Homospermidine, Spermidine, and Putrescine: The
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Histamine Degradation by Halophilic Archaea Wanaporn Tapingkae A
Histamine Degradation by Halophilic Archaea Wanaporn Tapingkae A Thesis Submitted in Partial Fulfillment of the Requirements for the Degree of Doctor of Philosophy in Food Technology Prince of Songkla University 2009 Copyright of Prince of Songkla University i Thesis Title Histamine Degradation by Halophilic Archaea Author Miss Wanaporn Tapingkae Major Program Food Technology Major Advisor Examining Committee : ……..…………………………………. ……..…………..…………….Chairperson (Dr. Wonnop Visessanguan) (Asst. Prof. Dr. Tipparat Hongpattarakere) ……..…………………..……………….… Co-advisors (Dr. Wonnop Visessanguan) ……..…………………………………. ……..…………………..……………….… (Prof. Dr. Soottawat Benjakul) (Prof. Dr. Soottawat Benjakul) ……..…………………………………. ……..…………………..……………….… (Prof. Dr. Kirk L. Parkin) (Assoc. Prof. Dr. Somboon Tanasupawat) ……..…………………..……………….… (Assoc. Prof. Dr. Borwonsak Leenanon) The Graduate School, Prince of Songkla University, has approved this thesis as partial fulfillment of the requirements for the Doctor of Philosophy Degree in Food Technology ………………………………………… (Assoc. Prof. Dr. Krerkchai Thongnoo) Dean of Graduate School ii ชือวทยานิ ิพนธ์ การยอยสลายฮ่ ีสตามีนโดยอาเคียที่ชอบเกลือ ผู้เขียน นางสาววรรณพร ทะพิงคแก์ สาขาวชาิ เทคโนโลยอาหารี ปี การศึกษา 2551 บทคดยั ่อ ปริมาณฮีสตามีนที่สูงก่อใหเก้ ิดผลเสียหายต่อทงคั ุณภาพและความปลอดภยของั นาปลาํ ดงนั นการทดลองนั ีจึงมีวตถั ุประสงคเพ์ ื่อคดแยกเชั ืออาเคียที่ชอบเกลือที่มีความสามารถยอย่ ฮีสตามีนในสภาวะที่มีเกลือสูง ศึกษาเอนไซมท์ ี่เกี่ยวของ้ และศึกษาความเป็นไปไดในการประย้ กตุ ์ ใชในกระบวนการผล้ ิตนาปลาํ จากเชืออาเคียที่ชอบเกลือ -

Supplementary Information
Supplementary Information Table S1. Categories of transcripts significantly regulated in salt-stressed Malus zumi. Number Expression a Putative Annotation Genome Genebank Identities p-value b locus Accession Signal transduction Kinase leucine-rich repeat transmembrane protein 5.41 × 10-5 1 I Chr 15 NP_199948 64% kinase 1 S leucine-rich repeat family protein kinase Chr 15 NP_179336 42% 1.02 × 10-4 1 I tousled-like serine/threonine kinase Chr 11 NP_568405 82% 3.20 × 10-5 1 I CIPK5 Chr 1 NP_568241 79% 7.74 × 10-5 1 S CIPK6 Chr 2 NP_194825 80% 1.98 × 10-7 1 S protein kinase family protein Chr 6 NP_194952 50% 2.25 × 10-5 Transcription factor 1 I IAA-LEUCINE RESISTANT3 Chr 3 NP_200279 89% 4.97 × 10-4 1 I IAA26 Chr 15 NP_188271 80% 3.58 × 10-6 1 S GT-like trihelix DNA-binding protein Chr 15 NP_177814 37% 3.04 × 10-6 1 S zinc finger (CCCH-type) family protein Chr 11 NP_200670 59% 8.13 × 10-5 1 I WRKY family transcription factor Chr 12 NP_001078015 50% 4.52 × 10-5 1 S GRAS family transcription factor Chr 10 XP_002322514 52% 3.58 × 10-4 2 S AP2 transcription factor Chr 15 NP_173355 70% 4.26 × 10-6 1 I SALT TOLERANCE homolog protein Chr 5 NP_849598 68% 5.37 × 10-7 1 S Auxin response factor Chr 7 NP_182176 70% 2.78 × 10-3 Int. J. Mol. Sci. 2013, 14 S2 Table S1. Cont. Number Expression a Putative Annotation Genome Genebank Identities p-value b locus Accession ROS elimination 1 S glutathione transferase Chr 3 NP_850479 62% 3.17 × 10-8 1 S peroxidase Chr 2 NP_201440 71% 4.26 × 10-3 1 S peroxidase Chr 10 NP_197022 54% 3.81 × 10-7 1 S peroxidase Chr 10 -

Symphytum Officinale) Hairy Roots by Rnai Silencing of Homospermidine Synthase
Published online: 2019-08-26 Original Papers Reduction of Pyrrolizidine Alkaloid Levels in Comfrey (Symphytum officinale) Hairy Roots by RNAi Silencing of Homospermidine Synthase Authors Lars H. Kruse 1, 3, Thomas Stegemann1, Julia Jensen-Kroll 1, Annika Engelhardt 1, Anne-Maria Wesseling 1, Annemarie Lippert 2, Jutta Ludwig-Müller 2,DietrichOber1 Affiliations ABSTRACT 1 Botanisches Institut, Kiel University, Kiel, Germany Comfrey is a medicinal plant, extracts of which are tradition- 2 Institut für Botanik, Technische Universität Dresden, ally used for the treatment of painful inflammatory muscle Dresden, Germany and joint problems, because the plant contains allantoin and 3 Plant Biology Section, School of Integrative Plant Science, rosmarinic acid. However, its medicinal use is limited because Cornell University, USA of its toxic pyrrolizidine alkaloid (PA) content. PAs encompass more than 400 different compounds that have been identified Key words from various plant lineages. To date, only the first pathway- Symphytum officinale, alkaloid extraction, transgenic tissue specific enzyme, homospermidine synthase (HSS), has been culture, transcript quantification, Boraginaceae characterized. HSS catalyzes the formation of homospermi- dine, which is exclusively incorporated into PAs. HSS has been received May 6, 2019 recruited several times independently in various plant line- revised August 5, 2019 ages during evolution by duplication of the gene encoding de- accepted August 15, 2019 oxyhypusine synthase (DHS), an enzyme of primary metabo- Bibliography lism. Here, we describe the establishment of RNAi knockdown DOI https://doi.org/10.1055/a-0998-5125 hairy root mutants of HSS in Symphytum officinale. A knock- – Published online August 26, 2019 | Planta Med 2019; 85: down of HSS by 60 80% resulted in a significant reduction of 1177–1186 © Georg Thieme Verlag KG Stuttgart · New York | homospermidine by ~ 86% and of the major PA components ‑ ‑ ISSN 0032‑0943 7 acetylintermedine N-oxide and 3 acetylmyoscorpine N-ox- ide by approximately 60%. -

Functional Characterization of Seven Γ-Glutamylpolyamine Synthetase
JOURNAL OF BACTERIOLOGY, Aug. 2011, p. 3923–3930 Vol. 193, No. 15 0021-9193/11/$12.00 doi:10.1128/JB.05105-11 Copyright © 2011, American Society for Microbiology. All Rights Reserved. Functional Characterization of Seven ␥-Glutamylpolyamine Synthetase Genes and the bauRABCD Locus for Polyamine and -Alanine Utilization in Pseudomonas aeruginosa PAO1ᰔ Xiangyu Yao,1† Weiqing He,1,2† and Chung-Dar Lu1,3* Department of Biology, Georgia State University, Atlanta, Georgia 303031; Institute of Medicinal Biotechnology, Chinese Academy of Medical Sciences and Peking Union Medical College, Beijing 100050, China2; and Department of Medical Laboratory Sciences and Biotechnology, China Medical University, Taichung, Taiwan 404023 Received 16 April 2011/Accepted 13 May 2011 Pseudomonas aeruginosa and many other bacteria can utilize biogenic polyamines, including diaminopropane (DAP), putrescine (Put), cadaverine (Cad), and spermidine (Spd), as carbon and/or nitrogen sources. Tran- scriptome analysis in response to exogenous Put and Spd led to the identification of a list of genes encoding putative enzymes for the catabolism of polyamines. Among them, pauA1 to pauA6, pauB1 to pauB4, pauC, and pauD1 and pauD2 (polyamine utilization) encode enzymes homologous to Escherichia coli PuuABCD of the ␥-glutamylation pathway in converting Put into GABA. A series of unmarked pauA mutants was constructed for growth phenotype analysis. The results revealed that it requires specific combinations of pauA knockouts to abolish utilization of different polyamines and support the importance of ␥-glutamylation for polyamine catabolism in P. aeruginosa. Another finding was that the list of Spd-inducible genes overlaps almost completely with that of Put-inducible ones except the pauA3B2 operon and the bauABCD operon (-alanine utilization). -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Pyrrolizidine Alkaloids: Biosynthesis, Biological Activities and Occurrence in Crop Plants
molecules Review Pyrrolizidine Alkaloids: Biosynthesis, Biological Activities and Occurrence in Crop Plants Sebastian Schramm, Nikolai Köhler and Wilfried Rozhon * Biotechnology of Horticultural Crops, TUM School of Life Sciences Weihenstephan, Technical University of Munich, Liesel-Beckmann-Straße 1, 85354 Freising, Germany; [email protected] (S.S.); [email protected] (N.K.) * Correspondence: [email protected]; Tel.: +49-8161-71-2023 Academic Editor: John C. D’Auria Received: 20 December 2018; Accepted: 29 January 2019; Published: 30 January 2019 Abstract: Pyrrolizidine alkaloids (PAs) are heterocyclic secondary metabolites with a typical pyrrolizidine motif predominantly produced by plants as defense chemicals against herbivores. They display a wide structural diversity and occur in a vast number of species with novel structures and occurrences continuously being discovered. These alkaloids exhibit strong hepatotoxic, genotoxic, cytotoxic, tumorigenic, and neurotoxic activities, and thereby pose a serious threat to the health of humans since they are known contaminants of foods including grain, milk, honey, and eggs, as well as plant derived pharmaceuticals and food supplements. Livestock and fodder can be affected due to PA-containing plants on pastures and fields. Despite their importance as toxic contaminants of agricultural products, there is limited knowledge about their biosynthesis. While the intermediates were well defined by feeding experiments, only one enzyme involved in PA biosynthesis has been characterized so far, the homospermidine synthase catalyzing the first committed step in PA biosynthesis. This review gives an overview about structural diversity of PAs, biosynthetic pathways of necine base, and necic acid formation and how PA accumulation is regulated. Furthermore, we discuss their role in plant ecology and their modes of toxicity towards humans and animals. -

REVIEW ARTICLES Pyrrolizidine Alkaloids
Review articles Pyrrolizidine alkaloids – chemistry, biosynthesis, pathway, toxicity, safety and perspectives of medicinal usage Mariola DREGER, MARZENA Stanisławska, ANNA Krajewska-Patan, SEBASTIAN Mielcarek, Przemysław Łukasz Mikołajczak, WALDEMAR Buchwald The Branch of Medicinal Plants Institute of Natural Fibres and Medicinal Plants Libelta 27 61-707 Poznań, Poland *corresponding author: [email protected] S u m m a r y Pyrrolizidine alkaloids (PAs) are the class of secondary metabolites that evolved as a powerful tool in the plant defensive interactions against herbivores. The occurrence of PAs in the plant world is scattered in several unrelated botanic families with special abun- dance in Asteraceae, Boraginaceae and Fabaceae. Homospermidine synthase (HSS) was recognized as a key enzyme that catalyzes homospermidine formation from polyamines. The studies of HSS kinetic and gene sequence revealed that it is of polyphyletic origin and raised as a result of deoxyhypusine synthase (DHS) gene duplication. The ability of PAs production occurred independently at least four times in course of plant evolution. The PAs biosynthesis is tightly correlated with growth phase and biomass production. It is sup- posed that PAs biosynthesis is individually regulated in different lineages of plants. The PAs with a 1,2 unsaturated necine skeleton show toxic activity (hepatoxicity, carcinogenicity, genotoxicity, teratogenocity and cytotoxicity). It is though that pyrrolic esters formation during the detoxication process in the liver is the main mechanism of PAs toxicity. The pyr- rolic esters are highly reactive and tend to bind rapidly with nucleophilic macromolecules including DNA and DNA-protein inducing hepatotoxicity or tumorigenecity. The problem of PAs toxicity cause the restrictions in the production and sale of herbal products. -

Spermidine Synthase (SPDS) Undergoes Concerted Structural Rearrangements Upon Ligand Binding – a Case Study of the Two SPDS Isoforms from Arabidopsis Thaliana
fpls-10-00555 May 4, 2019 Time: 16:20 # 1 ORIGINAL RESEARCH published: 07 May 2019 doi: 10.3389/fpls.2019.00555 Spermidine Synthase (SPDS) Undergoes Concerted Structural Rearrangements Upon Ligand Binding – A Case Study of the Two SPDS Isoforms From Arabidopsis thaliana Bartosz Sekula* and Zbigniew Dauter Synchrotron Radiation Research Section, Macromolecular Crystallography Laboratory, National Cancer Institute, Argonne, IL, United States Spermidine synthases (SPDSs) catalyze the production of the linear triamine, spermidine, from putrescine. They utilize decarboxylated S-adenosylmethionine (dc- SAM), a universal cofactor of aminopropyltransferases, as a donor of the aminopropyl Edited by: Antonio F. Tiburcio, moiety. In this work, we describe crystal structures of two SPDS isoforms from University of Barcelona, Spain Arabidopsis thaliana (AtSPDS1 and AtSPDS2). AtSPDS1 and AtSPDS2 are dimeric Reviewed by: enzymes that share the fold of the polyamine biosynthesis proteins. Subunits of both Taku Takahashi, Okayama University, Japan isoforms present the characteristic two-domain structure. Smaller, N-terminal domain Miguel A. Blazquez, is built of the two b-sheets, while the C-terminal domain has a Rossmann fold-like Spanish National Research Council topology. The catalytic cleft composed of two main compartments, the dc-SAM binding (CSIC), Spain site and the polyamine groove, is created independently in each AtSPDS subunits at *Correspondence: Bartosz Sekula the domain interface. We also provide the structural details about the dc-SAM binding [email protected]; mode and the inhibition of SPDS by a potent competitive inhibitor, cyclohexylamine [email protected] (CHA). CHA occupies the polyamine binding site of AtSPDS where it is bound at the Specialty section: bottom of the active site with the amine group placed analogously to the substrate. -

University of Florida Thesis Or Dissertation Formatting Template
METABOLOMIC CHARACTERIZATION OF THE INTESTINAL BACTERIAL OXALOBIOME By CASEY A. CHAMBERLAIN A DISSERTATION PRESENTED TO THE GRADUATE SCHOOL OF THE UNIVERSITY OF FLORIDA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY UNIVERSITY OF FLORIDA 2019 © 2019 Casey A. Chamberlain To my wife, Michelle ACKNOWLEDGMENTS Obtaining a PhD may only bestow the title of “Doctor” upon one individual, but the journey to such an achievement is often aided by, and impossible without, the loving support of family, friends, and others along the way. I am thankful for the Lord for His providence and love and for blessing me with the unique interests, talents, and gifts that gave me the opportunity to fulfill this achievement. I am eternally grateful for my wife, Michelle, for providing me with love, support, comfort, and guidance, which propagates confidence, happiness, and fulfillment into all areas of my life. Her guiding and strengthening example has molded me into the man I am today. I am blessed to have my daughter, Maci, who reminds me daily of the purpose of this life and the love the Lord feels for each of His children. I am fortunate to have received love and support from family, both immediate and extended, which helps foster a sense of wholeness in my life. I am uplifted and reinforced by the friends I have made during this season of life, both inside and outside the lab, particularly those whom I have come to know closely on a personal level. I respect, honor, and am indebted to Tim Garrett for providing me the opportunity for this journey. -

Supp Table 6.Pdf
Supplementary Table 6. Processes associated to the 2037 SCL candidate target genes ID Symbol Entrez Gene Name Process NM_178114 AMIGO2 adhesion molecule with Ig-like domain 2 adhesion NM_033474 ARVCF armadillo repeat gene deletes in velocardiofacial syndrome adhesion NM_027060 BTBD9 BTB (POZ) domain containing 9 adhesion NM_001039149 CD226 CD226 molecule adhesion NM_010581 CD47 CD47 molecule adhesion NM_023370 CDH23 cadherin-like 23 adhesion NM_207298 CERCAM cerebral endothelial cell adhesion molecule adhesion NM_021719 CLDN15 claudin 15 adhesion NM_009902 CLDN3 claudin 3 adhesion NM_008779 CNTN3 contactin 3 (plasmacytoma associated) adhesion NM_015734 COL5A1 collagen, type V, alpha 1 adhesion NM_007803 CTTN cortactin adhesion NM_009142 CX3CL1 chemokine (C-X3-C motif) ligand 1 adhesion NM_031174 DSCAM Down syndrome cell adhesion molecule adhesion NM_145158 EMILIN2 elastin microfibril interfacer 2 adhesion NM_001081286 FAT1 FAT tumor suppressor homolog 1 (Drosophila) adhesion NM_001080814 FAT3 FAT tumor suppressor homolog 3 (Drosophila) adhesion NM_153795 FERMT3 fermitin family homolog 3 (Drosophila) adhesion NM_010494 ICAM2 intercellular adhesion molecule 2 adhesion NM_023892 ICAM4 (includes EG:3386) intercellular adhesion molecule 4 (Landsteiner-Wiener blood group)adhesion NM_001001979 MEGF10 multiple EGF-like-domains 10 adhesion NM_172522 MEGF11 multiple EGF-like-domains 11 adhesion NM_010739 MUC13 mucin 13, cell surface associated adhesion NM_013610 NINJ1 ninjurin 1 adhesion NM_016718 NINJ2 ninjurin 2 adhesion NM_172932 NLGN3 neuroligin -

Functional Roles of Each of the 400 Putative Gene Products Used for Inferring Bradyrhizobium Phylogeny
Supplementary Table S3: Functional roles of each of the 400 putative gene products used for inferring Bradyrhizobium phylogeny. Kyoto Encyclopedia of Genes and Genomes (KEGG) Amino Gene Gene KEGG Acid number identifiera Orthology KEGG Annotation Functional class Definition model 1 BJ6T_00940 Uncharacterised Unclassified JTT NitT/TauT family transport system Mineral and organic ion transport 2 BJ6T_00960 K02050 permease protein system ABC.SN.P LG NitT/TauT family transport system ATP- Mineral and organic ion transport 3 BJ6T_00970 K02049 binding protein system ABC.SN.A JTT 4 BJ6T_01730 K05524 ferredoxin Energy metabolism fdxA Dayhoff 5 BJ6T_01770 K02902 large subunit ribosomal protein L28 Translation RP-L28 DCMut 6 BJ6T_01900 Uncharacterised Unclassified JTT Metabolism of cofactors and 7 BJ6T_01930 K09882 cobaltochelatase CobS vitamins cobS LG 8 BJ6T_01980 Uncharacterised Unclassified DCMut 9 BJ6T_02010 K01735 3-dehydroquinate synthase Amino acid metabolism aroB JTT Chromosome and associated 10 BJ6T_02040 K04763 integrase/recombinase XerD proteins xerD JTT DNA repair and recombination 11 BJ6T_02180 K03574 8-oxo-dGTP diphosphatase proteins mutT LG 12 BJ6T_02220 K11206 deaminated glutathione amidase Unclassified metabolism NIT1 CpREV 13 BJ6T_02230 Uncharacterised Unclassified JTT 14 BJ6T_02290 K02835 peptide chain release factor 1 Translation factors prfA JTT 15 BJ6T_02440 K00023 acetoacetyl-CoA reductase Carbohydrate metabolism phbB JTT 16 BJ6T_02460 Uncharacterised Unclassified JTT 17 BJ6T_02900 K01087 trehalose 6-phosphate phosphatase -
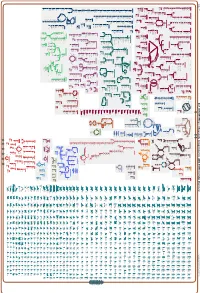
Generate Metabolic Map Poster
Authors: Pallavi Subhraveti Ron Caspi Peter Midford Peter D Karp An online version of this diagram is available at BioCyc.org. Biosynthetic pathways are positioned in the left of the cytoplasm, degradative pathways on the right, and reactions not assigned to any pathway are in the far right of the cytoplasm. Transporters and membrane proteins are shown on the membrane. Ingrid Keseler Periplasmic (where appropriate) and extracellular reactions and proteins may also be shown. Pathways are colored according to their cellular function. Gcf_003855395Cyc: Shewanella livingstonensis LMG 19866 Cellular Overview Connections between pathways are omitted for legibility.