Rapid Birth–Death Evolution Specific to Xenobiotic Cytochrome P450 Genes in Vertebrates
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

CYP26C1 Is a Hydroxylase of Multiple Active Retinoids and Interacts with Cellular Retinoic Acid Binding Proteins S
Supplemental material to this article can be found at: http://molpharm.aspetjournals.org/content/suppl/2018/02/23/mol.117.111039.DC1 1521-0111/93/5/489–503$35.00 https://doi.org/10.1124/mol.117.111039 MOLECULAR PHARMACOLOGY Mol Pharmacol 93:489–503, May 2018 Copyright ª 2018 by The American Society for Pharmacology and Experimental Therapeutics CYP26C1 Is a Hydroxylase of Multiple Active Retinoids and Interacts with Cellular Retinoic Acid Binding Proteins s Guo Zhong, David Ortiz, Alex Zelter, Abhinav Nath, and Nina Isoherranen Departments of Pharmaceutics (G.Z., N.I.) and Medicinal Chemistry (D.O., A.N.), School of Pharmacy, and Department of Biochemistry, School of Medicine (A.Z.), University of Washington, Seattle, Washington Received October 31, 2017; accepted February 22, 2018 ABSTRACT Downloaded from The clearance of retinoic acid (RA) and its metabolites is believed orientation of retinoids within the CYP26C1 active site. In compar- to be regulated by the CYP26 enzymes, but the specific roles of ison with other CYP26 family members, CYP26C1 was up to CYP26A1, CYP26B1, and CYP26C1 in clearing active vitamin A 10-fold more efficient in clearing 4-oxo-atRA (intrinsic clearance metabolites have not been defined. The goal of this study was to 153 ml/min/pmol) than CYP26A1 and CYP26B1, suggesting that establish the substrate specificity of CYP26C1, and determine CYP26C1 may be important in clearing this active retinoid. In whether CYP26C1 interacts with cellular retinoic acid binding support of this, CRABPs delivered 4-oxo-atRA and atRA for proteins (CRABPs). CYP26C1 was found to effectively metabo- metabolism by CYP26C1. -

Impaired Hepatic Drug and Steroid Metabolism in Congenital Adrenal
European Journal of Endocrinology (2010) 163 919–924 ISSN 0804-4643 CLINICAL STUDY Impaired hepatic drug and steroid metabolism in congenital adrenal hyperplasia due to P450 oxidoreductase deficiency Dorota Tomalik-Scharte1, Dominique Maiter2, Julia Kirchheiner3, Hannah E Ivison, Uwe Fuhr1 and Wiebke Arlt School of Clinical and Experimental Medicine, Centre for Endocrinology, Diabetes and Metabolism (CEDAM), University of Birmingham, Birmingham B15 2TT, UK, 1Department of Pharmacology, University Hospital, University of Cologne, 50931 Cologne, Germany, 2Department of Endocrinology, University Hospital Saint Luc, 1200 Brussels, Belgium and 3Department of Pharmacology of Natural Products and Clinical Pharmacology, University of Ulm, 89019 Ulm, Germany (Correspondence should be addressed to W Arlt; Email: [email protected]) Abstract Objective: Patients with congenital adrenal hyperplasia due to P450 oxidoreductase (POR) deficiency (ORD) present with disordered sex development and glucocorticoid deficiency. This is due to disruption of electron transfer from mutant POR to microsomal cytochrome P450 (CYP) enzymes that play a key role in glucocorticoid and sex steroid synthesis. POR also transfers electrons to all major drug- metabolizing CYP enzymes, including CYP3A4 that inactivates glucocorticoid and oestrogens. However, whether ORD results in impairment of in vivo drug metabolism has never been studied. Design: We studied an adult patient with ORD due to homozygous POR A287P, the most frequent POR mutation in Caucasians, and her clinically unaffected, heterozygous mother. The patient had received standard dose oestrogen replacement from 17 until 37 years of age when it was stopped after she developed breast cancer. Methods: Both subjects underwent in vivo cocktail phenotyping comprising the oral administration of caffeine, tolbutamide, omeprazole, dextromethorphan hydrobromide and midazolam to assess the five major drug-metabolizing CYP enzymes. -

Identification and Developmental Expression of the Full Complement Of
Goldstone et al. BMC Genomics 2010, 11:643 http://www.biomedcentral.com/1471-2164/11/643 RESEARCH ARTICLE Open Access Identification and developmental expression of the full complement of Cytochrome P450 genes in Zebrafish Jared V Goldstone1, Andrew G McArthur2, Akira Kubota1, Juliano Zanette1,3, Thiago Parente1,4, Maria E Jönsson1,5, David R Nelson6, John J Stegeman1* Abstract Background: Increasing use of zebrafish in drug discovery and mechanistic toxicology demands knowledge of cytochrome P450 (CYP) gene regulation and function. CYP enzymes catalyze oxidative transformation leading to activation or inactivation of many endogenous and exogenous chemicals, with consequences for normal physiology and disease processes. Many CYPs potentially have roles in developmental specification, and many chemicals that cause developmental abnormalities are substrates for CYPs. Here we identify and annotate the full suite of CYP genes in zebrafish, compare these to the human CYP gene complement, and determine the expression of CYP genes during normal development. Results: Zebrafish have a total of 94 CYP genes, distributed among 18 gene families found also in mammals. There are 32 genes in CYP families 5 to 51, most of which are direct orthologs of human CYPs that are involved in endogenous functions including synthesis or inactivation of regulatory molecules. The high degree of sequence similarity suggests conservation of enzyme activities for these CYPs, confirmed in reports for some steroidogenic enzymes (e.g. CYP19, aromatase; CYP11A, P450scc; CYP17, steroid 17a-hydroxylase), and the CYP26 retinoic acid hydroxylases. Complexity is much greater in gene families 1, 2, and 3, which include CYPs prominent in metabolism of drugs and pollutants, as well as of endogenous substrates. -

(12) Patent Application Publication (10) Pub. No.: US 2011/0190389 A1 Arterburn Et Al
US 2011 0190389A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2011/0190389 A1 Arterburn et al. (43) Pub. Date: Aug. 4, 2011 (54) OXYLIPINS FROM LONG CHAIN C07C 57/03 (2006.01) POLYUNSATURATED FATTY ACDS AND A6IP 29/00 (2006.01) METHODS OF MAKING AND USING THE CI2P 7/64 (2006.01) SAME CI2P I 7/02 (2006.01) C07D 303/38 (2006.01) (76) Inventors: Linda Arterburn, Ellicott City, A6IP 25/28 (2006.01) MD (US); William Barclay, Boulder, CO (US); Bindi Dangi, (52) U.S. Cl. ......... 514/475: 514/560; 554/124; 435/134; Elkridge, MD (US); James Flatt, 435/123:549/561 Colorado Springs, CO (US); Jung Lee, McLean, VA (US); Dutt (57) ABSTRACT Vinjamoori, Chesterfield, MO Disclosed are novel oxylipins, referred to herein as (US); Mary Van Elswyk, docosanoids and eicosanoids, that are derived from C22 poly Longmont, CO (US) unsaturated fatty acids and from C20 polyunsaturated fatty acids, respectively, and methods of making and using Such (21) Appl. No.: 12/531,344 oxylipins. Also disclosed is the use of docosapentaenoic acid 1-1. (C22:5n-6) (DPAn-6), docosapentaenoic acid (C22:5n-3) (22) PCT Filed: Feb. 20, 2008 (DPAn-3), and docosatetraenoic acid (DTAn-6: C22:4n-6), (86). PCT No.: PCT/USO8/54.456 docosatrienoic acid (C22:3n-3) (DTrAn-3), docosadienoic acid (C22:2n-6) (DDAn-6), eicosatrienoic acid (C20:3n-3) S371 (c)(1), (ETrAn-3) eicosapentaenoic acid and arachidonic acid as (2), (4) Date: Feb. 15, 2011 substrates for the production of novel oxylipins, and to the oxylipins produced thereby. -

Methylation Status of Vitamin D Receptor Gene Promoter in Adrenocortical Carcinoma
UNIVERSITÀ DEGLI STUDI DI PADOVA DEPARTMENT OF CARDIAC, THORACIC AND VASCULAR SCIENCES Ph.D Course Medical Clinical and Experimental Sciences Curriculum Clinical Methodology, Endocrinological, Diabetological and Nephrological Sciences XXIX° SERIES METHYLATION STATUS OF VITAMIN D RECEPTOR GENE PROMOTER IN ADRENOCORTICAL CARCINOMA Coordinator: Ch.mo Prof. Annalisa Angelini Supervisor: Ch.mo Prof. Francesco Fallo Ph.D Student: Andrea Rebellato TABLE OF CONTENTS SUMMARY 3 INTRODUCTION 4 PART 1: ADRENOCORTICAL CARCINOMA 4 1.1 EPIDEMIOLOGY 4 1.2 GENETIC PREDISPOSITION 4 1.3 CLINICAL PRESENTATION 6 1.4 DIAGNOSTIC WORK-UP 7 1.4.1 Biochemistry 7 1.4.2 Imaging 9 1.5 STAGING 10 1.6 PATHOLOGY 11 1.7 MOLECULAR PATHOLOGY 14 1.7.1 DNA content 15 1.7.2 Chromosomal aberrations 15 1.7.3 Differential gene expression 16 1.7.4 DNA methylation 17 1.7.5 microRNAs 18 1.7.6 Gene mutations 19 1.8 PATHOPHYSIOLOGY OF MOLECULAR SIGNALLING 21 PATHWAYS 1.8.1 IGF-mTOR pathway 21 1.8.2 WNTsignalling pathway 22 1.8.3 Vascular endothelial growth factor 23 1.9 THERAPY 24 1.9.1 Surgery 24 1.9.2 Adjuvant Therapy 27 1.9.2.1 Mitotane 27 1.9.2.2 Cytotoxic chemotherapy 30 1.9.2.3 Targeted therapy 31 1.9.2.4 Therapy for hormone excess 31 1.9.2.5 Radiation therapy 32 1.9.2.6 Other local therapies 32 1.10 PROGNOSTIC FACTORS AND PREDICTIVE MARKERS 32 PART 2: VITAMIN D 35 2.1 VITAMIN D AND ITS BIOACTIVATION 35 2.2 THE VITAMIN D RECEPTOR 37 2.3 GENOMIC MECHANISM OF 1,25(OH)2D3-VDR COMPLEX 38 2.4 CLASSICAL ROLES OF VITAMIN D 40 2.4.1 Intestine 40 2.4.2 Kidney 41 2.4.3 Bone 41 2.5 PLEIOTROPIC -

Cytochrome P450 Enzymes in Oxygenation of Prostaglandin Endoperoxides and Arachidonic Acid
Comprehensive Summaries of Uppsala Dissertations from the Faculty of Pharmacy 231 _____________________________ _____________________________ Cytochrome P450 Enzymes in Oxygenation of Prostaglandin Endoperoxides and Arachidonic Acid Cloning, Expression and Catalytic Properties of CYP4F8 and CYP4F21 BY JOHAN BYLUND ACTA UNIVERSITATIS UPSALIENSIS UPPSALA 2000 Dissertation for the Degree of Doctor of Philosophy (Faculty of Pharmacy) in Pharmaceutical Pharmacology presented at Uppsala University in 2000 ABSTRACT Bylund, J. 2000. Cytochrome P450 Enzymes in Oxygenation of Prostaglandin Endoperoxides and Arachidonic Acid: Cloning, Expression and Catalytic Properties of CYP4F8 and CYP4F21. Acta Universitatis Upsaliensis. Comprehensive Summaries of Uppsala Dissertations from Faculty of Pharmacy 231 50 pp. Uppsala. ISBN 91-554-4784-8. Cytochrome P450 (P450 or CYP) is an enzyme system involved in the oxygenation of a wide range of endogenous compounds as well as foreign chemicals and drugs. This thesis describes investigations of P450-catalyzed oxygenation of prostaglandins, linoleic and arachidonic acids. The formation of bisallylic hydroxy metabolites of linoleic and arachidonic acids was studied with human recombinant P450s and with human liver microsomes. Several P450 enzymes catalyzed the formation of bisallylic hydroxy metabolites. Inhibition studies and stereochemical analysis of metabolites suggest that the enzyme CYP1A2 may contribute to the biosynthesis of bisallylic hydroxy fatty acid metabolites in adult human liver microsomes. 19R-Hydroxy-PGE and 20-hydroxy-PGE are major components of human and ovine semen, respectively. They are formed in the seminal vesicles, but the mechanism of their biosynthesis is unknown. Reverse transcription-polymerase chain reaction using degenerate primers for mammalian CYP4 family genes, revealed expression of two novel P450 genes in human and ovine seminal vesicles. -

Transcriptomic Characterization of Fibrolamellar Hepatocellular
Transcriptomic characterization of fibrolamellar PNAS PLUS hepatocellular carcinoma Elana P. Simona, Catherine A. Freijeb, Benjamin A. Farbera,c, Gadi Lalazara, David G. Darcya,c, Joshua N. Honeymana,c, Rachel Chiaroni-Clarkea, Brian D. Dilld, Henrik Molinad, Umesh K. Bhanote, Michael P. La Quagliac, Brad R. Rosenbergb,f, and Sanford M. Simona,1 aLaboratory of Cellular Biophysics, The Rockefeller University, New York, NY 10065; bPresidential Fellows Laboratory, The Rockefeller University, New York, NY 10065; cDivision of Pediatric Surgery, Department of Surgery, Memorial Sloan-Kettering Cancer Center, New York, NY 10065; dProteomics Resource Center, The Rockefeller University, New York, NY 10065; ePathology Core Facility, Memorial Sloan-Kettering Cancer Center, New York, NY 10065; and fJohn C. Whitehead Presidential Fellows Program, The Rockefeller University, New York, NY 10065 Edited by Susan S. Taylor, University of California, San Diego, La Jolla, CA, and approved September 22, 2015 (received for review December 29, 2014) Fibrolamellar hepatocellular carcinoma (FLHCC) tumors all carry a exon of DNAJB1 and all but the first exon of PRKACA. This deletion of ∼400 kb in chromosome 19, resulting in a fusion of the produced a chimeric RNA transcript and a translated chimeric genes for the heat shock protein, DNAJ (Hsp40) homolog, subfam- protein that retains the full catalytic activity of wild-type PKA. ily B, member 1, DNAJB1, and the catalytic subunit of protein ki- This chimeric protein was found in 15 of 15 FLHCC patients nase A, PRKACA. The resulting chimeric transcript produces a (21) in the absence of any other recurrent mutations in the DNA fusion protein that retains kinase activity. -

Synonymous Single Nucleotide Polymorphisms in Human Cytochrome
DMD Fast Forward. Published on February 9, 2009 as doi:10.1124/dmd.108.026047 DMD #26047 TITLE PAGE: A BIOINFORMATICS APPROACH FOR THE PHENOTYPE PREDICTION OF NON- SYNONYMOUS SINGLE NUCLEOTIDE POLYMORPHISMS IN HUMAN CYTOCHROME P450S LIN-LIN WANG, YONG LI, SHU-FENG ZHOU Department of Nutrition and Food Hygiene, School of Public Health, Peking University, Beijing 100191, P. R. China (LL Wang & Y Li) Discipline of Chinese Medicine, School of Health Sciences, RMIT University, Bundoora, Victoria 3083, Australia (LL Wang & SF Zhou). 1 Copyright 2009 by the American Society for Pharmacology and Experimental Therapeutics. DMD #26047 RUNNING TITLE PAGE: a) Running title: Prediction of phenotype of human CYPs. b) Author for correspondence: A/Prof. Shu-Feng Zhou, MD, PhD Discipline of Chinese Medicine, School of Health Sciences, RMIT University, WHO Collaborating Center for Traditional Medicine, Bundoora, Victoria 3083, Australia. Tel: + 61 3 9925 7794; fax: +61 3 9925 7178. Email: [email protected] c) Number of text pages: 21 Number of tables: 10 Number of figures: 2 Number of references: 40 Number of words in Abstract: 249 Number of words in Introduction: 749 Number of words in Discussion: 1459 d) Non-standard abbreviations: CYP, cytochrome P450; nsSNP, non-synonymous single nucleotide polymorphism. 2 DMD #26047 ABSTRACT Non-synonymous single nucleotide polymorphisms (nsSNPs) in coding regions that can lead to amino acid changes may cause alteration of protein function and account for susceptivity to disease. Identification of deleterious nsSNPs from tolerant nsSNPs is important for characterizing the genetic basis of human disease, assessing individual susceptibility to disease, understanding the pathogenesis of disease, identifying molecular targets for drug treatment and conducting individualized pharmacotherapy. -

Colorectal Cancer and Omega Hydroxylases
1 The differential expression of omega-3 and omega-6 fatty acid metabolising enzymes in colorectal cancer and its prognostic significance Abdo Alnabulsi1,2, Rebecca Swan1, Beatriz Cash2, Ayham Alnabulsi2, Graeme I Murray1 1Pathology, School of Medicine, Medical Sciences and Nutrition, University of Aberdeen, Aberdeen, AB25, 2ZD, UK. 2Vertebrate Antibodies, Zoology Building, Tillydrone Avenue, Aberdeen, AB24 2TZ, UK. Address correspondence to: Professor Graeme I Murray Email [email protected] Phone: +44(0)1224 553794 Fax: +44(0)1224 663002 Running title: omega hydroxylases and colorectal cancer 2 Abstract Background: Colorectal cancer is a common malignancy and one of the leading causes of cancer related deaths. The metabolism of omega fatty acids has been implicated in tumour growth and metastasis. Methods: This study has characterised the expression of omega fatty acid metabolising enzymes CYP4A11, CYP4F11, CYP4V2 and CYP4Z1 using monoclonal antibodies we have developed. Immunohistochemistry was performed on a tissue microarray containing 650 primary colorectal cancers, 285 lymph node metastasis and 50 normal colonic mucosa. Results: The differential expression of CYP4A11 and CYP4F11 showed a strong association with survival in both the whole patient cohort (HR=1.203, 95% CI=1.092-1.324, χ2=14.968, p=0.001) and in mismatch repair proficient tumours (HR=1.276, 95% CI=1.095-1.488, χ2=9.988, p=0.007). Multivariate analysis revealed that the differential expression of CYP4A11 and CYP4F11 was independently prognostic in both the whole patient cohort (p = 0.019) and in mismatch repair proficient tumours (p=0.046). Conclusions: A significant and independent association has been identified between overall survival and the differential expression of CYP4A11 and CYP4F11 in the whole patient cohort and in mismatch repair proficient tumours. -

Comparative Proteomics Analysis of Human Liver Microsomes and S9
DMD Fast Forward. Published on November 7, 2019 as DOI: 10.1124/dmd.119.089235 This article has not been copyedited and formatted. The final version may differ from this version. DMD # 89235 Comparative Proteomics Analysis of Human Liver Microsomes and S9 Fractions Xinwen Wang, Bing He, Jian Shi, Qian Li, and Hao-Jie Zhu Department of Clinical Pharmacy, University of Michigan, Ann Arbor, Michigan (X.W., B.H., J.S., H.-J.Z.); and School of Life Science and Technology, China Pharmaceutical University, Nanjing, Jiangsu, 210009 (Q.L.) Downloaded from dmd.aspetjournals.org at ASPET Journals on October 2, 2021 1 DMD Fast Forward. Published on November 7, 2019 as DOI: 10.1124/dmd.119.089235 This article has not been copyedited and formatted. The final version may differ from this version. DMD # 89235 Running title: Comparative Proteomics of Human Liver Microsomes and S9 Corresponding author: Hao-Jie Zhu Ph.D. Department of Clinical Pharmacy University of Michigan College of Pharmacy 428 Church Street, Room 4565 Downloaded from Ann Arbor, MI 48109-1065 Tel: 734-763-8449, E-mail: [email protected] dmd.aspetjournals.org Number of words in Abstract: 250 at ASPET Journals on October 2, 2021 Number of words in Introduction: 776 Number of words in Discussion: 2304 2 DMD Fast Forward. Published on November 7, 2019 as DOI: 10.1124/dmd.119.089235 This article has not been copyedited and formatted. The final version may differ from this version. DMD # 89235 Non-standard ABBreviations: DMEs, drug metabolism enzymes; HLM, human liver microsomes; HLS9, -

Investigation of the Underlying Hub Genes and Molexular Pathogensis in Gastric Cancer by Integrated Bioinformatic Analyses
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Investigation of the underlying hub genes and molexular pathogensis in gastric cancer by integrated bioinformatic analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract The high mortality rate of gastric cancer (GC) is in part due to the absence of initial disclosure of its biomarkers. The recognition of important genes associated in GC is therefore recommended to advance clinical prognosis, diagnosis and and treatment outcomes. The current investigation used the microarray dataset GSE113255 RNA seq data from the Gene Expression Omnibus database to diagnose differentially expressed genes (DEGs). Pathway and gene ontology enrichment analyses were performed, and a proteinprotein interaction network, modules, target genes - miRNA regulatory network and target genes - TF regulatory network were constructed and analyzed. Finally, validation of hub genes was performed. The 1008 DEGs identified consisted of 505 up regulated genes and 503 down regulated genes. -
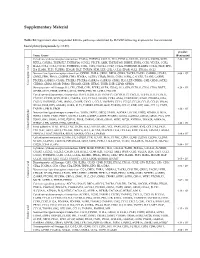
Supplementary Material
Supplementary Material Table S1: Significant downregulated KEGGs pathways identified by DAVID following exposure to five cinnamon- based phenylpropanoids (p < 0.05). p-value Term: Genes (Benjamini) Cytokine-cytokine receptor interaction: FASLG, TNFSF14, CXCL11, IL11, FLT3LG, CCL3L1, CCL3L3, CXCR6, XCR1, 2.43 × 105 RTEL1, CSF2RA, TNFRSF17, TNFRSF14, CCNL2, VEGFB, AMH, TNFRSF10B, INHBE, IFNB1, CCR3, VEGFA, CCR2, IL12A, CCL1, CCL3, CXCL5, TNFRSF25, CCR1, CSF1, CX3CL1, CCL7, CCL24, TNFRSF1B, IL12RB1, CCL21, FIGF, EPO, IL4, IL18R1, FLT1, TGFBR1, EDA2R, HGF, TNFSF8, KDR, LEP, GH2, CCL13, EPOR, XCL1, IFNA16, XCL2 Neuroactive ligand-receptor interaction: OPRM1, THRA, GRIK1, DRD2, GRIK2, TACR2, TACR1, GABRB1, LPAR4, 9.68 × 105 GRIK5, FPR1, PRSS1, GNRHR, FPR2, EDNRA, AGTR2, LTB4R, PRSS2, CNR1, S1PR4, CALCRL, TAAR5, GABRE, PTGER1, GABRG3, C5AR1, PTGER3, PTGER4, GABRA6, GABRA5, GRM1, PLG, LEP, CRHR1, GH2, GRM3, SSTR2, Chlorogenic acid Chlorogenic CHRM3, GRIA1, MC2R, P2RX2, TBXA2R, GHSR, HTR2C, TSHR, LHB, GLP1R, OPRD1 Hematopoietic cell lineage: IL4, CR1, CD8B, CSF1, FCER2, GYPA, ITGA2, IL11, GP9, FLT3LG, CD38, CD19, DNTT, 9.29 × 104 GP1BB, CD22, EPOR, CSF2RA, CD14, THPO, EPO, HLA-DRA, ITGA2B Cytokine-cytokine receptor interaction: IL6ST, IL21R, IL19, TNFSF15, CXCR3, IL15, CXCL11, TGFB1, IL11, FLT3LG, CXCL10, CCR10, XCR1, RTEL1, CSF2RA, IL21, CCNL2, VEGFB, CCR8, AMH, TNFRSF10C, IFNB1, PDGFRA, EDA, CXCL5, TNFRSF25, CSF1, IFNW1, CNTFR, CX3CL1, CCL5, TNFRSF4, CCL4, CCL27, CCL24, CCL25, CCL23, IFNA6, IFNA5, FIGF, EPO, AMHR2, IL2RA, FLT4, TGFBR2, EDA2R,