Gene Recquest) - Software to Assist in Identification and Selection of Candidate Genes from Genomic Regions
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Mouse Pign Conditional Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Pign Conditional Knockout Project (CRISPR/Cas9) Objective: To create a Pign conditional knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Pign gene (NCBI Reference Sequence: NM_013784 ; Ensembl: ENSMUSG00000056536 ) is located on Mouse chromosome 1. 31 exons are identified, with the ATG start codon in exon 4 and the TGA stop codon in exon 31 (Transcript: ENSMUST00000186485). Exon 8 will be selected as conditional knockout region (cKO region). Deletion of this region should result in the loss of function of the Mouse Pign gene. To engineer the targeting vector, homologous arms and cKO region will be generated by PCR using BAC clone RP23-213N12 as template. Cas9, gRNA and targeting vector will be co-injected into fertilized eggs for cKO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Mice homozygous for an ENU-induced allele exhibit abnormal gastrulation, forebrain hypoplasia, coloboma, and microphthalmia. Exon 8 starts from about 19.69% of the coding region. The knockout of Exon 8 will result in frameshift of the gene. The size of intron 7 for 5'-loxP site insertion: 3696 bp, and the size of intron 8 for 3'-loxP site insertion: 1043 bp. The size of effective cKO region: ~625 bp. The cKO region does not have any other known gene. Page 1 of 8 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele gRNA region 5' gRNA region 3' 1 8 9 31 Targeting vector Targeted allele Constitutive KO allele (After Cre recombination) Legends Exon of mouse Pign Homology arm cKO region loxP site Page 2 of 8 https://www.alphaknockout.com Overview of the Dot Plot Window size: 10 bp Forward Reverse Complement Sequence 12 Note: The sequence of homologous arms and cKO region is aligned with itself to determine if there are tandem repeats. -

Congenital Disorders of Glycosylation from a Neurological Perspective
brain sciences Review Congenital Disorders of Glycosylation from a Neurological Perspective Justyna Paprocka 1,* , Aleksandra Jezela-Stanek 2 , Anna Tylki-Szyma´nska 3 and Stephanie Grunewald 4 1 Department of Pediatric Neurology, Faculty of Medical Science in Katowice, Medical University of Silesia, 40-752 Katowice, Poland 2 Department of Genetics and Clinical Immunology, National Institute of Tuberculosis and Lung Diseases, 01-138 Warsaw, Poland; [email protected] 3 Department of Pediatrics, Nutrition and Metabolic Diseases, The Children’s Memorial Health Institute, W 04-730 Warsaw, Poland; [email protected] 4 NIHR Biomedical Research Center (BRC), Metabolic Unit, Great Ormond Street Hospital and Institute of Child Health, University College London, London SE1 9RT, UK; [email protected] * Correspondence: [email protected]; Tel.: +48-606-415-888 Abstract: Most plasma proteins, cell membrane proteins and other proteins are glycoproteins with sugar chains attached to the polypeptide-glycans. Glycosylation is the main element of the post- translational transformation of most human proteins. Since glycosylation processes are necessary for many different biological processes, patients present a diverse spectrum of phenotypes and severity of symptoms. The most frequently observed neurological symptoms in congenital disorders of glycosylation (CDG) are: epilepsy, intellectual disability, myopathies, neuropathies and stroke-like episodes. Epilepsy is seen in many CDG subtypes and particularly present in the case of mutations -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

A Genome-Wide Association Study of Calf Birth Weight in Holstein Cattle Using Single Nucleotide Polymorphisms and Phenotypes Predicted from Auxiliary Traits
J. Dairy Sci. 97 :3156–3172 http://dx.doi.org/ 10.3168/jds.2013-7409 © American Dairy Science Association®, 2014 . A genome-wide association study of calf birth weight in Holstein cattle using single nucleotide polymorphisms and phenotypes predicted from auxiliary traits J. B. Cole ,*1 B. Waurich ,† M. Wensch-Dorendorf ,† D. M. Bickhart ,* and H. H. Swalve † * Animal Improvement Programs Laboratory, Agricultural Research Service, USDA, Beltsville, MD 20705-2350 † Institute of Agricultural and Nutritional Sciences, Martin-Luther-University Halle-Wittenberg, Theodor-Lieser-Str. 11, D-06120 Halle / Saale, Germany ABSTRACT derived from one population are useful for identifying genes and gene networks associated with phenotypes Previous research has found that a quantitative trait that are not directly measured in a second population. locus exists affecting calving and conformation traits This approach will identify only genes associated with on Bos taurus autosome 18 that may be related to the traits used to construct the birth weight predictor, increased calf birth weights, which are not routinely and not loci that affect only birth weight. recorded in the United States. Birth weight data from Key words: birth weight , quantitative trait loci , se- large, intensively managed dairies in eastern Germany lection index , single nucleotide polymorphism with management systems similar to those commonly found in the United States were used to develop a selec- INTRODUCTION tion index predictor for predicted transmitting ability (PTA) of birth weight. The predictor included body Many studies have reported on QTL affecting calving depth, rump width, sire calving ease, sire gestation traits in several populations of Holstein cattle (Kühn et al., 2003; Schnabel et al., 2005; Holmberg and Ander- length, sire stillbirth, stature, and strength. -

Hypotonia and Intellectual Disability Without Dysmorphic Features in A
Thiffault et al. BMC Medical Genetics (2017) 18:124 DOI 10.1186/s12881-017-0481-9 CASE REPORT Open Access Hypotonia and intellectual disability without dysmorphic features in a patient with PIGN-related disease Isabelle Thiffault1,2,3* , Britton Zuccarelli4, Holly Welsh4, Xuan Yuan5, Emily Farrow1, Lee Zellmer1, Neil Miller1, Sarah Soden1,3,4, Ahmed Abdelmoity4, Robert A. Brodsky5 and Carol Saunders1,2,3 Abstract Background: Defects in the human glycosylphosphatidylinositol anchor biosynthetic pathway are associated with inherited glycosylphosphatidylinositol (GPI)-deficiencies characterized by a broad range of clinical phenotypes including multiple congenital anomalies, dysmorphic faces, developmental delay, hypotonia, and epilepsy. Biallelic variants in PIGN, encoding phosphatidylinositol-glycan biosynthesis class N have been recently associated with multiple congenital anomalies hypotonia seizure syndrome. Case presentation: Our patient is a 2 year old male with hypotonia, global developmental delay, and focal epilepsy. Trio whole-exome sequencing revealed heterozygous variants in PIGN, c.181G > T (p.Glu61*) and c.284G > A (p.Arg95Gln). Analysis of FLAER and anti-CD59 by flow-cytometry demonstrated a shift in this patient’s granulocytes, confirming a glycosylphosphatidylinositol-biosynthesis defect, consistent with PIGN-related disease. Conclusions: To date, a total of 18 patients have been reported, all but 2 of whom have congenital anomalies and/or obvious dysmorphic features. Our patient has no significant dysmorphic features or multiple congenital anomalies, which is consistent with recent reports linking non-truncating variants with a milder phenotype, highlighting the importance of functional studies in interpreting sequence variants. Keywords: PIGN, Developmental disorders, Intellectual disability, GPI deficiency, Seizures Background GPI proteins with important roles in embryogenesis require Biallelic variants in glycosylphosphatidylinositol (GPI)-an- GPI anchoring for expression on the cell surface [3]. -

Genome-Wide Association and Gene Enrichment Analyses of Meat Sensory Traits in a Crossbred Brahman-Angus
Proceedings of the World Congress on Genetics Applied to Livestock Production, 11. 124 Genome-wide association and gene enrichment analyses of meat tenderness in an Angus-Brahman cattle population J.D. Leal-Gutíerrez1, M.A. Elzo1, D. Johnson1 & R.G. Mateescu1 1 University of Florida, Department of Animal Sciences, 2250 Shealy Dr, 32608 Gainesville, Florida, United States. [email protected] Summary The objective of this study was to identify genomic regions associated with meat tenderness related traits using a whole-genome scan approach followed by a gene enrichment analysis. Warner-Bratzler shear force (WBSF) was measured on 673 steaks, and tenderness and connective tissue were assessed by a sensory panel on 496 steaks. Animals belong to the multibreed Angus-Brahman herd from University of Florida and range from 100% Angus to 100% Brahman. All animals were genotyped with the Bovine GGP F250 array. Gene enrichment was identified in two pathways; the first pathway is involved in negative regulation of transcription from RNA polymerase II, and the second pathway groups several cellular component of the endoplasmic reticulum membrane. Keywords: tenderness, gene enrichment, regulation of transcription, cell growth, cell proliferation Introduction Identification of quantitative trait loci (QTL) for any complex trait, including meat tenderness, is the first most important step in the process of understanding the genetic architecture underlying the phenotype. Given a large enough population and a dense coverage of the genome, a genome-wide association study (GWAS) is usually successful in uncovering major genes and QTLs with large and medium effect on these type of traits. Several GWA studies on Bos indicus (Magalhães et al., 2016; Tizioto et al., 2013) or crossbred beef cattle breeds (Bolormaa et al., 2011b; Hulsman Hanna et al., 2014; Lu et al., 2013) were successful at identifying QTL for meat tenderness; and most of them include the traditional candidate genes µ-calpain and calpastatin. -

Whole-Exome Sequencing Identifies Causative Mutations in Families
BASIC RESEARCH www.jasn.org Whole-Exome Sequencing Identifies Causative Mutations in Families with Congenital Anomalies of the Kidney and Urinary Tract Amelie T. van der Ven,1 Dervla M. Connaughton,1 Hadas Ityel,1 Nina Mann,1 Makiko Nakayama,1 Jing Chen,1 Asaf Vivante,1 Daw-yang Hwang,1 Julian Schulz,1 Daniela A. Braun,1 Johanna Magdalena Schmidt,1 David Schapiro,1 Ronen Schneider,1 Jillian K. Warejko,1 Ankana Daga,1 Amar J. Majmundar,1 Weizhen Tan,1 Tilman Jobst-Schwan,1 Tobias Hermle,1 Eugen Widmeier,1 Shazia Ashraf,1 Ali Amar,1 Charlotte A. Hoogstraaten,1 Hannah Hugo,1 Thomas M. Kitzler,1 Franziska Kause,1 Caroline M. Kolvenbach,1 Rufeng Dai,1 Leslie Spaneas,1 Kassaundra Amann,1 Deborah R. Stein,1 Michelle A. Baum,1 Michael J.G. Somers,1 Nancy M. Rodig,1 Michael A. Ferguson,1 Avram Z. Traum,1 Ghaleb H. Daouk,1 Radovan Bogdanovic,2 Natasa Stajic,2 Neveen A. Soliman,3,4 Jameela A. Kari,5,6 Sherif El Desoky,5,6 Hanan M. Fathy,7 Danko Milosevic,8 Muna Al-Saffar,1,9 Hazem S. Awad,10 Loai A. Eid,10 Aravind Selvin,11 Prabha Senguttuvan,12 Simone Sanna-Cherchi,13 Heidi L. Rehm,14 Daniel G. MacArthur,14,15 Monkol Lek,14,15 Kristen M. Laricchia,15 Michael W. Wilson,15 Shrikant M. Mane,16 Richard P. Lifton,16,17 Richard S. Lee,18 Stuart B. Bauer,18 Weining Lu,19 Heiko M. Reutter ,20,21 Velibor Tasic,22 Shirlee Shril,1 and Friedhelm Hildebrandt1 Due to the number of contributing authors, the affiliations are listed at the end of this article. -

A Case Report of Severe Childhood Obesity and Tall Stature
Hindawi Publishing Corporation Case Reports in Pediatrics Volume 2016, Article ID 6123150, 6 pages http://dx.doi.org/10.1155/2016/6123150 Case Report Melanocortin-4 Receptor Deficiency Phenotype with an Interstitial 18q Deletion: A Case Report of Severe Childhood Obesity and Tall Stature Sarah Abdullah,1 William Reginold,2 Courtney Kiss,3 Karen J. Harrison,4,5 and Jennifer J. MacKenzie1,3 1 School of Medicine, Queen’s University, Kingston, ON, Canada 2Department of Medical Imaging, University of Toronto, Toronto, ON, Canada 3Department of Pediatrics, Kingston General Hospital, Kingston, ON, Canada 4Department of Pathology and Laboratory Medicine, IWK Health Centre, Halifax, NS, Canada 5Department of Pathology, Dalhousie University, Halifax, NS, Canada Correspondence should be addressed to Jennifer J. MacKenzie; [email protected] Received 30 June 2016; Accepted 28 August 2016 Academic Editor: Nur Arslan Copyright © 2016 Sarah Abdullah et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Childhood obesity is a growing health concern, associated with significant physical and psychological morbidity. Childhood obesity is known to have a strong genetic component, with mutations in the melanocortin-4 receptor (MC4R) gene being the most common monogenetic cause of obesity. Over 166 different MC4R mutations have been identified in persons with hyperphagia, severe childhood obesity, and increased linear growth. However, it is unclear whether the MC4-R deficiency phenotype is due to haploinsufficiency or dominant-negative effects by the mutant receptor. We report the case of a four-and-a-half-year-old boy with an interstitial deletion involving the long arm of chromosome 18 (46,XY,del(18)(q21.32q22.1)) encompassing the MC4R gene. -
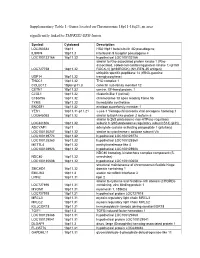
Supplementary Table 1: Genes Located on Chromosome 18P11-18Q23, an Area Significantly Linked to TMPRSS2-ERG Fusion
Supplementary Table 1: Genes located on Chromosome 18p11-18q23, an area significantly linked to TMPRSS2-ERG fusion Symbol Cytoband Description LOC260334 18p11 HSA18p11 beta-tubulin 4Q pseudogene IL9RP4 18p11.3 interleukin 9 receptor pseudogene 4 LOC100132166 18p11.32 hypothetical LOC100132166 similar to Rho-associated protein kinase 1 (Rho- associated, coiled-coil-containing protein kinase 1) (p160 LOC727758 18p11.32 ROCK-1) (p160ROCK) (NY-REN-35 antigen) ubiquitin specific peptidase 14 (tRNA-guanine USP14 18p11.32 transglycosylase) THOC1 18p11.32 THO complex 1 COLEC12 18pter-p11.3 collectin sub-family member 12 CETN1 18p11.32 centrin, EF-hand protein, 1 CLUL1 18p11.32 clusterin-like 1 (retinal) C18orf56 18p11.32 chromosome 18 open reading frame 56 TYMS 18p11.32 thymidylate synthetase ENOSF1 18p11.32 enolase superfamily member 1 YES1 18p11.31-p11.21 v-yes-1 Yamaguchi sarcoma viral oncogene homolog 1 LOC645053 18p11.32 similar to BolA-like protein 2 isoform a similar to 26S proteasome non-ATPase regulatory LOC441806 18p11.32 subunit 8 (26S proteasome regulatory subunit S14) (p31) ADCYAP1 18p11 adenylate cyclase activating polypeptide 1 (pituitary) LOC100130247 18p11.32 similar to cytochrome c oxidase subunit VIc LOC100129774 18p11.32 hypothetical LOC100129774 LOC100128360 18p11.32 hypothetical LOC100128360 METTL4 18p11.32 methyltransferase like 4 LOC100128926 18p11.32 hypothetical LOC100128926 NDC80 homolog, kinetochore complex component (S. NDC80 18p11.32 cerevisiae) LOC100130608 18p11.32 hypothetical LOC100130608 structural maintenance -

A Homozygous PIGN Missense Mutation in Soft-Coated Wheaten Terriers with a Canine Paroxysmal Dyskinesia
Neurogenetics DOI 10.1007/s10048-016-0502-4 ORIGINAL ARTICLE AhomozygousPIGN missense mutation in Soft-Coated Wheaten Terriers with a canine paroxysmal dyskinesia Ana L. Kolicheski1 & Gary S. Johnson1 & Tendai Mhlanga-Mutangadura1 & Jeremy F. Taylor2 & Robert D. Schnabel2,3 & Taroh Kinoshita 4 & Yoshiko Murakami 4 & Dennis P. O’Brien5 Received: 28 March 2016 /Accepted: 13 November 2016 # The Author(s) 2016. This article is published with open access at Springerlink.com Abstract Hereditary paroxysmal dyskinesias (PxD) are a het- PIGN:c.398C > T transition, which predicted the substitution of erogeneous group of movement disorders classified by frequen- an isoleucine for a highly conserved threonine in the encoded cy, duration, and triggers of the episodes. A young-adult onset enzyme. All 25 PxD-affected dogs were PIGN:c.398T allele canine PxD has segregated as an autosomal recessive trait in homozygotes, whereas there were no c.398T homozygotes Soft-Coated Wheaten Terriers. The medical records and videos among 1185 genotyped dogs without known histories of PxD. of episodes from 25 affected dogs were reviewed. The episodes PIGN encodes an enzyme involved in the biosynthesis of of hyperkinesia and dystonia lasted from several minutes to glycosylphosphatidylinositol (GPI), which anchors a variety several hours and could occur as often as >10/day. They were of proteins including CD59 to the cell surface. Flow cytometry not associated with strenuous exercise or fasting but were some- of PIGN-knockout HEK239 cells expressing recombinant hu- times triggered by excitement. The canine PxD phenotype most man PIGN with the c.398T variant showed reduced CD59 ex- closely resembled paroxysmal non-kinesigenic dyskinesia pression. -

Supplementary Table 1 Double Treatment Vs Single Treatment
Supplementary table 1 Double treatment vs single treatment Probe ID Symbol Gene name P value Fold change TC0500007292.hg.1 NIM1K NIM1 serine/threonine protein kinase 1.05E-04 5.02 HTA2-neg-47424007_st NA NA 3.44E-03 4.11 HTA2-pos-3475282_st NA NA 3.30E-03 3.24 TC0X00007013.hg.1 MPC1L mitochondrial pyruvate carrier 1-like 5.22E-03 3.21 TC0200010447.hg.1 CASP8 caspase 8, apoptosis-related cysteine peptidase 3.54E-03 2.46 TC0400008390.hg.1 LRIT3 leucine-rich repeat, immunoglobulin-like and transmembrane domains 3 1.86E-03 2.41 TC1700011905.hg.1 DNAH17 dynein, axonemal, heavy chain 17 1.81E-04 2.40 TC0600012064.hg.1 GCM1 glial cells missing homolog 1 (Drosophila) 2.81E-03 2.39 TC0100015789.hg.1 POGZ Transcript Identified by AceView, Entrez Gene ID(s) 23126 3.64E-04 2.38 TC1300010039.hg.1 NEK5 NIMA-related kinase 5 3.39E-03 2.36 TC0900008222.hg.1 STX17 syntaxin 17 1.08E-03 2.29 TC1700012355.hg.1 KRBA2 KRAB-A domain containing 2 5.98E-03 2.28 HTA2-neg-47424044_st NA NA 5.94E-03 2.24 HTA2-neg-47424360_st NA NA 2.12E-03 2.22 TC0800010802.hg.1 C8orf89 chromosome 8 open reading frame 89 6.51E-04 2.20 TC1500010745.hg.1 POLR2M polymerase (RNA) II (DNA directed) polypeptide M 5.19E-03 2.20 TC1500007409.hg.1 GCNT3 glucosaminyl (N-acetyl) transferase 3, mucin type 6.48E-03 2.17 TC2200007132.hg.1 RFPL3 ret finger protein-like 3 5.91E-05 2.17 HTA2-neg-47424024_st NA NA 2.45E-03 2.16 TC0200010474.hg.1 KIAA2012 KIAA2012 5.20E-03 2.16 TC1100007216.hg.1 PRRG4 proline rich Gla (G-carboxyglutamic acid) 4 (transmembrane) 7.43E-03 2.15 TC0400012977.hg.1 SH3D19 -

Supplementary Figure 1. Dystrophic Mice Show Unbalanced Stem Cell Niche
Supplementary Figure 1. Dystrophic mice show unbalanced stem cell niche. (A) Single channel images for the merged panels shown in Figure 1A, for of PAX7, MYOD and Laminin immunohistochemical staining in Lmna Δ8-11 mice of PAX7 and MYOD markers at the indicated days of post-natal growth. Basement membrane of muscle fibers was stained with Laminin. Scale bars, 50 µm. (B) Quantification of the % of PAX7+ MuSCs per 100 fibers at the indicated days of post-natal growth in (A). n =3-6 animals per genotype. (C) Immunohistochemical staining in Lmna Δ8-11 mice of activated, ASCs (PAX7+/KI67+) and quiescent QSCs (PAX7+/Ki67-) MuSCs at d19 and relative quantification (below). n= 4-6 animals per genotype. Scale bars, 50 µm. (D) Quantification of the number of cells per cluster in single myofibers extracted from d19 Lmna Δ8-11 mice and cultured 96h. n= 4-5 animals per group. Data are box with median and whiskers min to max. B, C, Data are mean ± s.e.m. Statistics by one-way (B) or two-way (C, D) analysis of variance (ANOVA) with multiple comparisons. * * P < 0.01, * * * P < 0.001. wt= Lmna Δ8-11 +/+; het= Lmna Δ8-11 +/; hom= Lmna Δ8-11 -/-. Supplementary Figure 2. Heterozygous mice show intermediate Lamin A levels. (A) RNA-seq signal tracks as the effective genome size normalized coverage of each biological replicate of Lmna Δ8-11 mice on Lmna locus. Neomycine cassette is indicated as a dark blue rectangle. (B) Western blot of total protein extracted from the whole Lmna Δ8-11 muscles at d19 hybridized with indicated antibodies.