Apo and Ligand-Bound High Resolution Cryo-EM Structures of the Human Kv3.1 Reveal a Novel Binding Site for Positive Modulators
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Mineralocorticoid Receptor Leads to Increased Expression of EGFR
www.nature.com/scientificreports OPEN The mineralocorticoid receptor leads to increased expression of EGFR and T‑type calcium channels that support HL‑1 cell hypertrophy Katharina Stroedecke1,2, Sandra Meinel1,2, Fritz Markwardt1, Udo Kloeckner1, Nicole Straetz1, Katja Quarch1, Barbara Schreier1, Michael Kopf1, Michael Gekle1 & Claudia Grossmann1* The EGF receptor (EGFR) has been extensively studied in tumor biology and recently a role in cardiovascular pathophysiology was suggested. The mineralocorticoid receptor (MR) is an important efector of the renin–angiotensin–aldosterone‑system and elicits pathophysiological efects in the cardiovascular system; however, the underlying molecular mechanisms are unclear. Our aim was to investigate the importance of EGFR for MR‑mediated cardiovascular pathophysiology because MR is known to induce EGFR expression. We identifed a SNP within the EGFR promoter that modulates MR‑induced EGFR expression. In RNA‑sequencing and qPCR experiments in heart tissue of EGFR KO and WT mice, changes in EGFR abundance led to diferential expression of cardiac ion channels, especially of the T‑type calcium channel CACNA1H. Accordingly, CACNA1H expression was increased in WT mice after in vivo MR activation by aldosterone but not in respective EGFR KO mice. Aldosterone‑ and EGF‑responsiveness of CACNA1H expression was confrmed in HL‑1 cells by Western blot and by measuring peak current density of T‑type calcium channels. Aldosterone‑induced CACNA1H protein expression could be abrogated by the EGFR inhibitor AG1478. Furthermore, inhibition of T‑type calcium channels with mibefradil or ML218 reduced diameter, volume and BNP levels in HL‑1 cells. In conclusion the MR regulates EGFR and CACNA1H expression, which has an efect on HL‑1 cell diameter, and the extent of this regulation seems to depend on the SNP‑216 (G/T) genotype. -

Table S1 the Four Gene Sets Derived from Gene Expression Profiles of Escs and Differentiated Cells
Table S1 The four gene sets derived from gene expression profiles of ESCs and differentiated cells Uniform High Uniform Low ES Up ES Down EntrezID GeneSymbol EntrezID GeneSymbol EntrezID GeneSymbol EntrezID GeneSymbol 269261 Rpl12 11354 Abpa 68239 Krt42 15132 Hbb-bh1 67891 Rpl4 11537 Cfd 26380 Esrrb 15126 Hba-x 55949 Eef1b2 11698 Ambn 73703 Dppa2 15111 Hand2 18148 Npm1 11730 Ang3 67374 Jam2 65255 Asb4 67427 Rps20 11731 Ang2 22702 Zfp42 17292 Mesp1 15481 Hspa8 11807 Apoa2 58865 Tdh 19737 Rgs5 100041686 LOC100041686 11814 Apoc3 26388 Ifi202b 225518 Prdm6 11983 Atpif1 11945 Atp4b 11614 Nr0b1 20378 Frzb 19241 Tmsb4x 12007 Azgp1 76815 Calcoco2 12767 Cxcr4 20116 Rps8 12044 Bcl2a1a 219132 D14Ertd668e 103889 Hoxb2 20103 Rps5 12047 Bcl2a1d 381411 Gm1967 17701 Msx1 14694 Gnb2l1 12049 Bcl2l10 20899 Stra8 23796 Aplnr 19941 Rpl26 12096 Bglap1 78625 1700061G19Rik 12627 Cfc1 12070 Ngfrap1 12097 Bglap2 21816 Tgm1 12622 Cer1 19989 Rpl7 12267 C3ar1 67405 Nts 21385 Tbx2 19896 Rpl10a 12279 C9 435337 EG435337 56720 Tdo2 20044 Rps14 12391 Cav3 545913 Zscan4d 16869 Lhx1 19175 Psmb6 12409 Cbr2 244448 Triml1 22253 Unc5c 22627 Ywhae 12477 Ctla4 69134 2200001I15Rik 14174 Fgf3 19951 Rpl32 12523 Cd84 66065 Hsd17b14 16542 Kdr 66152 1110020P15Rik 12524 Cd86 81879 Tcfcp2l1 15122 Hba-a1 66489 Rpl35 12640 Cga 17907 Mylpf 15414 Hoxb6 15519 Hsp90aa1 12642 Ch25h 26424 Nr5a2 210530 Leprel1 66483 Rpl36al 12655 Chi3l3 83560 Tex14 12338 Capn6 27370 Rps26 12796 Camp 17450 Morc1 20671 Sox17 66576 Uqcrh 12869 Cox8b 79455 Pdcl2 20613 Snai1 22154 Tubb5 12959 Cryba4 231821 Centa1 17897 -

Potassium Channels in Epilepsy
Downloaded from http://perspectivesinmedicine.cshlp.org/ on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press Potassium Channels in Epilepsy Ru¨diger Ko¨hling and Jakob Wolfart Oscar Langendorff Institute of Physiology, University of Rostock, Rostock 18057, Germany Correspondence: [email protected] This review attempts to give a concise and up-to-date overview on the role of potassium channels in epilepsies. Their role can be defined from a genetic perspective, focusing on variants and de novo mutations identified in genetic studies or animal models with targeted, specific mutations in genes coding for a member of the large potassium channel family. In these genetic studies, a demonstrated functional link to hyperexcitability often remains elusive. However, their role can also be defined from a functional perspective, based on dy- namic, aggravating, or adaptive transcriptional and posttranslational alterations. In these cases, it often remains elusive whether the alteration is causal or merely incidental. With 80 potassium channel types, of which 10% are known to be associated with epilepsies (in humans) or a seizure phenotype (in animals), if genetically mutated, a comprehensive review is a challenging endeavor. This goal may seem all the more ambitious once the data on posttranslational alterations, found both in human tissue from epilepsy patients and in chronic or acute animal models, are included. We therefore summarize the literature, and expand only on key findings, particularly regarding functional alterations found in patient brain tissue and chronic animal models. INTRODUCTION TO POTASSIUM evolutionary appearance of voltage-gated so- CHANNELS dium (Nav)andcalcium (Cav)channels, Kchan- nels are further diversified in relation to their otassium (K) channels are related to epilepsy newer function, namely, keeping neuronal exci- Psyndromes on many different levels, ranging tation within limits (Anderson and Greenberg from direct control of neuronal excitability and 2001; Hille 2001). -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -
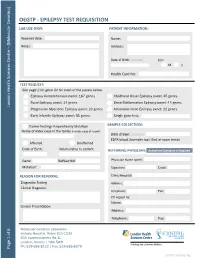
Oegtp - Epilepsy Test Requisition Lab Use Only: Patient Information
OEGTP - EPILEPSY TEST REQUISITION LAB USE ONLY: PATIENT INFORMATION: Received date: Name: Notes: Address: Date of Birth: YY/MM/DD Sex: M F Health Card No: TEST REQUEST: See page 2 for gene list for each of the panels below Epilepsy Comprehensive panel: 167 genes Childhood Onset Epilepsy panel: 45 genes Focal Epilepsy panel: 14 genes Brain Malformation Epilepsy panel: 44 genes London Health Sciences Centre – (Molecular Genetics) London Health Sciences Centre Progressive Myoclonic Epilepsy panel: 20 genes Actionable Gene Epilepsy panel: 22 genes Early Infantile Epilepsy panel: 51 genes Single gene test: Carrier Testing/ KnownFamily Mutation SAMPLE COLLECTION: Name of index case in the family (include copy of report) Date drawn: YY/MM/DD EDTA blood (lavender top) (5ml at room temp) Affected Unaffected Date of Birth: Relationship to patient: REFERRING PHYSICIAN: Authorized Signature is Required Gene: RefSeq:NM Physician Name (print): Mutation: Signature: Email: REASON FOR REFERRAL: Clinic/Hospital: Diagnostic Testing Address: Clinical Diagnosis: Telephone: Fax: CC report to: Name: Clinical Presentation: Address: Telephone: Fax: Molecular Genetics Laboratory Victoria Hospital, Room B10-123A 800 Commissioners Rd. E. London, Ontario | N6A 5W9 Pathology and Laboratory Medicine Ph: 519-685-8122 | Fax: 519-685-8279 Page 1 of 6 Page OEGTP (2021/05/28) OEGTP - EPILEPSY TEST PANELS Patient Identifier: COMPREHENSIVE EPILEPSY PANEL: 167 Genes ACTB, ACTG1, ADSL, AKT3, ALDH7A1, AMT, AP3B2, ARFGEF2, ARHGEF9, ARV1, ARX, ASAH1, ASNS, ATP1A3, ATP6V0A2, ATP7A, -

Transcriptomic Analysis of Native Versus Cultured Human and Mouse Dorsal Root Ganglia Focused on Pharmacological Targets Short
bioRxiv preprint doi: https://doi.org/10.1101/766865; this version posted September 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Transcriptomic analysis of native versus cultured human and mouse dorsal root ganglia focused on pharmacological targets Short title: Comparative transcriptomics of acutely dissected versus cultured DRGs Andi Wangzhou1, Lisa A. McIlvried2, Candler Paige1, Paulino Barragan-Iglesias1, Carolyn A. Guzman1, Gregory Dussor1, Pradipta R. Ray1,#, Robert W. Gereau IV2, # and Theodore J. Price1, # 1The University of Texas at Dallas, School of Behavioral and Brain Sciences and Center for Advanced Pain Studies, 800 W Campbell Rd. Richardson, TX, 75080, USA 2Washington University Pain Center and Department of Anesthesiology, Washington University School of Medicine # corresponding authors [email protected], [email protected] and [email protected] Funding: NIH grants T32DA007261 (LM); NS065926 and NS102161 (TJP); NS106953 and NS042595 (RWG). The authors declare no conflicts of interest Author Contributions Conceived of the Project: PRR, RWG IV and TJP Performed Experiments: AW, LAM, CP, PB-I Supervised Experiments: GD, RWG IV, TJP Analyzed Data: AW, LAM, CP, CAG, PRR Supervised Bioinformatics Analysis: PRR Drew Figures: AW, PRR Wrote and Edited Manuscript: AW, LAM, CP, GD, PRR, RWG IV, TJP All authors approved the final version of the manuscript. 1 bioRxiv preprint doi: https://doi.org/10.1101/766865; this version posted September 12, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Rescue of Motor Coordination by Purkinje Cell-Targeted Restoration of Kv3.3 Channels in Kcnc3-Null Mice Requires Kcnc1
The Journal of Neuroscience, December 16, 2009 • 29(50):15735–15744 • 15735 Cellular/Molecular Rescue of Motor Coordination by Purkinje Cell-Targeted Restoration of Kv3.3 Channels in Kcnc3-Null Mice Requires Kcnc1 Edward C. Hurlock, Mitali Bose, Ganon Pierce, and Rolf H. Joho Department of Neuroscience, The University of Texas Southwestern Medical Center, Dallas, Texas 75390-9111 The role of cerebellar Kv3.1 and Kv3.3 channels in motor coordination was examined with an emphasis on the deep cerebellar nuclei (DCN). Kv3 channel subunits encoded by Kcnc genes are distinguished by rapid activation and deactivation kinetics that support high-frequency, narrow action potential firing. Previously we reported that increased lateral deviation while ambulating and slips while traversing a narrow beam of ataxic Kcnc3-null mice were corrected by restoration of Kv3.3 channels specifically to Purkinje cells, whereas Kcnc3-mutant mice additionally lacking one Kcnc1 allele were partially rescued. Here, we report mice lacking all Kcnc1 and Kcnc3 alleles exhibit no such rescue. For Purkinje cell output to reach the rest of the brain it must be conveyed by neurons of the DCN or vestibular nuclei. As Kcnc1, but not Kcnc3, alleles are lost, mutant mice exhibit increasing gait ataxia accompanied by spike broadening and deceleration in DCN neurons, suggesting the facet of coordination rescued by Purkinje-cell-restricted Kv3.3 restoration in mice lacking just Kcnc3 is hypermetria, while gait ataxia emerges when additionally Kcnc1 alleles are lost. Thus, fast repolarization in Purkinje cells appears important for normal movement velocity, whereas DCN neurons are a prime candidate locus where fast repolarization is necessary for normal gait patterning. -

Global Gene Expression Analysis of Rodent Motor Neurons Following Spinal Cord Injury Associates Molecular Mechanisms with Development of Postinjury Spasticity
J Neurophysiol 103: 761–778, 2010. First published November 25, 2009; doi:10.1152/jn.00609.2009. Global Gene Expression Analysis of Rodent Motor Neurons Following Spinal Cord Injury Associates Molecular Mechanisms With Development of Postinjury Spasticity J. Wienecke,2,* A-C. Westerdahl,1 H. Hultborn,2 O. Kiehn,1 and J. Ryge1,* 1Mammalian Locomotor Laboratory, Department of Neuroscience, Karolinska Institutet, Stockholm, Sweden; and 2Department of Neuroscience and Pharmacology, University of Copenhagen, Copenhagen, Denmark Submitted 15 July 2009; accepted in final form 21 November 2009 Wienecke J, Westerdahl A-C, Hultborn H, Kiehn O, Ryge J. 2004). Significant effort has been devoted to understand the Global gene expression analysis of rodent motor neurons following pathological changes that occur as a response to spinal cord Downloaded from spinal cord injury associates molecular mechanisms with development injury as well as the mechanisms behind the subsequent de- of postinjury spasticity. J Neurophysiol 103: 761–778, 2010. First published November 25, 2009; doi:10.1152/jn.00609.2009. Spinal velopment of spasticity (Hultborn 2003; Little et al. 1999; cord injury leads to severe problems involving impaired motor, Nielsen et al. 2007). Research conducted on humans has sensory, and autonomic functions. After spinal injury there is an initial pointed toward changes in reflex transmission as a mechanism phase of hyporeflexia followed by hyperreflexia, often referred to as for the hyperreflexia (Nielsen et al. 2007; Pierrot-Deseilligny spasticity. Previous studies have suggested a relationship between the and Burke 2005). Animal studies suggest that many factors http://jn.physiology.org/ reappearance of endogenous plateau potentials in motor neurons and contribute to spasticity, including morphological changes such the development of spasticity after spinalization. -

Changes in Excitability and Ion Channel Expression in Neurons of the Major 2 Pelvic Ganglion in Female Type II Diabetic Mice
bioRxiv preprint doi: https://doi.org/10.1101/360826; this version posted July 4, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Changes in excitability and ion channel expression in neurons of the major 2 pelvic ganglion in female type II diabetic mice 3 4 5 Michael Gray1*, Kawasi M. Lett1*, Virginia B. Garcia1, Cindy Kyi1, Kathleen A. Pennington2, Laura C. 6 Schulz2, David J. Schulz1 7 8 1 Division of Biological Sciences, University of Missouri, Columbia, MO USA 65211 9 2 Department of Obstetrics, Gynecology and Women's Health, University of Missouri, Columbia, MO, 10 65211, USA. 11 * Denotes equal contribution by these authors 12 13 14 15 Abbreviated Title 16 Changes in parasympathetic neurons in Type II diabetes 17 18 Corresponding Author 19 David J. Schulz, Ph.D. 20 Division of Biological Sciences 21 University of Missouri-Columbia 22 218 LeFevre Hall 23 Columbia, MO 65211 24 Ph 573-882-4067 25 Fax 573-884-5020 26 Email: [email protected] 27 28 29 Acknowledgments 30 This work was funded by a grant from the Missouri Spinal Cord Injuries Research Program (D.J.S.), the 31 Craig H. Neilsen Foundation (D.J.S.), American Diabetes Association Grant 1-14-BS-181 (L.C.S.) and 32 American Heart Association Postdoctoral Fellowship 13POST16910108 (K.A.P.). The authors declare no 33 competing financial interests. -

Downloaded from the Cancer Tiation but Can Have Multiple Morphological Phenotypes, Since Genome Atlas Database
ONCOLOGY REPORTS 45: 73, 2021 Methylation gene KCNC1 is associated with overall survival in patients with seminoma SAIPENG CHEN1,2*, LONGFEI XIAO1,2*, HUAHONG PENG3, ZHEN WANG4 and JIANBING XIE1,2 1Affiliated Hospital of Putian University, Putian, Fujian 351100;2 Department of Reproductive Medicine, Xiangyang No. 1 People's Hospital, Hubei University of Medicine, Xiangyang, Hubei 441000; 3Tianjin Institute of Urology, The Second Hospital of Tianjin Medical University, Tianjin 300211; 4Taixing People's Hospital, Taixing, Jiangsu 225400, P.R. China Received September 13, 2020; Accepted March 1, 2021 DOI: 10.3892/or.2021.8024 Abstract. The aim of the present study was to explore and and non‑seminoma. The latter can be further subdivided into verify the potential mechanism of seminoma progression. Data embryonic cell carcinoma, choriocarcinoma, yolk sac tumor or on 132 RNA‑seq and 156 methylation sites from stage II/III teratoma (2). Non‑seminomas have a high degree of differen‑ and I seminoma specimens were downloaded from The Cancer tiation but can have multiple morphological phenotypes, since Genome Atlas database. An initial filter of |fold‑change| >2 they contain both stem cells and cells that have differentiated and false discovery rate <0.05 were used to identify differ‑ to varying degrees towards somatic lineages. Seminomas have entially expressed genes (DEGs) which were associated with a lower level of differentiation and resemble primordial germ differential methylation site genes; these genes were consid‑ cells and/or intratubular germ cell neoplasia of unclassified ered potential candidates for further investigation by survival type cells (3‑5). Seminomas can be staged according to the analysis. -

An Advance About the Genetic Causes of Epilepsy
E3S Web of Conferences 271, 03068 (2021) https://doi.org/10.1051/e3sconf/202127103068 ICEPE 2021 An advance about the genetic causes of epilepsy Yu Sun1, a, *, †, Licheng Lu2, b, *, †, Lanxin Li3, c, *, †, Jingbo Wang4, d, *, † 1The School of Molecular and Cellular Biology, University of Illinois at Urbana-Champaign, Urbana, IL 61801-3633, US 2High School Affiliated to Shanghai Jiao Tong University, Shanghai, 200441, China 3Applied Biology program, University of British Columbia, Vancouver, V6r3b1, Canada 4School of Chemical Machinery and Safety, Dalian University of Technology, Dalian, 116023, China †These authors contributed equally. Abstract: Human hereditary epilepsy has been found related to ion channel mutations in voltage-gated channels (Na+, K+, Ca2+, Cl-), ligand gated channels (GABA receptors), and G-protein coupled receptors, such as Mass1. In addition, some transmembrane proteins or receptor genes, including PRRT2 and nAChR, and glucose transporter genes, such as GLUT1 and SLC2A1, are also about the onset of epilepsy. The discovery of these genetic defects has contributed greatly to our understanding of the pathology of epilepsy. This review focuses on introducing and summarizing epilepsy-associated genes and related findings in recent decades, pointing out related mutant genes that need to be further studied in the future. 1 Introduction Epilepsy is a neurological disorder characterized by 2 Malfunction of Ion channel epileptic seizures caused by abnormal brain activity. 1 in Functional variation in voltage or ligand-gated ion 100 (50 million people) people are affected by symptoms channel mutations is a major cause of idiopathic epilepsy, of this disorder worldwide, with men, young children, and especially in rare genetic forms. -

Metastatic Adrenocortical Carcinoma Displays Higher Mutation Rate and Tumor Heterogeneity Than Primary Tumors
ARTICLE DOI: 10.1038/s41467-018-06366-z OPEN Metastatic adrenocortical carcinoma displays higher mutation rate and tumor heterogeneity than primary tumors Sudheer Kumar Gara1, Justin Lack2, Lisa Zhang1, Emerson Harris1, Margaret Cam2 & Electron Kebebew1,3 Adrenocortical cancer (ACC) is a rare cancer with poor prognosis and high mortality due to metastatic disease. All reported genetic alterations have been in primary ACC, and it is 1234567890():,; unknown if there is molecular heterogeneity in ACC. Here, we report the genetic changes associated with metastatic ACC compared to primary ACCs and tumor heterogeneity. We performed whole-exome sequencing of 33 metastatic tumors. The overall mutation rate (per megabase) in metastatic tumors was 2.8-fold higher than primary ACC tumor samples. We found tumor heterogeneity among different metastatic sites in ACC and discovered recurrent mutations in several novel genes. We observed 37–57% overlap in genes that are mutated among different metastatic sites within the same patient. We also identified new therapeutic targets in recurrent and metastatic ACC not previously described in primary ACCs. 1 Endocrine Oncology Branch, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892, USA. 2 Center for Cancer Research, Collaborative Bioinformatics Resource, National Cancer Institute, National Institutes of Health, Bethesda, MD 20892, USA. 3 Department of Surgery and Stanford Cancer Institute, Stanford University, Stanford, CA 94305, USA. Correspondence and requests for materials should be addressed to E.K. (email: [email protected]) NATURE COMMUNICATIONS | (2018) 9:4172 | DOI: 10.1038/s41467-018-06366-z | www.nature.com/naturecommunications 1 ARTICLE NATURE COMMUNICATIONS | DOI: 10.1038/s41467-018-06366-z drenocortical carcinoma (ACC) is a rare malignancy with types including primary ACC from the TCGA to understand our A0.7–2 cases per million per year1,2.