New Nomenclature Adhesion-Gpcrs April 22, 2015
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplemental Figure 1. Vimentin
Double mutant specific genes Transcript gene_assignment Gene Symbol RefSeq FDR Fold- FDR Fold- FDR Fold- ID (single vs. Change (double Change (double Change wt) (single vs. wt) (double vs. single) (double vs. wt) vs. wt) vs. single) 10485013 BC085239 // 1110051M20Rik // RIKEN cDNA 1110051M20 gene // 2 E1 // 228356 /// NM 1110051M20Ri BC085239 0.164013 -1.38517 0.0345128 -2.24228 0.154535 -1.61877 k 10358717 NM_197990 // 1700025G04Rik // RIKEN cDNA 1700025G04 gene // 1 G2 // 69399 /// BC 1700025G04Rik NM_197990 0.142593 -1.37878 0.0212926 -3.13385 0.093068 -2.27291 10358713 NM_197990 // 1700025G04Rik // RIKEN cDNA 1700025G04 gene // 1 G2 // 69399 1700025G04Rik NM_197990 0.0655213 -1.71563 0.0222468 -2.32498 0.166843 -1.35517 10481312 NM_027283 // 1700026L06Rik // RIKEN cDNA 1700026L06 gene // 2 A3 // 69987 /// EN 1700026L06Rik NM_027283 0.0503754 -1.46385 0.0140999 -2.19537 0.0825609 -1.49972 10351465 BC150846 // 1700084C01Rik // RIKEN cDNA 1700084C01 gene // 1 H3 // 78465 /// NM_ 1700084C01Rik BC150846 0.107391 -1.5916 0.0385418 -2.05801 0.295457 -1.29305 10569654 AK007416 // 1810010D01Rik // RIKEN cDNA 1810010D01 gene // 7 F5 // 381935 /// XR 1810010D01Rik AK007416 0.145576 1.69432 0.0476957 2.51662 0.288571 1.48533 10508883 NM_001083916 // 1810019J16Rik // RIKEN cDNA 1810019J16 gene // 4 D2.3 // 69073 / 1810019J16Rik NM_001083916 0.0533206 1.57139 0.0145433 2.56417 0.0836674 1.63179 10585282 ENSMUST00000050829 // 2010007H06Rik // RIKEN cDNA 2010007H06 gene // --- // 6984 2010007H06Rik ENSMUST00000050829 0.129914 -1.71998 0.0434862 -2.51672 -

Edinburgh Research Explorer
Edinburgh Research Explorer International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list Citation for published version: Davenport, AP, Alexander, SPH, Sharman, JL, Pawson, AJ, Benson, HE, Monaghan, AE, Liew, WC, Mpamhanga, CP, Bonner, TI, Neubig, RR, Pin, JP, Spedding, M & Harmar, AJ 2013, 'International Union of Basic and Clinical Pharmacology. LXXXVIII. G protein-coupled receptor list: recommendations for new pairings with cognate ligands', Pharmacological reviews, vol. 65, no. 3, pp. 967-86. https://doi.org/10.1124/pr.112.007179 Digital Object Identifier (DOI): 10.1124/pr.112.007179 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: Pharmacological reviews Publisher Rights Statement: U.S. Government work not protected by U.S. copyright General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 02. Oct. 2021 1521-0081/65/3/967–986$25.00 http://dx.doi.org/10.1124/pr.112.007179 PHARMACOLOGICAL REVIEWS Pharmacol Rev 65:967–986, July 2013 U.S. -

Supplementary Data
Supplemental Data A novel mouse model of X-linked nephrogenic diabetes insipidus: Phenotypic analysis and therapeutic implications Jian Hua Li, Chung-Lin Chou, Bo Li, Oksana Gavrilova, Christoph Eisner, Jürgen Schnermann, Stasia A. Anderson, Chu-Xia Deng, Mark A. Knepper, and Jürgen Wess Supplemental Methods Metabolic cage studies. Animals were maintained in mouse metabolic cages (Hatteras Instruments, Cary, NC) under controlled temperature and light conditions (12 hr light and dark cycles). Mice received a fixed daily ration of 6.5 g of gelled diet per 20 g of body weight per day. The gelled diet was composed of 4 g of Basal Diet 5755 (Test Diet, Richmond, IN), 2.5 ml of deionized water, and 65 mg agar. Preweighted drinking water was provided ad libitum during the course of the study. Mice were acclimated in the metabolic cages for 1-2 days. Urine was collected under mineral oil in preweighted collection vials for successive 24 hr periods. Analysis of GPCR expression in mouse IMCD cells via TaqMan real-time qRT-PCR. Total RNA prepared from mouse IMCD tubule suspensions was reverse transcribed as described under Experimental Procedures. Tissues from ten 10-week old C57BL/6 WT mice were collected and pooled for each individual experiment. cDNA derived from 640 ng of RNA was mixed with an equal volume of TaqMan gene expression 2 x master mix (Applied Biosystems, Foster City, CA). 100 μl-aliquots of this mixture (corresponding to 80 ng of RNA) were added to each of the 8 fill ports of a 384-well plate of a mouse GPCR array panel (Applied Biosystems). -

WO 2019/067413 Al 04 April 2019 (04.04.2019) W 1P O PCT
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2019/067413 Al 04 April 2019 (04.04.2019) W 1P O PCT (51) International Patent Classification: 16/139,608 24 September 2018 (24.09.2018) US A61K 31/137 (2006.01) A61K 45/06 (2006.01) 16/139,617 24 September 2018 (24.09.2018) US A61P 25/08 (2006.01) 16/139,698 24 September 2018 (24.09.2018) US 16/139,701 24 September 2018 (24.09.2018) US (21) International Application Number: 16/139,704 24 September 2018 (24.09.2018) US PCT/US2018/052580 16/139,763 24 September 2018 (24.09.2018) US (22) International Filing Date: (71) Applicant: ZOGENIX INTERNATIONAL LIMITED 25 September 2018 (25.09.2018) [GB/GB]; Siena Court Broadway, Maidenhead Berkshire (25) Filing Language: English SL6 1NJ (GB). (26) Publication Language: English (72) Inventors; and (71) Applicants: FARFEL, Gail [US/US]; c/o Zogenix, Inc., (30) 5858 Horton Street, Suite 455, Emeryville, California 94608 26 September 2017 (26.09.2017) US (US). LOCK, Michael [US/US]; c/o Zogenix, Inc., 5858 27 September 2017 (27.09.2017) US Horton Street, Suite 455, Emeryville, California 94608 31 October 2017 (3 1. 10.2017) US (US). GALER, Bradley S. [US/US]; 1740 Lenape Road, 30 November 2017 (30. 11.2017) US West Chester, Pennsylvania 19382 (US). MORRISON, 07 February 2018 (07.02.2018) US Glenn [CA/US]; c/o Zogenix, Inc., 5858 Horton Street, 10 May 2018 (10.05.2018) US Suite 455, Emeryville, California 94608 (US). -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Monitoring Nociception by Analyzing Gene Expression Changes in the Central Nervous System of Mice
Zurich Open Repository and Archive University of Zurich Main Library Strickhofstrasse 39 CH-8057 Zurich www.zora.uzh.ch Year: 2010 Monitoring nociception by analyzing gene expression changes in the central nervous system of mice Asner, I N Posted at the Zurich Open Repository and Archive, University of Zurich ZORA URL: https://doi.org/10.5167/uzh-46678 Dissertation Originally published at: Asner, I N. Monitoring nociception by analyzing gene expression changes in the central nervous system of mice. 2010, University of Zurich, Vetsuisse Faculty. Monitoring Nociception by Analyzing Gene Expression Changes in the Central Nervous System of Mice Dissertation zur Erlangung der naturwissenschaftlichen Doktorwürde (Dr. sc. nat) vorgelegt der Mathematisch-naturwissenschaftlichen Fakultät der Universität Zürich von Igor Asner von St. Cergue VD Promotionskomitee Prof. Dr. Peter Sonderegger Prof. Dr. Kurt Bürki Prof. Dr. Hanns Ulrich Zeilhofer Dr. Paolo Cinelli (Leitung der Dissertation) Zürich, 2010 Table of contents Table of content Curriculum vitae 6 Publications 9 Summary 11 Zusammenfassung 14 1. Introduction 17 1.1. Pain and nociception 17 1.1.1 Nociceptive neurons and Mechanoceptors 18 1.1.2 Activation of the nociceptive neurons at the periphery 21 1.1.2.1 Response to noxious heat 22 1.1.2.2 Response to noxious cold 23 1.1.2.3 Response to mechanical stress 24 1.1.3 Nociceptive message processing in the Spinal Cord 25 1.1.3.1 The lamina I and the ascending pathways 25 1.1.3.2 The lamina II and the descending pathways 26 1.1.4 Pain processing and integration in the brain 27 1.1.4.1 The Pain Matrix 27 1.1.4.2 Activation of the descending pathways 29 1.1.5 Inflammatory Pain 31 1.2. -

Flow Reagents Single Color Antibodies CD Chart
CD CHART CD N° Alternative Name CD N° Alternative Name CD N° Alternative Name Beckman Coulter Clone Beckman Coulter Clone Beckman Coulter Clone T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells CD1a T6, R4, HTA1 Act p n n p n n S l CD99 MIC2 gene product, E2 p p p CD223 LAG-3 (Lymphocyte activation gene 3) Act n Act p n CD1b R1 Act p n n p n n S CD99R restricted CD99 p p CD224 GGT (γ-glutamyl transferase) p p p p p p CD1c R7, M241 Act S n n p n n S l CD100 SEMA4D (semaphorin 4D) p Low p p p n n CD225 Leu13, interferon induced transmembrane protein 1 (IFITM1). p p p p p CD1d R3 Act S n n Low n n S Intest CD101 V7, P126 Act n p n p n n p CD226 DNAM-1, PTA-1 Act n Act Act Act n p n CD1e R2 n n n n S CD102 ICAM-2 (intercellular adhesion molecule-2) p p n p Folli p CD227 MUC1, mucin 1, episialin, PUM, PEM, EMA, DF3, H23 Act p CD2 T11; Tp50; sheep red blood cell (SRBC) receptor; LFA-2 p S n p n n l CD103 HML-1 (human mucosal lymphocytes antigen 1), integrin aE chain S n n n n n n n l CD228 Melanotransferrin (MT), p97 p p CD3 T3, CD3 complex p n n n n n n n n n l CD104 integrin b4 chain; TSP-1180 n n n n n n n p p CD229 Ly9, T-lymphocyte surface antigen p p n p n -

ADGRE1 (EMR1, F4/80) Is a Rapidly-Evolving Gene Expressed In
Edinburgh Research Explorer ADGRE1 (EMR1, F4/80) Is a Rapidly-Evolving Gene Expressed in Mammalian Monocyte-Macrophages Citation for published version: Waddell, L, Lefevre, L, Bush, S, Raper, A, Young, R, Lisowski, Z, McCulloch, M, Muriuki, C, Sauter, K, Clark, E, Irvine, KN, Pridans, C, Hope, J & Hume, D 2018, 'ADGRE1 (EMR1, F4/80) Is a Rapidly-Evolving Gene Expressed in Mammalian Monocyte-Macrophages' Frontiers in Immunology, vol. 9, 2246. DOI: 10.3389/fimmu.2018.02246 Digital Object Identifier (DOI): 10.3389/fimmu.2018.02246 Link: Link to publication record in Edinburgh Research Explorer Document Version: Publisher's PDF, also known as Version of record Published In: Frontiers in Immunology General rights Copyright for the publications made accessible via the Edinburgh Research Explorer is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The University of Edinburgh has made every reasonable effort to ensure that Edinburgh Research Explorer content complies with UK legislation. If you believe that the public display of this file breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. Download date: 05. Apr. 2019 ORIGINAL RESEARCH published: 01 October 2018 doi: 10.3389/fimmu.2018.02246 ADGRE1 (EMR1, F4/80) Is a Rapidly-Evolving Gene Expressed in Mammalian Monocyte-Macrophages Lindsey A. Waddell 1†, Lucas Lefevre 1†, Stephen J. Bush 2†, Anna Raper 1, Rachel Young 1, Zofia M. -

Supplemental Material For
Supplemental material for Epithelial to mesenchymal transition rewires the molecular path to PI3-Kinase-dependent proliferation Megan B. Salt1,2, Sourav Bandyopadhyay1,3 and Frank McCormick1 Authors Affiliations: 1-Helen Diller Family Comprehensive Cancer Center; 2-Biomedical Sciences Graduate Program, University of California San Francisco, San Francisco, California; 3-California Institute for Quantitative Biosciences, San Francisco, California. Supplemental Figure Legends Table S1. EMT associated changes in gene expression. Significantly (A) up- and (B) down- regulated genes in comparing H358-TwistER cells to control H358-GFP expressing cells treated with 100nM 4OHT for 12 days. The four columns on the right are associated with significantly upregulated genes, while the four columns furthest left described significantly downregulated genes. The first column for the up- and downregulated genes shows the Probe ID associated with each gene from the Affymetric Human Gene 1.0 ST microarrays. The gene name are shown in the next column, followed by the log2(fold change) in expression for each gene after 4OHT treatment in the H358-TwistER cells. The final column shows the p-value for each gene adjusted for the false discovery rate. Figure S1. Akt inhibition reduces serum-independent proliferation. (A) Proliferation of H358- TwistER cells with increasing concentrations (uM) of GSK-690693 (top) or MK-2206 (bottom). (B) Lysates from H358-TwistER cells treated for 48hrs in serum free media with indicated concentrations of GSK-690693 (top), or MK-2206 (bottom). Figure S2. The effects of TGFβ1 are specific to epithelial cells and lead to reduced serum- independent proliferation. (A) Proliferation of untreated (top) or TGFβ1 pre-treated (bottom) H358 and H441 cells in the presence (10%) or absence (SS) of serum. -
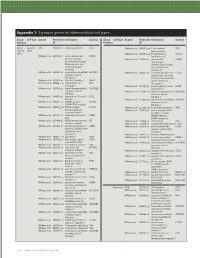
Technical Note, Appendix: an Analysis of Blood Processing Methods to Prepare Samples for Genechip® Expression Profiling (Pdf, 1
Appendix 1: Signature genes for different blood cell types. Blood Cell Type Source Probe Set Description Symbol Blood Cell Type Source Probe Set Description Symbol Fraction ID Fraction ID Mono- Lympho- GSK 203547_at CD4 antigen (p55) CD4 Whitney et al. 209813_x_at T cell receptor TRG nuclear cytes gamma locus cells Whitney et al. 209995_s_at T-cell leukemia/ TCL1A Whitney et al. 203104_at colony stimulating CSF1R lymphoma 1A factor 1 receptor, Whitney et al. 210164_at granzyme B GZMB formerly McDonough (granzyme 2, feline sarcoma viral cytotoxic T-lymphocyte- (v-fms) oncogene associated serine homolog esterase 1) Whitney et al. 203290_at major histocompatibility HLA-DQA1 Whitney et al. 210321_at similar to granzyme B CTLA1 complex, class II, (granzyme 2, cytotoxic DQ alpha 1 T-lymphocyte-associated Whitney et al. 203413_at NEL-like 2 (chicken) NELL2 serine esterase 1) Whitney et al. 203828_s_at natural killer cell NK4 (H. sapiens) transcript 4 Whitney et al. 212827_at immunoglobulin heavy IGHM Whitney et al. 203932_at major histocompatibility HLA-DMB constant mu complex, class II, Whitney et al. 212998_x_at major histocompatibility HLA-DQB1 DM beta complex, class II, Whitney et al. 204655_at chemokine (C-C motif) CCL5 DQ beta 1 ligand 5 Whitney et al. 212999_x_at major histocompatibility HLA-DQB Whitney et al. 204661_at CDW52 antigen CDW52 complex, class II, (CAMPATH-1 antigen) DQ beta 1 Whitney et al. 205049_s_at CD79A antigen CD79A Whitney et al. 213193_x_at T cell receptor beta locus TRB (immunoglobulin- Whitney et al. 213425_at Homo sapiens cDNA associated alpha) FLJ11441 fis, clone Whitney et al. 205291_at interleukin 2 receptor, IL2RB HEMBA1001323, beta mRNA sequence Whitney et al. -

The Expanded Endocannabinoid System/Endocannabinoidome As a Potential Target for Treating Diabetes Mellitus
Current Diabetes Reports (2019) 19:117 https://doi.org/10.1007/s11892-019-1248-9 OBESITY (KM GADDE, SECTION EDITOR) The Expanded Endocannabinoid System/Endocannabinoidome as a Potential Target for Treating Diabetes Mellitus Alain Veilleux1,2,3 & Vincenzo Di Marzo1,2,3,4,5 & Cristoforo Silvestri3,4,5 # Springer Science+Business Media, LLC, part of Springer Nature 2019 Abstract Purpose of Review The endocannabinoid (eCB) system, i.e. the receptors that respond to the psychoactive component of cannabis, their endogenous ligands and the ligand metabolic enzymes, is part of a larger family of lipid signals termed the endocannabinoidome (eCBome). We summarize recent discoveries of the roles that the eCBome plays within peripheral tissues in diabetes, and how it is being targeted, in an effort to develop novel therapeutics for the treatment of this increasingly prevalent disease. Recent Findings As with the eCB system, many eCBome members regulate several physiological processes, including energy intake and storage, glucose and lipid metabolism and pancreatic health, which contribute to the development of type 2 diabetes (T2D). Preclinical studies increasingly support the notion that targeting the eCBome may beneficially affect T2D. Summary The eCBome is implicated in T2D at several levels and in a variety of tissues, making this complex lipid signaling system a potential source of many potential therapeutics for the treatments for T2D. Keywords Endocannabinoidome . Bioactive lipids . Peripheral tissues . Glucose . Insulin Introduction: The Endocannabinoid System cannabis-derived natural product, Δ9-tetrahydrocannabinol and its Subsequent Expansion (THC), responsible for most of the psychotropic, euphoric to the “Endocannabinoidome” and appetite-stimulating actions (via CB1 receptors) and immune-modulatory effects (via CB2 receptors) of marijuana, The discovery of two G protein-coupled receptors, the canna- opened the way to the identification of the endocannabinoids binoid receptor type-1 (CB1) and − 2 (CB2) [1, 2], for the (eCBs). -
![EMR3 (ADGRE3) Hamster Monoclonal Antibody [Clone ID: 3D7] Product Data](https://docslib.b-cdn.net/cover/2432/emr3-adgre3-hamster-monoclonal-antibody-clone-id-3d7-product-data-802432.webp)
EMR3 (ADGRE3) Hamster Monoclonal Antibody [Clone ID: 3D7] Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for AM01316FC-N EMR3 (ADGRE3) Hamster Monoclonal Antibody [Clone ID: 3D7] Product data: Product Type: Primary Antibodies Clone Name: 3D7 Applications: FC Recommended Dilution: Flow cytometry ( Neat - 1/10): use 10 µl of the suggested working dilution to label 10e6 cells in 100 µl. Reactivity: Human Host: Hamster Isotype: IgG Clonality: Monoclonal Immunogen: ARHO-EMR3-CD97 (EGF1) transfectants. Spleen cells from immunised Armenian hamster were fused with cells of the mouse SP2/0 myeloma cell line. Specificity: This antibody recognises epidermal growth factor (EGF) module-containing mucin-like hormone receptor 3 (EMR3), which is a 56kD member of the EGF-7 transmembrane (TM7) family of adhesion receptors. Formulation: PBS, pH 7.4, containing 0.09% Sodium Azide Label: FITC State: Liquid purified IgG Label: Fluorescein Isothiocyanate Isomer I Concentration: lot specific Purification: Affinity chromatography on Protein G Conjugation: FITC Storage: Store the antibody undiluted at 2-8°C for one month or (in aliquots) at -20°C for longer. Avoid repeated freezing and thawing. This product is photosensitive and should be protected from light. Stability: Shelf life: one year from despatch. Database Link: Entrez Gene 84658 Human Q9BY15 This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 EMR3 (ADGRE3) Hamster Monoclonal Antibody [Clone ID: 3D7] – AM01316FC-N Background: EMR3 is expressed at the cell surface as a heterodimer.