Context-Specific Regulation of Monocyte Surface IL7R Expression And
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Expression, Network Analysis, and Drug Discovery of Neurofibromatosis Type 2-Associated Vestibular Schwannomas Based on Bioinformatics Analysis
Hindawi Journal of Oncology Volume 2020, Article ID 5976465, 9 pages https://doi.org/10.1155/2020/5976465 Research Article Gene Expression, Network Analysis, and Drug Discovery of Neurofibromatosis Type 2-Associated Vestibular Schwannomas Based on Bioinformatics Analysis Qiao Huang , Si-Jia Zhai , Xing-Wei Liao , Yu-Chao Liu, and Shi-Hua Yin Department of Otolaryngology & Head and Neck Surgery, e Second Affiliated Hospital of Guangxi Medical University, Nanning 530007, China Correspondence should be addressed to Shi-Hua Yin; [email protected] Received 26 March 2020; Revised 27 May 2020; Accepted 1 June 2020; Published 15 July 2020 Academic Editor: Pierfrancesco Franco Copyright © 2020 Qiao Huang et al. ,is is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Neurofibromatosis Type 2- (NF2-) associated vestibular schwannomas (VSs) are histologically benign tumors. ,is study aimed to determine disease-related genes, pathways, and potential therapeutic drugs associated with NF2-VSs using the bioinformatics method. Microarray data of GSE108524 were downloaded from the Gene Expression Omnibus (GEO) database, and differentially expressed genes (DEGs) were screened using GEO2R. ,e functional enrichment and pathway enrichment of DEGs were performed using Gene Ontology (GO) and Kyoto Encyclopedia of Genes Genomes (KEGG). Furthermore, the STRING and Cytoscape were used to analyze the protein-protein interaction (PPI) network of all differentially expressed genes and identify hub genes. Finally, the enriched gene sets belonging to the identified pathways were queried against the Drug-Gene Interaction database to find drug candidates for topical use in NF2-associated VSs. -

Galectin-4 Interaction with CD14 Triggers the Differentiation of Monocytes Into Macrophage-Like Cells Via the MAPK Signaling Pathway
Immune Netw. 2019 Jun;19(3):e17 https://doi.org/10.4110/in.2019.19.e17 pISSN 1598-2629·eISSN 2092-6685 Original Article Galectin-4 Interaction with CD14 Triggers the Differentiation of Monocytes into Macrophage-like Cells via the MAPK Signaling Pathway So-Hee Hong 1,2,3,4,5, Jun-Seop Shin 1,2,3,5, Hyunwoo Chung 1,2,4,5, Chung-Gyu Park 1,2,3,4,5,* 1Xenotransplantation Research Center, Seoul National University College of Medicine, Seoul 03080, Korea 2Institute of Endemic Diseases, Seoul National University College of Medicine, Seoul 03080, Korea 3Cancer Research Institute, Seoul National University College of Medicine, Seoul 03080, Korea 4Department of Biomedical Sciences, Seoul National University College of Medicine, Seoul 03080, Korea 5Department of Microbiology and Immunology, Seoul National University College of Medicine, Seoul 03080, Korea Received: Jan 28, 2019 ABSTRACT Revised: May 13, 2019 Accepted: May 19, 2019 Galectin-4 (Gal-4) is a β-galactoside-binding protein mostly expressed in the gastrointestinal *Correspondence to tract of animals. Although intensive functional studies have been done for other galectin Chung-Gyu Park isoforms, the immunoregulatory function of Gal-4 still remains ambiguous. Here, we Department of Microbiology and Immunology, Seoul National University College of Medicine, demonstrated that Gal-4 could bind to CD14 on monocytes and induce their differentiation 103 Daehak-ro, Jongno-gu, Seoul 03080, into macrophage-like cells through the MAPK signaling pathway. Gal-4 induced the phenotypic Korea. changes on monocytes by altering the expression of various surface molecules, and induced E-mail: [email protected] functional changes such as increased cytokine production and matrix metalloproteinase expression and reduced phagocytic capacity. -

Associations Between FCGR3A Polymorphisms and Susceptibility to Rheumatoid Arthritis: a Metaanalysis YOUNG HO LEE, JONG DAE JI, and GWAN GYU SONG
Associations Between FCGR3A Polymorphisms and Susceptibility to Rheumatoid Arthritis: A Metaanalysis YOUNG HO LEE, JONG DAE JI, and GWAN GYU SONG ABSTRACT. Objective. To investigate whether the Fcγ receptor (FCGR) polymorphism confers susceptibility to rheumatoid arthritis (RA). Methods. We conducted metaanalyses on the associations between FCGR polymorphisms and RA susceptibility as determined using (1) allele contrast, (2) recessive models, (3) dominant models, and (4) contrast of homozygotes, using fixed or random effects models. Results. A total of 10 separate comparisons were considered, which comprised 6 European and 4 Asian population samples. Metaanalysis of FCGR3A polymorphism revealed a significant associa- tion between the VV genotype and the risk of developing RA relative to the VF+FF genotype (OR 1.256, 95% CI 1.045–1.510, p = 0.015), with no evidence of between-study heterogeneity (p = 0.167). In subjects of European descent, a stronger association was observed between the VV geno- type and RA than for the FF genotype (OR 1.374, 95% CI 1.101–1.714, p = 0.005). In Asians, no such association was found. Metaanalysis of the VV vs FF genotype revealed a significantly increased OR in Europeans (OR 1.399, 95% CI 1.107–1.769, p = 0.005), but not in Asians. No asso- ciation was found between RA and the FCGR2A and FCGR3B polymorphisms in all subjects and in European and Asian populations, except for the NA22 vs NA11 of FCGR3B in Europeans. Conclusion. No relation was found between the FCGR2A polymorphism and susceptibility to RA in Europeans or Asians. The FCGR3A polymorphism was found to be associated with RA in Europeans but not in Asians. -

IL21R Expressing CD14+CD16+ Monocytes Expand in Multiple
Plasma Cell Disorders SUPPLEMENTARY APPENDIX IL21R expressing CD14 +CD16 + monocytes expand in multiple myeloma patients leading to increased osteoclasts Marina Bolzoni, 1 Domenica Ronchetti, 2,3 Paola Storti, 1,4 Gaetano Donofrio, 5 Valentina Marchica, 1,4 Federica Costa, 1 Luca Agnelli, 2,3 Denise Toscani, 1 Rosanna Vescovini, 1 Katia Todoerti, 6 Sabrina Bonomini, 7 Gabriella Sammarelli, 1,7 Andrea Vecchi, 8 Daniela Guasco, 1 Fabrizio Accardi, 1,7 Benedetta Dalla Palma, 1,7 Barbara Gamberi, 9 Carlo Ferrari, 8 Antonino Neri, 2,3 Franco Aversa 1,4,7 and Nicola Giuliani 1,4,7 1Myeloma Unit, Dept. of Medicine and Surgery, University of Parma; 2Dept. of Oncology and Hemato-Oncology, University of Milan; 3Hematology Unit, “Fondazione IRCCS Ca’ Granda”, Ospedale Maggiore Policlinico, Milan; 4CoreLab, University Hospital of Parma; 5Dept. of Medical-Veterinary Science, University of Parma; 6Laboratory of Pre-clinical and Translational Research, IRCCS-CROB, Referral Cancer Center of Basilicata, Rionero in Vulture; 7Hematology and BMT Center, University Hospital of Parma; 8Infectious Disease Unit, University Hospital of Parma and 9“Dip. Oncologico e Tecnologie Avanzate”, IRCCS Arcispedale Santa Maria Nuova, Reggio Emilia, Italy ©2017 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol. 2016.153841 Received: August 5, 2016. Accepted: December 23, 2016. Pre-published: January 5, 2017. Correspondence: [email protected] SUPPLEMENTAL METHODS Immunophenotype of BM CD14+ in patients with monoclonal gammopathies. Briefly, 100 μl of total BM aspirate was incubated in the dark with anti-human HLA-DR-PE (clone L243; BD), anti-human CD14-PerCP-Cy 5.5, anti-human CD16-PE-Cy7 (clone B73.1; BD) and anti-human CD45-APC-H 7 (clone 2D1; BD) for 20 min. -

Identification of 18 Immune Cell Subsets Using 13-Color Panel
Immunophenotyping Identification of 18 immune cell subsets in human blood using a 13-color panel Background Cell type Function Phenotype Flow cytometry has become the method of choice for Eosinophils Parasitic immunity CD45+, SSCmid/hi, CD14 –, CD16 –, CD19– immunophenotyping and identifying specific cellular + mid/hi subsets. Within seconds, it provides a thorough overview of Neutrophils Innate Immunity CD45 , SSC , CD14 –, CD16+, CD19– the major cell types that constitute a sample. Using multiple + mid markers simultaneously increases the number of parameters Classical Phagocytosis of CD45 , SSC , monocytes pathogens and CD14+, CD16– that can be analyzed per run and decreases the amount of antigen presentation starting material required to perform an assay. This can be Intermediate Phagocytosis of CD45+, SSCmid, critical for precious sample material and long-term immune- monocytes pathogens and CD14+, CD16mid monitoring studies. In this application note, we demonstrate antigen presentation 13-color immunophenotyping of human peripheral blood Non-classical Phagocytosis of CD45+, SSCmid, + + mononuclear cells (PBMCs) using the MACSQuant® Analyzer 16, monocytes pathogens and CD14 , CD16 antigen presentation a compact and reliable benchtop flow cytometer equipped + low + with three lasers. The markers selected allow for the Class-switched Adaptive immunity CD45 , SSC , CD19 , memory B cells CD27+, IgD–, CD14– simultaneous identification and analysis of 18 different cell Non-switched Adaptive immunity CD45+, SSClow, CD19+, populations, thus maximizing the amount of information that memory B cells CD27+, IgD+, CD14– can be retrieved from the sample material analyzed. This is Naive B cells Adaptive immunity – CD45+, SSClow, CD19+, critical when input material is limited, as is often the case for non-antigen CD27–, IgD+, CD14– pediatric or disease studies. -

Combined Carriership of TLR9 -1237C and CD14 -260T Alleles Enhances the Risk of Developing Chronic Relapsing Pouchitis
CORE Metadata, citation and similar papers at core.ac.uk Provided by DSpace at VU PO Box 2345, Beijing 100023, China World J Gastroenterol 2005;11(46):7323-7329 www.wjgnet.com World Journal of Gastroenterology ISSN 1007-9327 [email protected] © 2005 The WJG Press and Elsevier Inc. All rights reserved. ELSEVIER • RAPID COMMUNICATION • Combined carriership of TLR9 -1237C and CD14 -260T alleles enhances the risk of developing chronic relapsing pouchitis KM Lammers, S Ouburg, SA Morré, JBA Crusius, P Gionchetti, F Rizzello, C Morselli, E Caramelli, R Conte, G Poggioli, M Campieri, AS Peña KM Lammers, P Gionchetti, F Rizzello, C Morselli, M CONCLUSION: There is no evidence that the SNPs Campieri, Department of Internal Medicine and Gastroenterology, predispose to the need for IPAA surgery. The signifi cant Policlinico S. Orsola, University of Bologna, Bologna, Italy increase of the combined carriership of the CD14 S Ouburg, SA Morré, JBA Crusius, AS Peña, Laboratory of -260T and TLR9 -1237C alleles in the chronic relapsing Immunogenetics, VU University Medical Center, Amsterdam, The pouchitis group suggests that these markers identify Netherlands a subgroup of IPAA patients with a risk of developing E Caramelli, Institute of Histology and General Embryology, University of Bologna, Bologna, Italy chronic or refractory pouchitis. R Conte, Department of Immunohaematology and Blood Transfusion, Policlinico S. Orsola, University of Bologna, © 2005 The WJG Press and Elsevier Inc. All rights reserved. Bologna, Italy AS Peña, Laboratory of Immunogenetics and Department of Key words: Pouchitis; Innate immunity; Single nucleotide Gastroenterology, VU University Medical Center, Amsterdam, polymorphisms; CD14 ; TLR9 The Netherlands G Poggioli, Department of Surgery and Organ Transplantation, Lammers KM, Ouburg S, Morré SA, Crusius JBA, Gionchetti Policlinic S. -

Single‑Nucleotide Polymorphisms and Copy Number Variations of the FCGR2A and FCGR3A Genes in Healthy Japanese Subjects
BIOMEDICAL REPORTS 2: 265-269, 2014 Single‑nucleotide polymorphisms and copy number variations of the FCGR2A and FCGR3A genes in healthy Japanese subjects HIROYUKI MORIYA1, KATSUHIKO SAITO1,2, NUALA HELSBY3, NAOMI HAYASHI1, SHIGEKAZU SUGINO4, MICHIAKI YAMAKAGE4, TAKERU SAWAGUCHI5, MASAHIKO TAKASAKI5, MASATO TAKAHASHI6 and NAHOKO KUROSAWA1 1Department of Pharmacy, Hokkaido Pharmaceutical University School of Pharmacy, Otaru, Hokkaido 047-0264; 2Department of Pharmacy, Sapporo Hokuyu Hospital, Sapporo, Hokkaido 003-0006, Japan; 3Department of Molecular Medicine and Pathology, Faculty of Medical and Health Sciences, University of Auckland, Auckland 1142, New Zealand; 4Department of Anesthesiology, Sapporo Medical University School of Medicine, Sapporo, Hokkaido 060-8543; Departments of 5Pharmacy and 6Breast Surgery, National Hospital Organization Hokkaido Cancer Center, Sapporo, Hokkaido 003-0804, Japan Received October 24, 2013; Accepted November 18, 2013 DOI: 10.3892/br.2013.210 Abstract. FcγRII and FcγRIII are low-affinity Fcγ recep- previous findings regarding FCGR2A and FCGR3A alleles and tors that are encoded by the FCGR2A and FCGR3A genes, CNVs. These assays may provide a basis for the investigation of respectively. These genes contain functional single-nucleotide the role of these genes in the efficacy of antibody-based drugs, polymorphisms (SNPs), which alter the binding affinities of such as trastuzumab and rituximab, in Japanese subjects. these receptors for the γ chain of the Fc fragment of immu- noglobulin G. The known SNPs in FCGR2A and FCGR3A Introduction are rs1801274 (A>G; H131R) and rs396991 (T>G; F158V), respectively. It is also known that there are copy number varia- Fcγ receptors (FcγRs) bind specifically to the γ chain of the tions (CNVs) in the genetic locus (1q23) where FCGR2A and Fc fragment of immunoglobulin G (IgG) and are located on the FCGR3A are located. -

Integrated Weighted Gene Co-Expression Network Analysis Identi Ed That TLR2 and CD40 Are Related to Coronary Artery Disease
Integrated weighted gene co-expression network analysis identied that TLR2 and CD40 are related to coronary artery disease Bin Qi Liuzhou People's Hospital Jian-Hong Chen Liuzhou People's Hospital Lin Tao Liuzhou People's Hospital Chuan-Meng Zhu Liuzhou People's Hospital Yong Wang Liuzhou People's Hospital Guo-Xiong Deng The First People's of Nanning Liu Miao ( [email protected] ) LiuZhou People's Hospital https://orcid.org/0000-0001-6642-7005 Research article Keywords: Gene Expression Omnibus, Integrated weighted gene co-expression network analysis (WGCNA), Functional enrichment, Functional validation and prognostic analysis. Posted Date: October 7th, 2020 DOI: https://doi.org/10.21203/rs.3.rs-86115/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/16 Abstract Background: The current research attempted to identify possible hub genes and pathways of coronary artery disease (CAD) and to detect the possible mechanisms. Methods: Array data from GSE90074 were downloaded from the Gene Expression Omnibus (GEO) database. Integrated weighted gene co-expression network analysis (WGCNA) was performed to analyze the gene module and clinical characteristics. Gene Ontology annotation, Disease Ontology and the Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses were performed by clusterProler and the DOSE package in R. A protein-protein interaction (PPI) network was established using Cytoscape software, and signicant modules were analyzed using Molecular Complex Detection to identify hub genes. Then, further functional validation of hub genes in other microarrays and population samples was performed, and survival analysis was performed to investigate the prognosis. -

First Infusion Reactions Are Mediated by Fcγriiib and Neutrophils
Pharm Res (2018) 35: 169 https://doi.org/10.1007/s11095-018-2448-8 RESEARCH PAPER First Infusion Reactions are Mediated by FcγRIIIb and Neutrophils Felix Weber 1 & Daniel Breustedt 1,2 & Sonja Schlicht 3 & Claas A. Meyer 3 & Jens Niewoehner 4 & Martin Ebeling 1 & Per-Ola Freskgard5 & Peter Bruenker6 & Thomas Singer1 & Michael Reth7 & Antonio Iglesias1 Received: 29 March 2018 /Accepted: 15 June 2018 /Published online: 27 June 2018 # The Author(s) 2018 ABSTRACT FIR. FcγRIIIb-mediated FIR was abolished by depleting neu- Purpose Administration of therapeutic monoclonal antibod- trophils or blocking FcγRIIIb with CD11b antibodies. ies (mAbs) is frequently accompanied by severe first infusion Conclusions Human FcγRIIIb and neutrophils are primarily reactions (FIR). The mechanism driving FIR is still unclear. responsible for triggering FIR. Clinical strategies to prevent This study aimed to investigate the cellular and molecular FIR in patients should focus on this pathway and may include mechanisms causing FIR in humanized mouse models and transient depletion of neutrophils or blocking FcγRIIIb with their potential for evaluating FIR risk in patients. specific mAbs. Methods Mice humanized for Fc gamma receptors (FcγRs) were generated by recombination-mediated genomic replace- KEY WORDS human FcγRIIIb . humanized mouse model . ment. Body temperature, cytokine release and reactive oxygen immunotoxicology . infusion reactions . neutrophils species (ROS) were measured to assess FIR to mAbs. Results Infusion of human mAb specific for mouse transferrin receptor (HamTfR) into FcγR-humanized mice, produced marked transient hypothermia accompanied by an increase ABBREVIATIONS in inflammatory cytokines KC and MIP-2, and ROS. FIR ADA Anti-drug antibodies were dependent on administration route and Fc-triggered ef- ADCC Antibody-dependent cellular cytotoxicity fector functions mediated by neutrophils. -
Type of the Paper (Article
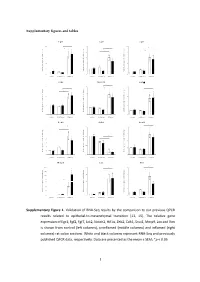
Supplementary figures and tables E g r 1 F g f2 F g f7 1 0 * 5 1 0 * * e e e * g g g * n n n * a a a 8 4 * 8 h h h * c c c d d d * l l l o o o * f f f * n n n o o o 6 3 6 i i i s s s s s s e e e r r r p p p x x x e e e 4 2 4 e e e n n n e e e g g g e e e v v v i i i t t t 2 1 2 a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d J a k 2 N o tc h 2 H if1 * 3 4 6 * * * e e e g g g n n n a a * * a * h h * h c c c 3 * d d * d l l l * o o o f f 2 f 4 n n n o o o i i i s s s s s s e e e r r 2 r p p p x x x e e e e e e n n n e e 1 e 2 g g g e e 1 e v v v i i i t t t a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Z e b 2 C d h 1 S n a i1 * * 7 1 .5 4 * * e e e g g g 6 n n n * a a a * h h h c c c 3 * d d d l l l 5 o o o f f f 1 .0 * n n n * o o o i i i 4 * s s s s s s e e e r r r 2 p p p x x x 3 e e e e e e n n n e e e 0 .5 g g g 2 e e e 1 v v v i i i t t t a a a * l l l e e e 1 * R R R 0 0 .0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d M m p 9 L o x V im 2 0 0 2 0 8 * * * e e e * g g g 1 5 0 * n n n * a a a * h h h * c c c 1 5 * 6 d d d l l l 1 0 0 o o o f f f n n n o o o i i i 5 0 s s s s s s * e e e r r r 1 0 4 3 0 p p p * x x x e e e * e e e n n n e e e 2 0 g g g e e e 5 2 v v v i i i t t t a a a l l l 1 0 e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Supplementary Figure 1. -

Development and Validation of a Protein-Based Risk Score for Cardiovascular Outcomes Among Patients with Stable Coronary Heart Disease
Supplementary Online Content Ganz P, Heidecker B, Hveem K, et al. Development and validation of a protein-based risk score for cardiovascular outcomes among patients with stable coronary heart disease. JAMA. doi: 10.1001/jama.2016.5951 eTable 1. List of 1130 Proteins Measured by Somalogic’s Modified Aptamer-Based Proteomic Assay eTable 2. Coefficients for Weibull Recalibration Model Applied to 9-Protein Model eFigure 1. Median Protein Levels in Derivation and Validation Cohort eTable 3. Coefficients for the Recalibration Model Applied to Refit Framingham eFigure 2. Calibration Plots for the Refit Framingham Model eTable 4. List of 200 Proteins Associated With the Risk of MI, Stroke, Heart Failure, and Death eFigure 3. Hazard Ratios of Lasso Selected Proteins for Primary End Point of MI, Stroke, Heart Failure, and Death eFigure 4. 9-Protein Prognostic Model Hazard Ratios Adjusted for Framingham Variables eFigure 5. 9-Protein Risk Scores by Event Type This supplementary material has been provided by the authors to give readers additional information about their work. Downloaded From: https://jamanetwork.com/ on 10/02/2021 Supplemental Material Table of Contents 1 Study Design and Data Processing ......................................................................................................... 3 2 Table of 1130 Proteins Measured .......................................................................................................... 4 3 Variable Selection and Statistical Modeling ........................................................................................ -

Arming Tumor-Associated Macrophages to Reverse Epithelial
Published OnlineFirst August 15, 2019; DOI: 10.1158/0008-5472.CAN-19-1246 Cancer Tumor Biology and Immunology Research Arming Tumor-Associated Macrophages to Reverse Epithelial Cancer Progression Hiromi I.Wettersten1,2,3, Sara M.Weis1,2,3, Paulina Pathria1,2,Tami Von Schalscha1,2,3, Toshiyuki Minami1,2,3, Judith A. Varner1,2, and David A. Cheresh1,2,3 Abstract Tumor-associated macrophages (TAM) are highly expressed it engaged macrophages but not natural killer (NK) cells to within the tumor microenvironment of a wide range of cancers, induce antibody-dependent cellular cytotoxicity (ADCC) of where they exert a protumor phenotype by promoting tumor avb3-expressing tumor cells despite their expression of the cell growth and suppressing antitumor immune function. CD47 "don't eat me" signal. In contrast to strategies designed Here, we show that TAM accumulation in human and mouse to eliminate TAMs, these findings suggest that anti-avb3 tumors correlates with tumor cell expression of integrin avb3, represents a promising immunotherapeutic approach to redi- a known driver of epithelial cancer progression and drug rect TAMs to serve as tumor killers for late-stage or drug- resistance. A monoclonal antibody targeting avb3 (LM609) resistant cancers. exploited the coenrichment of avb3 and TAMs to not only eradicate highly aggressive drug-resistant human lung and Significance: Therapeutic antibodies are commonly engi- pancreas cancers in mice, but also to prevent the emergence neered to optimize engagement of NK cells as effectors. In of circulating tumor cells. Importantly, this antitumor activity contrast, LM609 targets avb3 to suppress tumor progression in mice was eliminated following macrophage depletion.