Cataloguing Functionally Relevant Polymorphisms in Gene DNA Ligase I: a Computational Approach
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genome-Wide Approach to Identify Risk Factors for Therapy-Related Myeloid Leukemia
Leukemia (2006) 20, 239–246 & 2006 Nature Publishing Group All rights reserved 0887-6924/06 $30.00 www.nature.com/leu ORIGINAL ARTICLE Genome-wide approach to identify risk factors for therapy-related myeloid leukemia A Bogni1, C Cheng2, W Liu2, W Yang1, J Pfeffer1, S Mukatira3, D French1, JR Downing4, C-H Pui4,5,6 and MV Relling1,6 1Department of Pharmaceutical Sciences, The University of Tennessee, Memphis, TN, USA; 2Department of Biostatistics, The University of Tennessee, Memphis, TN, USA; 3Hartwell Center, The University of Tennessee, Memphis, TN, USA; 4Department of Pathology, The University of Tennessee, Memphis, TN, USA; 5Department of Hematology/Oncology St Jude Children’s Research Hospital, The University of Tennessee, Memphis, TN, USA; and 6Colleges of Medicine and Pharmacy, The University of Tennessee, Memphis, TN, USA Using a target gene approach, only a few host genetic risk therapy increases, the importance of identifying host factors for factors for treatment-related myeloid leukemia (t-ML) have been secondary neoplasms increases. defined. Gene expression microarrays allow for a more 4 genome-wide approach to assess possible genetic risk factors Because DNA microarrays interrogate multiple ( 10 000) for t-ML. We assessed gene expression profiles (n ¼ 12 625 genes in one experiment, they allow for a ‘genome-wide’ probe sets) in diagnostic acute lymphoblastic leukemic cells assessment of genes that may predispose to leukemogenesis. from 228 children treated on protocols that included leukemo- DNA microarray analysis of gene expression has been used to genic agents such as etoposide, 13 of whom developed t-ML. identify distinct expression profiles that are characteristic of Expression of 68 probes, corresponding to 63 genes, was different leukemia subtypes.13,14 Studies using this method have significantly related to risk of t-ML. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

The Middle Temporal Gyrus Is Transcriptionally Altered in Patients with Alzheimer’S Disease
1 The middle temporal gyrus is transcriptionally altered in patients with Alzheimer’s Disease. 2 1 3 Shahan Mamoor 1Thomas Jefferson School of Law 4 East Islip, NY 11730 [email protected] 5 6 We sought to understand, at the systems level and in an unbiased fashion, how gene 7 expression was most different in the brains of patients with Alzheimer’s Disease (AD) by mining published microarray datasets (1, 2). Comparing global gene expression profiles between 8 patient and control revealed that a set of 84 genes were expressed at significantly different levels in the middle temporal gyrus (MTG) of patients with Alzheimer’s Disease (1, 2). We used 9 computational analyses to classify these genes into known pathways and existing gene sets, 10 and to describe the major differences in the epigenetic marks at the genomic loci of these genes. While a portion of these genes is computationally cognizable as part of a set of genes 11 up-regulated in the brains of patients with AD (3), many other genes in the gene set identified here have not previously been studied in association with AD. Transcriptional repression, both 12 pre- and post-transcription appears to be affected; nearly 40% of these genes are transcriptional 13 targets of MicroRNA-19A/B (miR-19A/B), the zinc finger protein 10 (ZNF10), or of the AP-1 repressor jun dimerization protein 2 (JDP2). 14 15 16 17 18 19 20 21 22 23 24 25 26 Keywords: Alzheimer’s Disease, systems biology of Alzheimer’s Disease, differential gene 27 expression, middle temporal gyrus. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Bioinformatic Evaluation of Transcriptional Regulation of WNT Pathway Genes with Reference to Diabetic Nephropathy
Bioinformatic evaluation of transcriptional regulation of WNT pathway genes with reference to diabetic nephropathy McKay, G. J., Kavanagh, D. H., Crean, J. K., & Maxwell, A. P. (2015). Bioinformatic evaluation of transcriptional regulation of WNT pathway genes with reference to diabetic nephropathy. Journal of Diabetes Research, [608691]. https://doi.org/10.1155/2016/7684038 Published in: Journal of Diabetes Research Document Version: Publisher's PDF, also known as Version of record Queen's University Belfast - Research Portal: Link to publication record in Queen's University Belfast Research Portal Publisher rights Copyright © 2015 Gareth J. McKay et al.This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. General rights Copyright for the publications made accessible via the Queen's University Belfast Research Portal is retained by the author(s) and / or other copyright owners and it is a condition of accessing these publications that users recognise and abide by the legal requirements associated with these rights. Take down policy The Research Portal is Queen's institutional repository that provides access to Queen's research output. Every effort has been made to ensure that content in the Research Portal does not infringe any person's rights, or applicable UK laws. If you discover content in the Research Portal that you believe breaches copyright or violates any law, please contact [email protected]. Download date:27. Sep. 2021 Hindawi Publishing Corporation Journal of Diabetes Research Article ID 608691 Research Article Bioinformatic Evaluation of Transcriptional Regulation of WNT Pathway Genes with reference to Diabetic Nephropathy Gareth J. -
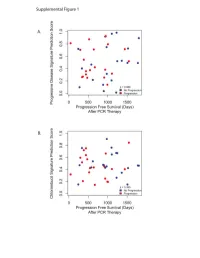
Supplementary Data
Progressive Disease Signature Upregulated probes with progressive disease U133Plus2 ID Gene Symbol Gene Name 239673_at NR3C2 nuclear receptor subfamily 3, group C, member 2 228994_at CCDC24 coiled-coil domain containing 24 1562245_a_at ZNF578 zinc finger protein 578 234224_at PTPRG protein tyrosine phosphatase, receptor type, G 219173_at NA NA 218613_at PSD3 pleckstrin and Sec7 domain containing 3 236167_at TNS3 tensin 3 1562244_at ZNF578 zinc finger protein 578 221909_at RNFT2 ring finger protein, transmembrane 2 1552732_at ABRA actin-binding Rho activating protein 59375_at MYO15B myosin XVB pseudogene 203633_at CPT1A carnitine palmitoyltransferase 1A (liver) 1563120_at NA NA 1560098_at AKR1C2 aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding pro 238576_at NA NA 202283_at SERPINF1 serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), m 214248_s_at TRIM2 tripartite motif-containing 2 204766_s_at NUDT1 nudix (nucleoside diphosphate linked moiety X)-type motif 1 242308_at MCOLN3 mucolipin 3 1569154_a_at NA NA 228171_s_at PLEKHG4 pleckstrin homology domain containing, family G (with RhoGef domain) member 4 1552587_at CNBD1 cyclic nucleotide binding domain containing 1 220705_s_at ADAMTS7 ADAM metallopeptidase with thrombospondin type 1 motif, 7 232332_at RP13-347D8.3 KIAA1210 protein 1553618_at TRIM43 tripartite motif-containing 43 209369_at ANXA3 annexin A3 243143_at FAM24A family with sequence similarity 24, member A 234742_at SIRPG signal-regulatory protein gamma -

Alterations of the Pro-Survival Bcl-2 Protein Interactome in Breast Cancer
bioRxiv preprint doi: https://doi.org/10.1101/695379; this version posted July 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Alterations of the pro-survival Bcl-2 protein interactome in 2 breast cancer at the transcriptional, mutational and 3 structural level 4 5 Simon Mathis Kønig1, Vendela Rissler1, Thilde Terkelsen1, Matteo Lambrughi1, Elena 6 Papaleo1,2 * 7 1Computational Biology Laboratory, Danish Cancer Society Research Center, 8 Strandboulevarden 49, 2100, Copenhagen 9 10 2Translational Disease Systems Biology, Faculty of Health and Medical Sciences, Novo 11 Nordisk Foundation Center for Protein Research University of Copenhagen, Copenhagen, 12 Denmark 13 14 Abstract 15 16 Apoptosis is an essential defensive mechanism against tumorigenesis. Proteins of the B-cell 17 lymphoma-2 (Bcl-2) family regulates programmed cell death by the mitochondrial apoptosis 18 pathway. In response to intracellular stresses, the apoptotic balance is governed by interactions 19 of three distinct subgroups of proteins; the activator/sensitizer BH3 (Bcl-2 homology 3)-only 20 proteins, the pro-survival, and the pro-apoptotic executioner proteins. Changes in expression 21 levels, stability, and functional impairment of pro-survival proteins can lead to an imbalance 22 in tissue homeostasis. Their overexpression or hyperactivation can result in oncogenic effects. 23 Pro-survival Bcl-2 family members carry out their function by binding the BH3 short linear 24 motif of pro-apoptotic proteins in a modular way, creating a complex network of protein- 25 protein interactions. -

The Mutational Landscape of the Oncogenic MZF1 SCAN Domain in Cancer
The Mutational Landscape of the Oncogenic MZF1 SCAN Domain in Cancer Nygaard, Mads; Terkelsen, Thilde; Vidas Olsen, André; Sora, Valentina; Salamanca Viloria, Juan; Rizza, Fabio; Bergstrand-Poulsen, Sanne; Di Marco, Miriam; Vistesen, Mette; Tiberti, Matteo; Lambrughi, Matteo; Jäättelä, Marja; Kallunki, Tuula; Papaleo, Elena Published in: Frontiers in Molecular Biosciences DOI: 10.3389/fmolb.2016.00078 Publication date: 2016 Document version Publisher's PDF, also known as Version of record Citation for published version (APA): Nygaard, M., Terkelsen, T., Vidas Olsen, A., Sora, V., Salamanca Viloria, J., Rizza, F., Bergstrand-Poulsen, S., Di Marco, M., Vistesen, M., Tiberti, M., Lambrughi, M., Jäättelä, M., Kallunki, T., & Papaleo, E. (2016). The Mutational Landscape of the Oncogenic MZF1 SCAN Domain in Cancer. Frontiers in Molecular Biosciences, 3, 1-18. [78]. https://doi.org/10.3389/fmolb.2016.00078 Download date: 27. sep.. 2021 ORIGINAL RESEARCH published: 15 December 2016 doi: 10.3389/fmolb.2016.00078 The Mutational Landscape of the Oncogenic MZF1 SCAN Domain in Cancer Mads Nygaard 1 †, Thilde Terkelsen 1 †, André Vidas Olsen 1, Valentina Sora 1, Juan Salamanca Viloria 1, Fabio Rizza 2, Sanne Bergstrand-Poulsen 1, Miriam Di Marco 1, Mette Vistesen 3, Matteo Tiberti 4, Matteo Lambrughi 1, Marja Jäättelä 5, Tuula Kallunki 5 and Elena Papaleo 1* 1 Computational Biology Laboratory and Center for Autophagy, Recycling and Disease, Danish Cancer Society Research Center, Copenhagen, Denmark, 2 Department of Biomedical Sciences, University -

The Changing Chromatome As a Driver of Disease: a Panoramic View from Different Methodologies
The changing chromatome as a driver of disease: A panoramic view from different methodologies Isabel Espejo1, Luciano Di Croce,1,2,3 and Sergi Aranda1 1. Centre for Genomic Regulation (CRG), Barcelona Institute of Science and Technology, Dr. Aiguader 88, Barcelona 08003, Spain 2. Universitat Pompeu Fabra (UPF), Barcelona, Spain 3. ICREA, Pg. Lluis Companys 23, Barcelona 08010, Spain *Corresponding authors: Luciano Di Croce ([email protected]) Sergi Aranda ([email protected]) 1 GRAPHICAL ABSTRACT Chromatin-bound proteins regulate gene expression, replicate and repair DNA, and transmit epigenetic information. Several human diseases are highly influenced by alterations in the chromatin- bound proteome. Thus, biochemical approaches for the systematic characterization of the chromatome could contribute to identifying new regulators of cellular functionality, including those that are relevant to human disorders. 2 SUMMARY Chromatin-bound proteins underlie several fundamental cellular functions, such as control of gene expression and the faithful transmission of genetic and epigenetic information. Components of the chromatin proteome (the “chromatome”) are essential in human life, and mutations in chromatin-bound proteins are frequently drivers of human diseases, such as cancer. Proteomic characterization of chromatin and de novo identification of chromatin interactors could thus reveal important and perhaps unexpected players implicated in human physiology and disease. Recently, intensive research efforts have focused on developing strategies to characterize the chromatome composition. In this review, we provide an overview of the dynamic composition of the chromatome, highlight the importance of its alterations as a driving force in human disease (and particularly in cancer), and discuss the different approaches to systematically characterize the chromatin-bound proteome in a global manner. -

Tommune Touttmann
TOMMUNEUS009952216B2TOUTTMANN (12 ) United States Patent ( 10 ) Patent No. : US 9 , 952 ,216 B2 Visel et al. (45 ) Date of Patent: Apr. 24 , 2018 ( 54 ) BRAIN - SPECIFIC ENHANCERS FOR C12N 15 / 86 (2006 .01 ) CELL -BASED THERAPY A61K 48 /00 (2006 .01 ) (52 ) U .S . CI. (71 ) Applicants :Axel Visel, El Cerrito , CA (US ) ; John CPC . .. .. GOIN 33 / 56966 (2013 .01 ) ; A61K 35 / 545 L . R . Rubenstein , San Francisco , CA ( 2013 .01 ) ; C12N 15 /85 ( 2013 .01 ) ; C12N (US ) ; Ying- Jiun ( Jasmine) Chen , 15 / 86 (2013 .01 ) ; GOIN 33/ 5014 ( 2013 .01 ) ; South San Francisco , CA ( US) ; Len A . A61K 48 / 00 ( 2013 .01 ) ; C12N 2830 /008 Pennacchio , Sebastopol, CA (US ) ; ( 2013 .01 ) Daniel Vogt, Burlingame, CA (US ) ; ( 58 ) Field of Classification Search Cory Nicholas, San Francisco , CA None (US ) ; Arnold Kriegstein , Mill Valley, See application file for complete search history . CA (US ) (56 ) References Cited ( 72 ) Inventors : Axel Visel , El Cerrito , CA ( US ) ; John L . R . Rubenstein , San Francisco , CA U . S . PATENT DOCUMENTS (US ) ; Ying - Jiun ( Jasmine ) Chen , 5 ,436 , 128 A * 7 /1995 Harpold . .. GOIN 33/ 6872 South San Francisco , CA (US ) ; Len A . 435 / 365 Pennacchio , Sebastopol, CA (US ) ; Daniel Vogt, Burlingame, CA (US ) ; OTHER PUBLICATIONS Cory Nicholas, San Francisco , CA (US ) ; Arnold Kriegstein , Mill Valley , Ferguson et al . The human synaptotagmin IV gene defines an CA (US ) evolutionary break point between syntenic mouse and human chro mosome regions but retains ligand inducibility and tissue specific ( 73 ) Assignee: The Regents of the University of ity . The Journal of Biological Chemistry , vol . 275 , No . 47 , pp . 36920 - 36926 , 2000 . -

An Evaluation of Cancer Subtypes and Glioma Stem Cell Characterisation Unifying Tumour Transcriptomic Features with Cell Line Expression and Chromatin Accessibility
An evaluation of cancer subtypes and glioma stem cell characterisation Unifying tumour transcriptomic features with cell line expression and chromatin accessibility Ewan Roderick Johnstone EMBL-EBI, Darwin College University of Cambridge This dissertation is submitted for the degree of Doctor of Philosophy Darwin College December 2016 Dedicated to Klaudyna. Declaration • I hereby declare that except where specific reference is made to the work of others, the contents of this dissertation are original and have not been submitted in whole or in part for consideration for any other degree or qualification in this, or any other university. • This dissertation is my own work and contains nothing which is the outcome of work done in collaboration with others, except as specified in the text and Acknowledge- ments. • This dissertation is typeset in LATEX using one-and-a-half spacing, contains fewer than 60,000 words including appendices, footnotes, tables and equations and has fewer than 150 figures. Ewan Roderick Johnstone December 2016 Acknowledgements This work was funded by the Biotechnology and Biological Sciences Research Council (BBSRC, Ref:1112564) and supported by the European Molecular Biology Laboratory (EMBL) and its outstation, the European Bioinformatics Institute (EBI). I have many people to thank for assistance in preparing this thesis. First and foremost I must thank my supervisor, Paul Bertone for his support and willingness to take me on as a student. My thanks are also extended to present and past members of the Bertone group, particularly Pär Engström and Remco Loos who have provided a great deal of guidance over the course of my studentship. -

Global Gene Expression of the Inner Cell Mass and Trophectoderm of The
Ozawa et al. BMC Developmental Biology 2012, 12:33 http://www.biomedcentral.com/1471-213X/12/33 RESEARCHARTICLE Open Access Global gene expression of the inner cell mass and trophectoderm of the bovine blastocyst Manabu Ozawa1,4, Miki Sakatani3, JiQiang Yao2, Savita Shanker2, Fahong Yu2, Rui Yamashita5, Shunichi Wakabayashi5, Kenta Nakai5, Kyle B Dobbs1, Mateus José Sudano6, William G Farmerie2 and Peter J Hansen1* Abstract Background: The first distinct differentiation event in mammals occurs at the blastocyst stage when totipotent blastomeres differentiate into either pluripotent inner cell mass (ICM) or multipotent trophectoderm (TE). Here we determined, for the first time, global gene expression patterns in the ICM and TE isolated from bovine blastocysts. The ICM and TE were isolated from blastocysts harvested at day 8 after insemination by magnetic activated cell sorting, and cDNA sequenced using the SOLiD 4.0 system. Results: A total of 870 genes were differentially expressed between ICM and TE. Several genes characteristic of ICM (for example, NANOG, SOX2, and STAT3) and TE (ELF5, GATA3, and KRT18) in mouse and human showed similar patterns in bovine. Other genes, however, showed differences in expression between ICM and TE that deviates from the expected based on mouse and human. Conclusion: Analysis of gene expression indicated that differentiation of blastomeres of the morula-stage embryo into the ICM and TE of the blastocyst is accompanied by differences between the two cell lineages in expression of genes controlling metabolic processes, endocytosis, hatching from the zona pellucida, paracrine and endocrine signaling with the mother, and genes supporting the changes in cellular architecture, stemness, and hematopoiesis necessary for development of the trophoblast.