Magnesium Deficiency Alters Expression of Genes Critical For
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

ATG7 Promotes Autophagy in Sepsis‑Induced Acute Kidney Injury and Is Inhibited by Mir‑526B
MOLECULAR MEDICINE REPORTS 21: 2193-2201, 2020 ATG7 promotes autophagy in sepsis‑induced acute kidney injury and is inhibited by miR‑526b YING LIU1*, JILAI XIAO1*, JIAKUI SUN1*, WENXIU CHEN1, SHU WANG1, RUN FU1, HAN LIU1 and HONGGUANG BAO2 Departments of 1Critical Care Medicine and 2Anesthesiology, The Affiliated Nanjing First Hospital of Nanjing Medical University, Nanjing, Jiangsu 210006, P.R. China Received July 12, 2019; Accepted January 14, 2020 DOI: 10.3892/mmr.2020.11001 Abstract. Sepsis is considered to be the most common Autophagy is a highly regulated lysosomal intracellular contributing factor in the development of acute kidney degradation pathway involved in removing aggregated protein injury (AKI). However, the mechanisms by which sepsis and maintaining intracellular homeostasis (4,5). Autophagy is leads to AKI remain unclear. Autophagy is important for a associated with several diseases, including kidney disease (6). number of fundamental biological activities and plays a key Autophagy is considered to be a degradation system that occurs role in numerous different diseases. The present study demon- under conditions of stress in order to meet energy and nutrient strated that autophagy is involved in sepsis-induced kidney requirements. Autophagy is very important for a number of injury and upregulates ATG7, LC3 and Beclin I. In addition, fundamental biological activities (4,5). Dysregulation of it was revealed that miR‑526b is decreased in sepsis-induced autophagy has been emphasized in the occurrence of a variety kidney injury, and miR‑526b was identified as a direct regu- of diseases, as the targets of selective autophagy, including key lator of ATG7. Furthermore, the present study investigated organelles, such as mitochondria and lysosomes, are involved the biological effects of ATG7 inhibited by miR‑526b and in diseases (7,8). -

Bioinformatic Analyses of Integral Membrane Transport Proteins Encoded Within the Genome of the Planctomycetes Species, Rhodopirellula Baltica
UC San Diego UC San Diego Previously Published Works Title Bioinformatic analyses of integral membrane transport proteins encoded within the genome of the planctomycetes species, Rhodopirellula baltica. Permalink https://escholarship.org/uc/item/0f85q1z7 Journal Biochimica et biophysica acta, 1838(1 Pt B) ISSN 0006-3002 Authors Paparoditis, Philipp Västermark, Ake Le, Andrew J et al. Publication Date 2014 DOI 10.1016/j.bbamem.2013.08.007 Peer reviewed eScholarship.org Powered by the California Digital Library University of California Biochimica et Biophysica Acta 1838 (2014) 193–215 Contents lists available at ScienceDirect Biochimica et Biophysica Acta journal homepage: www.elsevier.com/locate/bbamem Bioinformatic analyses of integral membrane transport proteins encoded within the genome of the planctomycetes species, Rhodopirellula baltica Philipp Paparoditis a, Åke Västermark a,AndrewJ.Lea, John A. Fuerst b, Milton H. Saier Jr. a,⁎ a Department of Molecular Biology, Division of Biological Sciences, University of California at San Diego, La Jolla, CA, 92093–0116, USA b School of Chemistry and Molecular Biosciences, University of Queensland, Brisbane, Queensland, 9072, Australia article info abstract Article history: Rhodopirellula baltica (R. baltica) is a Planctomycete, known to have intracellular membranes. Because of its un- Received 12 April 2013 usual cell structure and ecological significance, we have conducted comprehensive analyses of its transmembrane Received in revised form 8 August 2013 transport proteins. The complete proteome of R. baltica was screened against the Transporter Classification Data- Accepted 9 August 2013 base (TCDB) to identify recognizable integral membrane transport proteins. 342 proteins were identified with a Available online 19 August 2013 high degree of confidence, and these fell into several different classes. -

Reconstructability Analysis As a Tool for Identifying Gene-Gene Interactions in Studies of Human Diseases
Portland State University PDXScholar Systems Science Faculty Publications and Presentations Systems Science 3-2010 Reconstructability Analysis As A Tool For Identifying Gene-Gene Interactions In Studies Of Human Diseases Stephen Shervais Eastern Washington University Patricia L. Kramer Oregon Health & Science University Shawn K. Westaway Oregon Health & Science University Nancy J. Cox University of Chicago Martin Zwick Portland State University, [email protected] Follow this and additional works at: https://pdxscholar.library.pdx.edu/sysc_fac Part of the Bioinformatics Commons, Diseases Commons, and the Genomics Commons Let us know how access to this document benefits ou.y Citation Details Shervais, S., Kramer, P. L., Westaway, S. K., Cox, N. J., & Zwick, M. (2010). Reconstructability Analysis as a Tool for Identifying Gene-Gene Interactions in Studies of Human Diseases. Statistical Applications In Genetics & Molecular Biology, 9(1), 1-25. This Article is brought to you for free and open access. It has been accepted for inclusion in Systems Science Faculty Publications and Presentations by an authorized administrator of PDXScholar. Please contact us if we can make this document more accessible: [email protected]. Statistical Applications in Genetics and Molecular Biology Volume 9, Issue 1 2010 Article 18 Reconstructability Analysis as a Tool for Identifying Gene-Gene Interactions in Studies of Human Diseases Stephen Shervais∗ Patricia L. Kramery Shawn K. Westawayz Nancy J. Cox∗∗ Martin Zwickyy ∗Eastern Washington University, [email protected] yOregon Health & Science University, [email protected] zOregon Health & Science University, [email protected] ∗∗University of Chicago, [email protected] yyPortland State University, [email protected] Copyright c 2010 The Berkeley Electronic Press. -

Autophagy-Related 7 Modulates Tumor Progression in Triple-Negative Breast Cancer
Laboratory Investigation (2019) 99:1266–1274 https://doi.org/10.1038/s41374-019-0249-2 ARTICLE Autophagy-related 7 modulates tumor progression in triple-negative breast cancer 1 1 1 1 1 2 3 1 Mingyang Li ● Jingwei Liu ● Sihui Li ● Yanling Feng ● Fei Yi ● Liang Wang ● Shi Wei ● Liu Cao Received: 6 November 2018 / Revised: 11 February 2019 / Accepted: 14 February 2019 / Published online: 15 April 2019 © United States & Canadian Academy of Pathology 2019 Abstract The exact role of autophagy in breast cancers remains elusive. In this study, we explored the potential functions of autophagy-related 7 (Atg7) in breast cancer cell lines and tissues. Compared to normal breast tissue, a significantly lower expression of Atg7 was observed in triple-negative breast cancer (TNBC), but not other subtypes. A higher Atg7 expression was significantly associated with favorable clinicopathologic factors and better prognostic outcomes in patients with TNBC. Reflecting the clinical and pathologic observations, Atg7 was found to inhibit proliferation and migration, but promotes apoptosis in TNBC cell lines. Furthermore, Atg7 suppressed epithelial–mesenchymal transition through inhibiting aerobic glycolysis metabolism of TNBC cells. These findings provided novel molecular and clinical evidence of Atg7 in modulating 1234567890();,: 1234567890();,: the biological behavior of TNBC, thus warranting further investigation. Introduction and epidermal growth factor receptor 2 (HER-2), accounts for 15–20% of breast cancers [2]. Cytotoxic chemotherapy Breast cancer is the most commonly diagnosed cancer and remains the mainstay for the treatment of TNBC due to lack the leading cause of cancer death among women worldwide of targeted therapies. Moreover, TNBC is prone to recur- [1]. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Slc1a3-2A-Creert2 Mice Reveal Unique Features of Bergmann Glia and Augment a Growing Collection of Cre Drivers and Effectors In
www.nature.com/scientificreports OPEN Slc1a3‑2A‑CreERT2 mice reveal unique features of Bergmann glia and augment a growing collection of Cre drivers and efectors in the 129S4 genetic background Lech Kaczmarczyk1,2, Nicole Reichenbach2, Nelli Blank2, Maria Jonson1, Lars Dittrich2, Gabor C. Petzold2,3 & Walker S. Jackson1,2* Genetic variation is a primary determinant of phenotypic diversity. In laboratory mice, genetic variation can be a serious experimental confounder, and thus minimized through inbreeding. However, generalizations of results obtained with inbred strains must be made with caution, especially when working with complex phenotypes and disease models. Here we compared behavioral characteristics of C57Bl/6—the strain most widely used in biomedical research—with those of 129S4. In contrast to 129S4, C57Bl/6 demonstrated high within‑strain and intra‑litter behavioral hyperactivity. Although high consistency would be advantageous, the majority of disease models and transgenic tools are in C57Bl/6. We recently established six Cre driver lines and two Cre efector lines in 129S4. To augment this collection, we genetically engineered a Cre line to study astrocytes in 129S4. It was validated with two Cre efector lines: calcium indicator gCaMP5g‑tdTomato and RiboTag—a tool widely used to study cell type‑specifc translatomes. These reporters are in diferent genomic loci, and in both the Cre was functional and astrocyte‑specifc. We found that calcium signals lasted longer and had a higher amplitude in cortical compared to hippocampal astrocytes, genes linked to a single neurodegenerative disease have highly divergent expression patterns, and that ribosome proteins are non‑uniformly expressed across brain regions and cell types. -

Cellular and Molecular Signatures in the Disease Tissue of Early
Cellular and Molecular Signatures in the Disease Tissue of Early Rheumatoid Arthritis Stratify Clinical Response to csDMARD-Therapy and Predict Radiographic Progression Frances Humby1,* Myles Lewis1,* Nandhini Ramamoorthi2, Jason Hackney3, Michael Barnes1, Michele Bombardieri1, Francesca Setiadi2, Stephen Kelly1, Fabiola Bene1, Maria di Cicco1, Sudeh Riahi1, Vidalba Rocher-Ros1, Nora Ng1, Ilias Lazorou1, Rebecca E. Hands1, Desiree van der Heijde4, Robert Landewé5, Annette van der Helm-van Mil4, Alberto Cauli6, Iain B. McInnes7, Christopher D. Buckley8, Ernest Choy9, Peter Taylor10, Michael J. Townsend2 & Costantino Pitzalis1 1Centre for Experimental Medicine and Rheumatology, William Harvey Research Institute, Barts and The London School of Medicine and Dentistry, Queen Mary University of London, Charterhouse Square, London EC1M 6BQ, UK. Departments of 2Biomarker Discovery OMNI, 3Bioinformatics and Computational Biology, Genentech Research and Early Development, South San Francisco, California 94080 USA 4Department of Rheumatology, Leiden University Medical Center, The Netherlands 5Department of Clinical Immunology & Rheumatology, Amsterdam Rheumatology & Immunology Center, Amsterdam, The Netherlands 6Rheumatology Unit, Department of Medical Sciences, Policlinico of the University of Cagliari, Cagliari, Italy 7Institute of Infection, Immunity and Inflammation, University of Glasgow, Glasgow G12 8TA, UK 8Rheumatology Research Group, Institute of Inflammation and Ageing (IIA), University of Birmingham, Birmingham B15 2WB, UK 9Institute of -

Magnesium Is a Key Player in Neuronal Maturation and Neuropathology
International Journal of Molecular Sciences Review Magnesium Is a Key Player in Neuronal Maturation and Neuropathology Ryu Yamanaka 1,2 , Yutaka Shindo 1 and Kotaro Oka 1,3,4,* 1 Center for Biosciences and Informatics, School of Fundamental Science and Technology Graduate School of Science and Technology, Keio University, Yokohama, Kanagawa 223-8522, Japan 2 Faculty of Pharmaceutical Sciences, Sanyo-Onoda City University, Sanyo-Onoda, Yamaguchi 756-0884, Japan 3 Graduate Institute of Medicine, College of Medicine, Kaohsiung Medical University, Kaohsiung City 80708, Taiwan 4 Waseda Research Institute for Science and Engineering, Waseda University, 2-2 Wakamatsucho, Shinjuku, Tokyo 162-8480, Japan * Correspondence: [email protected]; Tel.: +81-045-566-1789 Received: 16 June 2019; Accepted: 9 July 2019; Published: 12 July 2019 Abstract: Magnesium (Mg) is the second most abundant cation in mammalian cells, and it is essential for numerous cellular processes including enzymatic reactions, ion channel functions, metabolic cycles, cellular signaling, and DNA/RNA stabilities. Because of the versatile and universal nature of Mg2+, the homeostasis of intracellular Mg2+ is physiologically linked to growth, proliferation, differentiation, energy metabolism, and death of cells. On the cellular and tissue levels, maintaining Mg2+ within optimal levels according to the biological context, such as cell types, developmental stages, extracellular environments, and pathophysiological conditions, is crucial for development, normal functions, and diseases. Hence, Mg2+ is pathologically involved in cancers, diabetes, and neurodegenerative diseases, such as Parkinson’s disease, Alzheimer’s disease, and demyelination. In the research field regarding the roles and mechanisms of Mg2+ regulation, numerous controversies caused by its versatility and complexity still exist. -
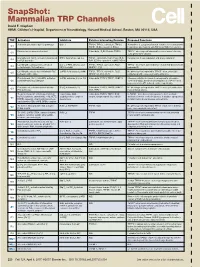
Snapshot: Mammalian TRP Channels David E
SnapShot: Mammalian TRP Channels David E. Clapham HHMI, Children’s Hospital, Department of Neurobiology, Harvard Medical School, Boston, MA 02115, USA TRP Activators Inhibitors Putative Interacting Proteins Proposed Functions Activation potentiated by PLC pathways Gd, La TRPC4, TRPC5, calmodulin, TRPC3, Homodimer is a purported stretch-sensitive ion channel; form C1 TRPP1, IP3Rs, caveolin-1, PMCA heteromeric ion channels with TRPC4 or TRPC5 in neurons -/- Pheromone receptor mechanism? Calmodulin, IP3R3, Enkurin, TRPC6 TRPC2 mice respond abnormally to urine-based olfactory C2 cues; pheromone sensing 2+ Diacylglycerol, [Ca ]I, activation potentiated BTP2, flufenamate, Gd, La TRPC1, calmodulin, PLCβ, PLCγ, IP3R, Potential role in vasoregulation and airway regulation C3 by PLC pathways RyR, SERCA, caveolin-1, αSNAP, NCX1 La (100 µM), calmidazolium, activation [Ca2+] , 2-APB, niflumic acid, TRPC1, TRPC5, calmodulin, PLCβ, TRPC4-/- mice have abnormalities in endothelial-based vessel C4 i potentiated by PLC pathways DIDS, La (mM) NHERF1, IP3R permeability La (100 µM), activation potentiated by PLC 2-APB, flufenamate, La (mM) TRPC1, TRPC4, calmodulin, PLCβ, No phenotype yet reported in TRPC5-/- mice; potentially C5 pathways, nitric oxide NHERF1/2, ZO-1, IP3R regulates growth cones and neurite extension 2+ Diacylglycerol, [Ca ]I, 20-HETE, activation 2-APB, amiloride, Cd, La, Gd Calmodulin, TRPC3, TRPC7, FKBP12 Missense mutation in human focal segmental glomerulo- C6 potentiated by PLC pathways sclerosis (FSGS); abnormal vasoregulation in TRPC6-/- -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

The Association of ATG16L1 Variations with Clinical Phenotypes of Adult-Onset Still’S Disease
G C A T T A C G G C A T genes Article The Association of ATG16L1 Variations with Clinical Phenotypes of Adult-Onset Still’s Disease Wei-Ting Hung 1,2, Shuen-Iu Hung 3 , Yi-Ming Chen 4,5,6 , Chia-Wei Hsieh 6,7, Hsin-Hua Chen 6,7,8 , Kuo-Tung Tang 5,6,7 , Der-Yuan Chen 9,10,11,* and Tsuo-Hung Lan 1,5,12,13,* 1 Institute of Clinical Medicine, National Yang-Ming Chiao Tung University, Taipei 11221, Taiwan; [email protected] 2 Department of Medical Education, Taichung Veterans General Hospital, Taichung 40705, Taiwan 3 Cancer Vaccine and Immune Cell Therapy Core Laboratory, Chang Gung Immunology Consortium, Chang Gung Memorial Hospital, Linkou, Taoyuan 33305, Taiwan; [email protected] 4 Department of Medical Research, Taichung Veterans General Hospital, Taichung 40705, Taiwan; [email protected] 5 School of Medicine, College of Medicine, National Yang Ming Chiao Tung University, Taipei 11221, Taiwan; [email protected] 6 Rong Hsing Research Center for Translational Medicine & Ph.D. Program in Translational Medicine, National Chung Hsing University, Taichung 40227, Taiwan; [email protected] (C.-W.H.); [email protected] (H.-H.C.) 7 Division of Allergy, Immunology, and Rheumatology, Taichung Veterans General Hospital, Taichung 40705, Taiwan 8 Department of Industrial Engineering and Enterprise Information, Tunghai University, Taichung 40705, Taiwan 9 Translational Medicine Laboratory, Rheumatology and Immunology Center, China Medical University Hospital, Taichung 40447, Taiwan 10 Rheumatology and Immunology Center, China Medical University Hospital, Taichung 40447, Taiwan Citation: Hung, W.-T.; Hung, S.-I.; 11 School of Medicine, China Medical University, Taichung 40447, Taiwan Chen, Y.-M.; Hsieh, C.-W.; Chen, 12 Tsao-Tun Psychiatric Center, Ministry of Health and Welfare, Nantou 54249, Taiwan H.-H.; Tang, K.-T.; Chen, D.-Y.; Lan, 13 Center for Neuropsychiatric Research, National Health Research Institutes, Miaoli 35053, Taiwan T.-H. -

Ca Signaling in Cardiac Fibroblasts and Fibrosis-Associated Heart
Journal of Cardiovascular Development and Disease Review Ca2+ Signaling in Cardiac Fibroblasts and Fibrosis-Associated Heart Diseases Jianlin Feng 1, Maria K. Armillei 1, Albert S. Yu 1, Bruce T. Liang 1, Loren W. Runnels 2,* and Lixia Yue 1,* 1 Calhoun Cardiology Center, Department of Cell Biology, University of Connecticut Health Center, Farmington, CT 06030, USA; [email protected] (J.F.); [email protected] (M.K.A.); [email protected] (A.S.Y.); [email protected] (B.T.L.) 2 Department of Pharmacology, Rutgers, Robert Wood Johnson Medical School, Piscataway, NJ 08854, USA * Correspondence: [email protected] (L.W.R.); [email protected] (L.Y.) Received: 11 August 2019; Accepted: 18 September 2019; Published: 23 September 2019 Abstract: Cardiac fibrosis is the excessive deposition of extracellular matrix proteins by cardiac fibroblasts and myofibroblasts, and is a hallmark feature of most heart diseases, including arrhythmia, hypertrophy, and heart failure. This maladaptive process occurs in response to a variety of stimuli, including myocardial injury, inflammation, and mechanical overload. There are multiple signaling pathways and various cell types that influence the fibrogenesis cascade. Fibroblasts and myofibroblasts are central effectors. Although it is clear that Ca2+ signaling plays a vital role in this pathological process, what contributes to Ca2+ signaling in fibroblasts and myofibroblasts is still not wholly understood, chiefly because of the large and diverse number of receptors, transporters, and ion channels that influence intracellular Ca2+ signaling. Intracellular Ca2+ signals are generated by Ca2+ release from intracellular Ca2+ stores and by Ca2+ entry through a multitude of Ca2+-permeable ion channels in the plasma membrane.