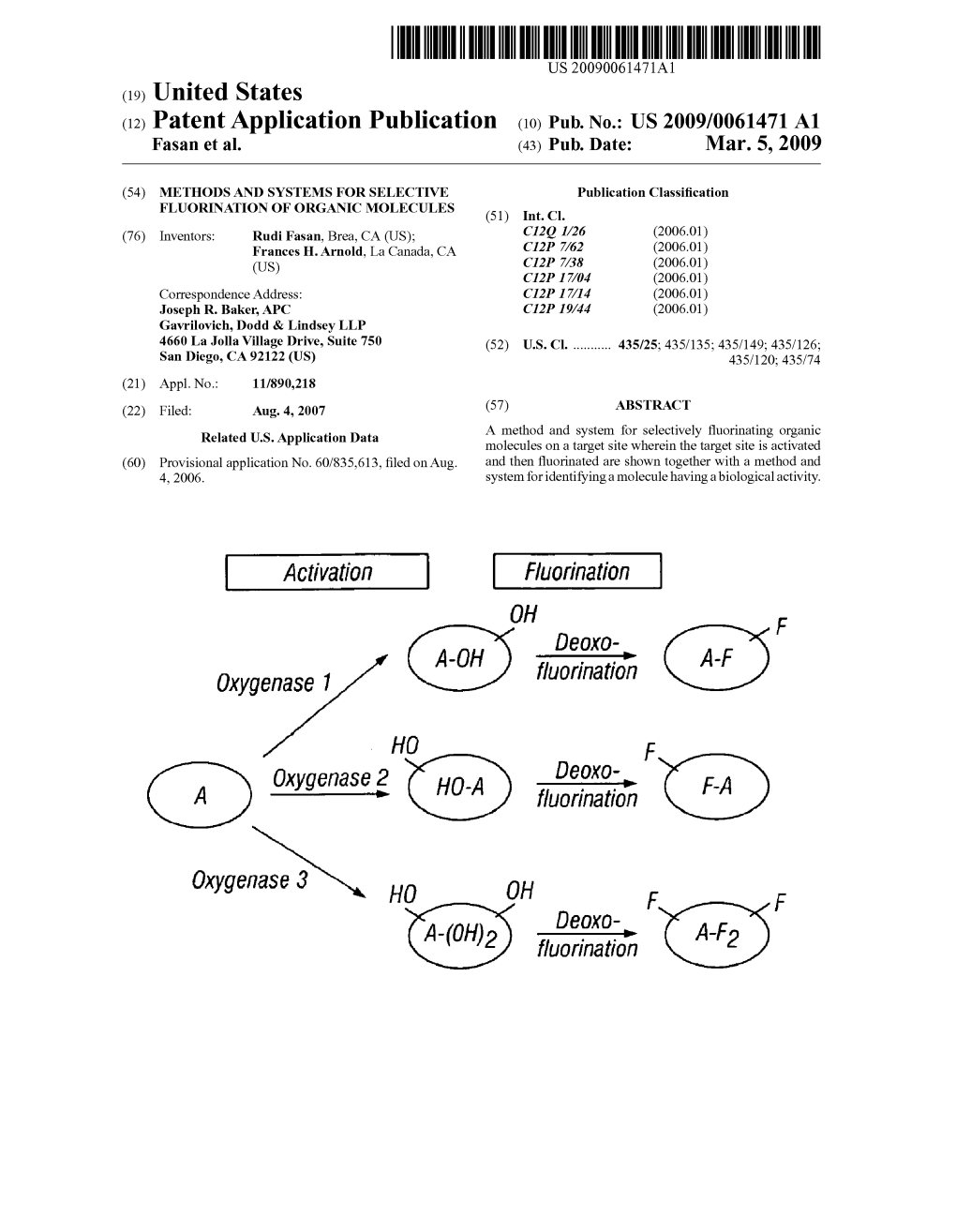
(E) Fluorination
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

(12) United States Patent (10) Patent No.: US 8,323,956 B2 Reardon Et Al
USOO8323956B2 (12) United States Patent (10) Patent No.: US 8,323,956 B2 Reardon et al. (45) Date of Patent: Dec. 4, 2012 (54) DISTAL TIP OF BIOSENSORTRANSDUCER 6,159,681 A 12/2000 Zebala COMPRISINGENZYME FOR DEAMINATION 6,265,201 B1 7/2001 Wackett et al. 6,271,015 B1* 8/2001 Gilula et al. .................. 435,228 6,284.522 B1 9, 2001 Wackett et al. (75) Inventors: Kenneth F. Reardon, Fort Collins, CO 6,291,200 B1 9, 2001 LeJeune et al. (US); Lawrence Philip Wackett, St. 6,369,299 B1 4/2002 Sadowsky et al. 6.825,001 B2 11/2004 Wackett et al. Paul, MN (US) 7,381,538 B2 * 6/2008 Reardon et al. ................. 435/23 7.595,181 B2 * 9/2009 Gruning et al. ............... 435/197 (73) Assignee: Colorado State University Research 7,709,221 B2 5, 2010 Rose et al. Foundation, Fort Collins, CO (US) 2004/0265811 A1 12/2004 Reardon et al. 2006/0275855 A1 12/2006 Blackburn et al. .............. 435/15 (*) Notice: Subject to any disclaimer, the term of this FOREIGN PATENT DOCUMENTS patent is extended or adjusted under 35 U.S.C. 154(b) by 340 days. WO WOO3,O25892 12/1993 OTHER PUBLICATIONS (21) Appl. No.: 12/358,140 PCT/US2009/040121, International Search Report & Written Opin (22) Filed: Jan. 22, 2009 ion mailed Jul. 14, 2009, 8 pages. Adachi, K., et al; Purification and properties of homogentisate (65) Prior Publication Data oxygenase from Pseudomonas fluorescens. Biochim. Biophys. Acta 118 (1966) 88-97. US 2009/0221014 A1 Sep. 3, 2009 Aldridge, W.N.; Serum esterases. -

Potential for and Distribution of Enzymatic Biodegradation of Polystyrene by Environmental Microorganisms
materials Communication Potential for and Distribution of Enzymatic Biodegradation of Polystyrene by Environmental Microorganisms Liyuan Hou and Erica L.-W. Majumder * Department of Chemistry, SUNY College of Environmental Science and Forestry, Syracuse, NY 13210, USA; [email protected] * Correspondence: [email protected] or [email protected]; Tel.: +1-3154706854 Abstract: Polystyrene (PS) is one of the main polymer types of plastic wastes and is known to be resistant to biodegradation, resulting in PS waste persistence in the environment. Although previous studies have reported that some microorganisms can degrade PS, enzymes and mechanisms of microorganism PS biodegradation are still unknown. In this study, we summarized microbial species that have been identified to degrade PS. By screening the available genome information of microorganisms that have been reported to degrade PS for enzymes with functional potential to depolymerize PS, we predicted target PS-degrading enzymes. We found that cytochrome P4500s, alkane hydroxylases and monooxygenases ranked as the top potential enzyme classes that can degrade PS since they can break C–C bonds. Ring-hydroxylating dioxygenases may be able to break the side-chain of PS and oxidize the aromatic ring compounds generated from the decomposition of PS. These target enzymes were distributed in Proteobacteria, Actinobacteria, Bacteroidetes, and Firmicutes, suggesting a broad potential for PS biodegradation in various earth environments and microbiomes. Our results provide insight into the enzymatic degradation of PS and suggestions for realizing the biodegradation of this recalcitrant plastic. Citation: Hou, L.; Majumder, E.L. Keywords: plastics; polystyrene biodegradation; enzymatic biodegradation; monooxygenase; alkane Potential for and Distribution of hydroxylase; cytochrome P450 Enzymatic Biodegradation of Polystyrene by Environmental Microorganisms. -

Thermophilic Bacteria Are Potential Sources of Novel Rieske Non-Heme
Chakraborty et al. AMB Expr (2017) 7:17 DOI 10.1186/s13568-016-0318-5 ORIGINAL ARTICLE Open Access Thermophilic bacteria are potential sources of novel Rieske non‑heme iron oxygenases Joydeep Chakraborty, Chiho Suzuki‑Minakuchi, Kazunori Okada and Hideaki Nojiri* Abstract Rieske non-heme iron oxygenases, which have a Rieske-type [2Fe–2S] cluster and a non-heme catalytic iron center, are an important family of oxidoreductases involved mainly in regio- and stereoselective transformation of a wide array of aromatic hydrocarbons. Though present in all domains of life, the most widely studied Rieske non-heme iron oxygenases are found in mesophilic bacteria. The present study explores the potential for isolating novel Rieske non- heme iron oxygenases from thermophilic sources. Browsing the entire bacterial genome database led to the identifi‑ cation of 45 homologs from thermophilic bacteria distributed mainly among Chloroflexi, Deinococcus–Thermus and Firmicutes. Thermostability, measured according to the aliphatic index, showed higher values for certain homologs compared with their mesophilic relatives. Prediction of substrate preferences indicated that a wide array of aromatic hydrocarbons could be transformed by most of the identified oxygenase homologs. Further identification of putative genes encoding components of a functional oxygenase system opens up the possibility of reconstituting functional thermophilic Rieske non-heme iron oxygenase systems with novel properties. Keywords: Rieske non-heme iron oxygenase, Oxidoreductase, Thermophiles, Aromatic hydrocarbons, Biotransformation Introduction of a wide array of agrochemically and pharmaceutically Rieske non-heme iron oxygenases (ROs) constitute a important compounds (Ensley et al. 1983; Wackett et al. large family of oxidoreductase enzymes involved primar- 1988; Hudlicky et al. -

Flavoprotein Hydroxylases and Epoxidases
277 Flavins on the Move: Flavoprotein Hydroxylases and Epoxidases Willem J.H. van Berkel1, Stefania Montersino1, Dirk Tischler2, Stefan Kaschabek2, Michael Schlömann2, George T. Gassner3 1Laboratory of Biochemistry, Wageningen University, Wageningen, The Netherlands 2Environmental Microbiology, TU Bergakademie Freiberg, Germany 3Department of Chemistry and Biochemistry, San Francisco State University, San Francisco, USA Introduction Flavoprotein monooxygenases perform chemo-, regio- and/or enantioselective oxygenations of organic substrates under mild reaction conditions [1]. These properties along with effective preparation methods turn flavoprotein monooxygenases in focus of industrial biocatalysis. Here we describe two biocatalytically relevant subclasses of flavoprotein monooxygenases with a close evolutionary relation: class A represented by p-hydroxybenzoate hydroxylase (PHBH) and class E formed by styrene monooxygenases (SMOs). PHBH family members perform highly regioselective hydroxylations on a wide variety of aromatic compounds. A rapid increase in available crystal structures and detailed mechanistic studies of such enzymes [2] are opening a new season of research in the field. SMOs catalyze a number of stereoselective epoxidation and sulfoxidation reactions [3]. Mechanistic and structural studies expose distinct characteristics, which provide a promising source for future biocatalyst development [4]. Nearly all bacterial SMOs are two- component proteins comprising a reductase and a monooxygenase. Remarkably, in few cases, the reductase is fused to the monooxygenase [5]. Such a self-sufficient enzyme can also cooperate with a single monooxygenase, resulting in a novel type of two-component SMO [6]. Results & Discussion Flavoprotein monooxygenases can be divided in six different subclasses based on structural features and oxygenation chemistry [1]. Table 1 gives an overview of the crystal structures of single-component flavoprotein aromatic hydroxylases (class A) and two-component styrene monooxygenases (SMO; class E). -

An Overview of Microbial Indigo-Forming Enzymes
Applied Microbiology and Biotechnology (2020) 104:925–933 https://doi.org/10.1007/s00253-019-10292-5 MINI-REVIEW An overview of microbial indigo-forming enzymes Andrea N. Fabara1 & Marco W. Fraaije1 Received: 11 October 2019 /Revised: 23 November 2019 /Accepted: 28 November 2019 /Published online: 13 December 2019 # The Author(s) 2019 Abstract Indigo is one of the oldest textile dyes and was originally prepared from plant material. Nowadays, indigo is chemically synthesized at a large scale to satisfy the demand for dyeing jeans. The current indigo production processes are based on fossil feedstocks; therefore, it is highly attractive to develop a more sustainable and environmentally friendly biotechnological process for the production of this popular dye. In the past decades, a number of natural and engineered enzymes have been identified that can be used for the synthesis of indigo. This mini-review provides an overview of the various microbial enzymes which are able to produce indigo and discusses the advantages and disadvantages of each biocatalytic system. Keywords Indigo . Indole . Naphthalene dioxygenase . Styrene monoxygenase . P450 monoxygenase . Peroxygenase . Flavoprotein monooxygenase Introduction vats, hence the classification as a vat-dye. The deposition of the insoluble indigo on the fabric results in a special decora- Humans have decorated textiles with dyes and pigments, often tion of the fabric material as the fabric is not penetrated by the derived from plant material, since ancient times (Aino et al. dye. This gives denim and other indigo-dyed garments their 2018). Indigo blue (in short: indigo) is one of the oldest dyes special appearance, where wear or abrasion exposes the white to be used for textile dyeing (Fig. -

Relating Metatranscriptomic Profiles to the Micropollutant
1 Relating Metatranscriptomic Profiles to the 2 Micropollutant Biotransformation Potential of 3 Complex Microbial Communities 4 5 Supporting Information 6 7 Stefan Achermann,1,2 Cresten B. Mansfeldt,1 Marcel Müller,1,3 David R. Johnson,1 Kathrin 8 Fenner*,1,2,4 9 1Eawag, Swiss Federal Institute of Aquatic Science and Technology, 8600 Dübendorf, 10 Switzerland. 2Institute of Biogeochemistry and Pollutant Dynamics, ETH Zürich, 8092 11 Zürich, Switzerland. 3Institute of Atmospheric and Climate Science, ETH Zürich, 8092 12 Zürich, Switzerland. 4Department of Chemistry, University of Zürich, 8057 Zürich, 13 Switzerland. 14 *Corresponding author (email: [email protected] ) 15 S.A and C.B.M contributed equally to this work. 16 17 18 19 20 21 This supporting information (SI) is organized in 4 sections (S1-S4) with a total of 10 pages and 22 comprises 7 figures (Figure S1-S7) and 4 tables (Table S1-S4). 23 24 25 S1 26 S1 Data normalization 27 28 29 30 Figure S1. Relative fractions of gene transcripts originating from eukaryotes and bacteria. 31 32 33 Table S1. Relative standard deviation (RSD) for commonly used reference genes across all 34 samples (n=12). EC number mean fraction bacteria (%) RSD (%) RSD bacteria (%) RSD eukaryotes (%) 2.7.7.6 (RNAP) 80 16 6 nda 5.99.1.2 (DNA topoisomerase) 90 11 9 nda 5.99.1.3 (DNA gyrase) 92 16 10 nda 1.2.1.12 (GAPDH) 37 39 6 32 35 and indicates not determined. 36 37 38 39 S2 40 S2 Nitrile hydration 41 42 43 44 Figure S2: Pearson correlation coefficients r for rate constants of bromoxynil and acetamiprid with 45 gene transcripts of ECs describing nucleophilic reactions of water with nitriles. -

An Overview of Microbial Indigo-Forming Enzymes Fabara, Andrea N.; Fraaije, Marco W
University of Groningen An overview of microbial indigo-forming enzymes Fabara, Andrea N.; Fraaije, Marco W. Published in: Applied Microbiology and Biotechnology DOI: 10.1007/s00253-019-10292-5 IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please check the document version below. Document Version Publisher's PDF, also known as Version of record Publication date: 2020 Link to publication in University of Groningen/UMCG research database Citation for published version (APA): Fabara, A. N., & Fraaije, M. W. (2020). An overview of microbial indigo-forming enzymes. Applied Microbiology and Biotechnology, 104(3), 925-933. https://doi.org/10.1007/s00253-019-10292-5 Copyright Other than for strictly personal use, it is not permitted to download or to forward/distribute the text or part of it without the consent of the author(s) and/or copyright holder(s), unless the work is under an open content license (like Creative Commons). The publication may also be distributed here under the terms of Article 25fa of the Dutch Copyright Act, indicated by the “Taverne” license. More information can be found on the University of Groningen website: https://www.rug.nl/library/open-access/self-archiving-pure/taverne- amendment. Take-down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your claim. Downloaded from the University of Groningen/UMCG research database (Pure): http://www.rug.nl/research/portal. For technical reasons the number of authors shown on this cover page is limited to 10 maximum. -

Structure-Function Studies of Iron-Sulfur Enzyme Systems
Structure-Function Studies of Iron-Sulfur Enzyme Systems Rosmarie Friemann Department of Molecular Biology Uppsala Doctoral thesis Swedish University of Agricultural Sciences Uppsala 2005 1 Acta Universitatis Agriculturae Sueciae Agraria 504 ISSN 1401-6249 ISBN 91-576-6783-7 © 2004 Rosmarie Friemann, Uppsala Tryck: SLU Service/Repro, Uppsala 2004 2 Abstract Friemann, R., 2005, Structure-Function Studies of Iron-Sulfur Enzyme Systems. Doctorial dissertation. ISSN 1401-6249, ISBN 91-576-6783-7 Iron-sulfur clusters are among the most ancient of metallocofactors and serve a variety of biological functions in proteins, including electron transport, catalytic, and structural roles. Two kinds of multicomponent enzyme systems have been investigated by X-ray crystallography, the ferredoxin/thioredoxin system and bacterial Rieske non- heme iron dioxygenase (RDO) systems. The ferredoxin/thioredoxin system is a light sensitive system controlling the activities of key enzymes involved in the assimilatory (photosynthetic) and dissimilatory pathways in chloroplasts and photosynthetic bacteria. The system consists of a ferredoxin, ferredoxin:thioredoxin reductase (FTR), and two thioredoxins, Trx-m and Trx-f. In light, photosystem I reduces ferredoxin that reduces Trx-m and Trx- f. This two-electron reduction is catalyzed by FTR that contains a [4Fe-4S] center and a proximal disulfide bridge. When the first electron is delivered by the ferredoxin, an intermediate is formed where one thiol of the proximal disulfide attacks the disulfide bridge of thioredoxin. This results in a transient protein-protein complex held together by a mixed disulfide between FTR and Trx-m. This complex is stabilized by using a C40S mutant Trx-m and its structure have been determined. -

Structure-Based Redesign of a Self-Sufficient Flavin-Containing Monooxygenase Towards Indigo Production Lončar, Nikola; Van Beek, Hugo L.; Fraaije, Marco W
University of Groningen Structure-Based Redesign of a Self-Sufficient Flavin-Containing Monooxygenase towards Indigo Production Lončar, Nikola; van Beek, Hugo L.; Fraaije, Marco W. Published in: International Journal of Molecular Sciences DOI: 10.3390/ijms20246148 IMPORTANT NOTE: You are advised to consult the publisher's version (publisher's PDF) if you wish to cite from it. Please check the document version below. Document Version Publisher's PDF, also known as Version of record Publication date: 2019 Link to publication in University of Groningen/UMCG research database Citation for published version (APA): Lonar, N., van Beek, H. L., & Fraaije, M. W. (2019). Structure-Based Redesign of a Self-Sufficient Flavin- Containing Monooxygenase towards Indigo Production. International Journal of Molecular Sciences, 20(24). https://doi.org/10.3390/ijms20246148 Copyright Other than for strictly personal use, it is not permitted to download or to forward/distribute the text or part of it without the consent of the author(s) and/or copyright holder(s), unless the work is under an open content license (like Creative Commons). Take-down policy If you believe that this document breaches copyright please contact us providing details, and we will remove access to the work immediately and investigate your claim. Downloaded from the University of Groningen/UMCG research database (Pure): http://www.rug.nl/research/portal. For technical reasons the number of authors shown on this cover page is limited to 10 maximum. Download date: 13-01-2020 International Journal of Molecular Sciences Article Structure-Based Redesign of a Self-Sufficient Flavin-Containing Monooxygenase towards Indigo Production 1, 2, 2, Nikola Lonˇcar y, Hugo L. -

Biotechnological Potential of Rhodococcus Biodegradative Pathways Dockyu Kim1*, Ki Young Choi2, Miyoun Yoo3, Gerben J
J. Microbiol. Biotechnol. (2018), 28(7), 1037–1051 https://doi.org/10.4014/jmb.1712.12017 Research Article Review jmb Biotechnological Potential of Rhodococcus Biodegradative Pathways Dockyu Kim1*, Ki Young Choi2, Miyoun Yoo3, Gerben J. Zylstra4, and Eungbin Kim5 1Division of Polar Life Sciences, Korea Polar Research Institute, Incheon 21990, Republic of Korea 2University College, Yonsei University, Incheon 21983, Republic of Korea 3Korea Research Institute of Chemical Technology, Daejeon 34114, Republic of Korea 4Department of Biochemistry and Microbiology, School of Environmental and Biological Sciences, Rutgers University, NJ 08901-8520, USA 5Department of Systems Biology, Yonsei University, Seoul 03722, Republic of Korea Received: December 8, 2017 Revised: March 26, 2018 The genus Rhodococcus is a phylogenetically and catabolically diverse group that has been Accepted: May 1, 2018 isolated from diverse environments, including polar and alpine regions, for its versatile ability First published online to degrade a wide variety of natural and synthetic organic compounds. Their metabolic May 8, 2018 capacity and diversity result from their diverse catabolic genes, which are believed to be *Corresponding author obtained through frequent recombination events mediated by large catabolic plasmids. Many Phone: +82-32-760-5525; rhodococci have been used commercially for the biodegradation of environmental pollutants Fax: +82-32-760-5509; E-mail: [email protected] and for the biocatalytic production of high-value chemicals from low-value materials. -

Supplementary Information
Supplementary information (a) (b) Figure S1. Resistant (a) and sensitive (b) gene scores plotted against subsystems involved in cell regulation. The small circles represent the individual hits and the large circles represent the mean of each subsystem. Each individual score signifies the mean of 12 trials – three biological and four technical. The p-value was calculated as a two-tailed t-test and significance was determined using the Benjamini-Hochberg procedure; false discovery rate was selected to be 0.1. Plots constructed using Pathway Tools, Omics Dashboard. Figure S2. Connectivity map displaying the predicted functional associations between the silver-resistant gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S3. Connectivity map displaying the predicted functional associations between the silver-sensitive gene hits; disconnected gene hits not shown. The thicknesses of the lines indicate the degree of confidence prediction for the given interaction, based on fusion, co-occurrence, experimental and co-expression data. Figure produced using STRING (version 10.5) and a medium confidence score (approximate probability) of 0.4. Figure S4. Metabolic overview of the pathways in Escherichia coli. The pathways involved in silver-resistance are coloured according to respective normalized score. Each individual score represents the mean of 12 trials – three biological and four technical. Amino acid – upward pointing triangle, carbohydrate – square, proteins – diamond, purines – vertical ellipse, cofactor – downward pointing triangle, tRNA – tee, and other – circle. -

Monohydroxylation of Phenoland 2,5-Dichlorophenol by Toluene
APPLIED AND ENVIRONMENTAL MICROBIOLOGY, OCt. 1989, p. 2648-2652 Vol. 55, No. 10 0099-2240/89/102648-05$02.00/0 Copyright © 1989, American Society for Microbiology Monohydroxylation of Phenol and 2,5-Dichlorophenol by Toluene Dioxygenase in Pseudomonas putida Fl J. C. SPAIN,1* G. J. ZYLSTRA,2 C. K. BLAKE,2 AND D. T. GIBSON2 Air Force Engineering and Services Laboratory, Tyndall Air Force Base, Florida 32403,1 and Department of Microbiology, University of Iowa, Iowa City, Iowa 522422 Received 13 April 1989/Accepted 21 July 1989 Pseudomonas putida Fl contains a multicomponent enzyme system, toluene dioxygenase, that converts toluene and a variety of substituted benzenes to cis-dihydrodiols by the addition of one molecule of molecular oxygen. Toluene-grown cells of P. putida Fl also catalyze the monohydroxylation of phenols to the corresponding catechols by an unknown mechanism. Respirometric studies with washed cells revealed similar enzyme induction patterns in cells grown on toluene or phenol. Induction of toluene dioxygenase and subsequent enzymes for catechol oxidation allowed growth on phenol. Tests with specific mutants of P. putida Fl indicated that the ability to hydroxylate phenols was only expressed in cells that contained an active toluene dioxygenase enzyme system. 1802 experiments indicated that the overall reaction involved the incorporation of only one atom of oxygen in the catechol, which suggests either a monooxygenase mechanism or a dioxygenase reaction with subsequent specific elimination of water. The phenomenon of enzymatic oxygen fixation was first P. putida F39/D is a mutant strain of P. putida Fl that described in 1955 (11, 15).