controls. In lung and pancreatic cells, gene ontology analyses of DM promoters show an enrichment for genes involved in diferentiation and development. Taken all together, KRAS-mediated DNA methylation are stochastic and independent of canonical downstream efector signaling. These epigenetically altered genes associated with KRAS expression could represent potential therapeutic targets in KRAS-driven cancer. Activating KRAS mutations can be found in nearly 25 percent of all cancers1. -

Datasheet: MCA740PE Product Details
Datasheet: MCA740PE Description: MOUSE ANTI HUMAN CD42b:RPE Specificity: CD42b Other names: GPIB-ALPHA Format: RPE Product Type: Monoclonal Antibody Clone: AK2 Isotype: IgG1 Quantity: 100 TESTS Product Details Applications This product has been reported to work in the following applications. This information is derived from testing within our laboratories, peer-reviewed publications or personal communications from the originators. Please refer to references indicated for further information. For general protocol recommendations, please visit www.bio-rad-antibodies.com/protocols. Yes No Not Determined Suggested Dilution Flow Cytometry Neat Where this antibody has not been tested for use in a particular technique this does not necessarily exclude its use in such procedures. The suggested working dilution is given as a guide only. It is recommended that the user titrates the antibody for use in his/her own system using appropriate negative/positive controls. Target Species Human Product Form Purified IgG conjugated to R. Phycoerythrin (RPE) - lyophilized Reconstitution Reconstitute with 1 ml distilled water Max Ex/Em Fluorophore Excitation Max (nm) Emission Max (nm) RPE 488nm laser 496 578 Preparation Purified IgG prepared by affinity chromatography on Protein A from tissue culture supernatant Buffer Solution Phosphate buffered saline Preservative 0.09% Sodium Azide Stabilisers 1% Bovine Serum Albumin 5% Sucrose External Database Links UniProt: P07359 Related reagents Entrez Gene: 2811 GP1BA Related reagents Page 1 of 3 Specificity Mouse anti Human CD42b antibody, clone AK2 recognizes the human CD42b cell surface antigen, also known as platelet glycoprotein GP1B. CD42b is expressed by platelets and megakaryocytes. Clone AK2 has been reported to block the binding of von Willebrand Factor (VWF) to platelets Flow Cytometry Use 10ul of the suggested working dilution to label 100ul whole blood. -

NICU Gene List Generator.Xlsx
Neonatal Crisis Sequencing Panel Gene List Genes: A2ML1 - B3GLCT A2ML1 ADAMTS9 ALG1 ARHGEF15 AAAS ADAMTSL2 ALG11 ARHGEF9 AARS1 ADAR ALG12 ARID1A AARS2 ADARB1 ALG13 ARID1B ABAT ADCY6 ALG14 ARID2 ABCA12 ADD3 ALG2 ARL13B ABCA3 ADGRG1 ALG3 ARL6 ABCA4 ADGRV1 ALG6 ARMC9 ABCB11 ADK ALG8 ARPC1B ABCB4 ADNP ALG9 ARSA ABCC6 ADPRS ALK ARSL ABCC8 ADSL ALMS1 ARX ABCC9 AEBP1 ALOX12B ASAH1 ABCD1 AFF3 ALOXE3 ASCC1 ABCD3 AFF4 ALPK3 ASH1L ABCD4 AFG3L2 ALPL ASL ABHD5 AGA ALS2 ASNS ACAD8 AGK ALX3 ASPA ACAD9 AGL ALX4 ASPM ACADM AGPS AMELX ASS1 ACADS AGRN AMER1 ASXL1 ACADSB AGT AMH ASXL3 ACADVL AGTPBP1 AMHR2 ATAD1 ACAN AGTR1 AMN ATL1 ACAT1 AGXT AMPD2 ATM ACE AHCY AMT ATP1A1 ACO2 AHDC1 ANK1 ATP1A2 ACOX1 AHI1 ANK2 ATP1A3 ACP5 AIFM1 ANKH ATP2A1 ACSF3 AIMP1 ANKLE2 ATP5F1A ACTA1 AIMP2 ANKRD11 ATP5F1D ACTA2 AIRE ANKRD26 ATP5F1E ACTB AKAP9 ANTXR2 ATP6V0A2 ACTC1 AKR1D1 AP1S2 ATP6V1B1 ACTG1 AKT2 AP2S1 ATP7A ACTG2 AKT3 AP3B1 ATP8A2 ACTL6B ALAS2 AP3B2 ATP8B1 ACTN1 ALB AP4B1 ATPAF2 ACTN2 ALDH18A1 AP4M1 ATR ACTN4 ALDH1A3 AP4S1 ATRX ACVR1 ALDH3A2 APC AUH ACVRL1 ALDH4A1 APTX AVPR2 ACY1 ALDH5A1 AR B3GALNT2 ADA ALDH6A1 ARFGEF2 B3GALT6 ADAMTS13 ALDH7A1 ARG1 B3GAT3 ADAMTS2 ALDOB ARHGAP31 B3GLCT Updated: 03/15/2021; v.3.6 1 Neonatal Crisis Sequencing Panel Gene List Genes: B4GALT1 - COL11A2 B4GALT1 C1QBP CD3G CHKB B4GALT7 C3 CD40LG CHMP1A B4GAT1 CA2 CD59 CHRNA1 B9D1 CA5A CD70 CHRNB1 B9D2 CACNA1A CD96 CHRND BAAT CACNA1C CDAN1 CHRNE BBIP1 CACNA1D CDC42 CHRNG BBS1 CACNA1E CDH1 CHST14 BBS10 CACNA1F CDH2 CHST3 BBS12 CACNA1G CDK10 CHUK BBS2 CACNA2D2 CDK13 CILK1 BBS4 CACNB2 CDK5RAP2 -
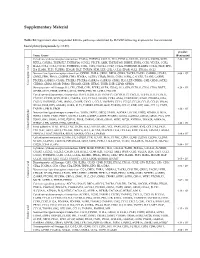
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R, -

Path Ggf 5 2020.Pdf
Hemostasis Hemostasis and Thrombosis Normal hemostasis is a consequence of tightly regulated processes that maintain blood in a fluid state in normal vessels, yet also permit the rapid formation of a hemostatic clot at the site of a vascular injury. Thrombosis involves blood clot formation within intact vessels. Both hemostasis and thrombosis involve three components: the vascular wall, platelets and the coagulation cascade. Elements of the Hemostatic process • Endothelium • Anti-thrombosis • Pro-thrombosis • Platelets • Platelet-endothelial cell interaction • Coagulation cascade http://www.as.miami.edu/chemistry/2086/chapter_21/NEW-Chap21_class_part1_files/image002.jpg After initial injury there is a brief period of arteriolar vasoconstriction mediated by reflex neurogenic mechanisms and augmented by the local secretion of factors such as endothelin (a potent endothelium-derived vasoconstrictor) The effect is transient, however, and bleeding would resume if not for activation of the platelet and coagulation systems. Endothelial injury exposes highly thrombogenic subendothelial extracellular matrix (ECM), facilitating platelet adherence and activation. Activation of platelets results in a dramatic shape change (from small rounded discs to flat plates with markedly increased surface area), as well as the release of secretory granules. Within minutes the secreted products recruit additional platelets (aggregation) to form a hemostatic plug; this process is referred to as primary hemostasis. http://www.ouhsc.edu/platelets/Platelet%20Pic s/Platelets3.jpg http://medcell.med.yale.edu/histology/blood_bone_marr ow_lab/images/platelets_em.jpg Tissue factor is also exposed at the site of injury. Also known as factor III and thromboplastin, tissue factor is a membrane-bound procoagulant glycoprotein synthesized by endothelial cells. It acts in conjunction with factor VII (see below) as the major in vivo initiator of the coagulation cascade, eventually culminating in thrombin generation. -

Biomechanical Thrombosis: the Dark Side of Force and Dawn of Mechano- Medicine
Open access Review Stroke Vasc Neurol: first published as 10.1136/svn-2019-000302 on 15 December 2019. Downloaded from Biomechanical thrombosis: the dark side of force and dawn of mechano- medicine Yunfeng Chen ,1 Lining Arnold Ju 2 To cite: Chen Y, Ju LA. ABSTRACT P2Y12 receptor antagonists (clopidogrel, pras- Biomechanical thrombosis: the Arterial thrombosis is in part contributed by excessive ugrel, ticagrelor), inhibitors of thromboxane dark side of force and dawn platelet aggregation, which can lead to blood clotting and A2 (TxA2) generation (aspirin, triflusal) or of mechano- medicine. Stroke subsequent heart attack and stroke. Platelets are sensitive & Vascular Neurology 2019;0. protease- activated receptor 1 (PAR1) antag- to the haemodynamic environment. Rapid haemodynamcis 1 doi:10.1136/svn-2019-000302 onists (vorapaxar). Increasing the dose of and disturbed blood flow, which occur in vessels with these agents, especially aspirin and clopi- growing thrombi and atherosclerotic plaques or is caused YC and LAJ contributed equally. dogrel, has been employed to dampen the by medical device implantation and intervention, promotes Received 12 November 2019 platelet thrombotic functions. However, this platelet aggregation and thrombus formation. In such 4 Accepted 14 November 2019 situations, conventional antiplatelet drugs often have also increases the risk of excessive bleeding. suboptimal efficacy and a serious side effect of excessive It has long been recognized that arterial bleeding. Investigating the mechanisms of platelet thrombosis -

Human Induced Pluripotent Stem Cell–Derived Podocytes Mature Into Vascularized Glomeruli Upon Experimental Transplantation
BASIC RESEARCH www.jasn.org Human Induced Pluripotent Stem Cell–Derived Podocytes Mature into Vascularized Glomeruli upon Experimental Transplantation † Sazia Sharmin,* Atsuhiro Taguchi,* Yusuke Kaku,* Yasuhiro Yoshimura,* Tomoko Ohmori,* ‡ † ‡ Tetsushi Sakuma, Masashi Mukoyama, Takashi Yamamoto, Hidetake Kurihara,§ and | Ryuichi Nishinakamura* *Department of Kidney Development, Institute of Molecular Embryology and Genetics, and †Department of Nephrology, Faculty of Life Sciences, Kumamoto University, Kumamoto, Japan; ‡Department of Mathematical and Life Sciences, Graduate School of Science, Hiroshima University, Hiroshima, Japan; §Division of Anatomy, Juntendo University School of Medicine, Tokyo, Japan; and |Japan Science and Technology Agency, CREST, Kumamoto, Japan ABSTRACT Glomerular podocytes express proteins, such as nephrin, that constitute the slit diaphragm, thereby contributing to the filtration process in the kidney. Glomerular development has been analyzed mainly in mice, whereas analysis of human kidney development has been minimal because of limited access to embryonic kidneys. We previously reported the induction of three-dimensional primordial glomeruli from human induced pluripotent stem (iPS) cells. Here, using transcription activator–like effector nuclease-mediated homologous recombination, we generated human iPS cell lines that express green fluorescent protein (GFP) in the NPHS1 locus, which encodes nephrin, and we show that GFP expression facilitated accurate visualization of nephrin-positive podocyte formation in -

Gene Expression Analysis of Mevalonate Kinase Deficiency
International Journal of Environmental Research and Public Health Article Gene Expression Analysis of Mevalonate Kinase Deficiency Affected Children Identifies Molecular Signatures Related to Hematopoiesis Simona Pisanti * , Marianna Citro , Mario Abate , Mariella Caputo and Rosanna Martinelli * Department of Medicine, Surgery and Dentistry ‘Scuola Medica Salernitana’, University of Salerno, Via Salvatore Allende, 84081 Baronissi (SA), Italy; [email protected] (M.C.); [email protected] (M.A.); [email protected] (M.C.) * Correspondence: [email protected] (S.P.); [email protected] (R.M.) Abstract: Mevalonate kinase deficiency (MKD) is a rare autoinflammatory genetic disorder charac- terized by recurrent fever attacks and systemic inflammation with potentially severe complications. Although it is recognized that the lack of protein prenylation consequent to mevalonate pathway blockade drives IL1β hypersecretion, and hence autoinflammation, MKD pathogenesis and the molecular mechanisms underlaying most of its clinical manifestations are still largely unknown. In this study, we performed a comprehensive bioinformatic analysis of a microarray dataset of MKD patients, using gene ontology and Ingenuity Pathway Analysis (IPA) tools, in order to identify the most significant differentially expressed genes and infer their predicted relationships into biological processes, pathways, and networks. We found that hematopoiesis linked biological functions and pathways are predominant in the gene ontology of differentially expressed genes in MKD, in line with the observed clinical feature of anemia. We also provided novel information about the molecular Citation: Pisanti, S.; Citro, M.; Abate, M.; Caputo, M.; Martinelli, R. mechanisms at the basis of the hematological abnormalities observed, that are linked to the chronic Gene Expression Analysis of inflammation and to defective prenylation.