Amino Acid-Based Polymeric Scaffold Fabrication And
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Hydrogels and Dentin–Pulp Complex Regeneration: from the Benchtop to Clinical Translation
polymers Review Hydrogels and Dentin–Pulp Complex Regeneration: From the Benchtop to Clinical Translation Marwa M. S. Abbass 1,2 , Aiah A. El-Rashidy 2,3 , Khadiga M. Sadek 2,3 , Sara El Moshy 1,2, Israa Ahmed Radwan 1,2 , Dina Rady 1,2 , Christof E. Dörfer 4 and Karim M. Fawzy El-Sayed 2,4,5,* 1 Oral Biology Department, Faculty of Dentistry, Cairo University, Cairo 11562, Egypt; [email protected] (M.M.S.A.); [email protected] (S.E.M.); [email protected] (I.A.R.); [email protected] (D.R.) 2 Stem Cells and Tissue Engineering Research Group, Faculty of Dentistry, Cairo University, Cairo 11562, Egypt; [email protected] (A.A.E.-R.); [email protected] (K.M.S.) 3 Biomaterials Department, Faculty of Dentistry, Cairo University, Cairo 11562, Egypt 4 Clinic for Conservative Dentistry and Periodontology, School of Dental Medicine, Christian Albrechts University, 24105 Kiel, Germany; [email protected] 5 Oral Medicine and Periodontology Department, Faculty of Dentistry, Cairo University, Cairo 11562, Egypt * Correspondence: [email protected] Received: 22 October 2020; Accepted: 10 November 2020; Published: 9 December 2020 Abstract: Dentin–pulp complex is a term which refers to the dental pulp (DP) surrounded by dentin along its peripheries. Dentin and dental pulp are highly specialized tissues, which can be affected by various insults, primarily by dental caries. Regeneration of the dentin–pulp complex is of paramount importance to regain tooth vitality. The regenerative endodontic procedure (REP) is a relatively current approach, which aims to regenerate the dentin–pulp complex through stimulating the differentiation of resident or transplanted stem/progenitor cells. -

Barriers to the Identification of Occupational Asthma
BARRIERS TO THE IDENTIFICATION OF OCCUPATIONAL ASTHMA By Gareth Iestyn Walters A thesis submitted to The University of Birmingham for the degree of Doctor of Medicine School of Health and Populations Sciences College of Medical and Dental Sciences The University of Birmingham August 2014 DEDICATION This is for my father Derek Walters (1940-2011)… who would have kept a copy on the bookshelf, but probably wouldn’t have read it unless it was about Frederick the Great or the Stuart monarchy… which unfortunately it isn’t. ABSTRACT Occupational asthma accounts for 1 in 6 cases of new-onset adult asthma and is associated with an estimated societal cost in the UK of £100 million per annum. The cost is somewhat avoidable if workers with occupational asthma are identified quickly and removed from exposure to a sensitizing agent. However many workers with occupational asthma go undiagnosed or experience a lengthy delay in diagnosis. The aim of this work was to identify the barriers to diagnosis of occupational asthma on the part of the worker and of the healthcare professional. The first study evaluated current practice in assessing working-age asthmatics for occupational asthma in a West Midlands primary care population, using UK national guidelines as a reference standard. The recorded prevalence of occupational asthma was much lower than expected (0-0.8%) and there was poor enquiry regarding occupation (14% of cases) and the effect of work on asthma symptoms (2% of cases) by primary healthcare professionals. The second study used a qualitative methodology to explore and define health beliefs and behaviours in workers with occupational asthma symptoms. -

Ncounter Human Inflammation V2 Panel Gene List
nCounter Human Inflammation V2 Panel Gene List Official Symbol Accession Alias / Prev Symbol GO Annotation* Official Full Name Intrinsic To Plasma Membrane,Intrinsic To Membrane,Membrane Part,Membrane,Integral To Membrane,Integral To Plasma Membrane,Plasma Membrane Part,Plasma Membrane,Signal Transduction,Cell Surface Receptor Linked Signal Transduction Go 0007166,Defense Response,Inflammatory Response,Response To Stress,Response To External AGER NM_001136.3 RAGE Stimulus,Response To W advanced glycosylation end product-specific receptor Cytoplasmic Part,Membrane,Cytoplasm,Cytosol,Plasma Membrane,Regulation Of Biological Quality,Positive Regulation Of Cell Proliferation,Cell Development,Negative Regulation Of Apoptosis,Programmed Cell Death,Regulation Of Growth,Carboxylic Acid Metabolic ALOX12 NM_000697.1 12-LOX, 12S-LOX, LOG12 Process,Fatty Acid Oxidation,Negative Regulation Of Cellular Process,Regulation Of Cell arachidonate 12-lipoxygenase Defense Response,Inflammatory Response,Response To Stress,Response To External ALOX15 NM_001140.3 15-LOX-1, 15LOX-1 Stimulus,Response To Wounding,Oxidoreductase Activity arachidonate 15-lipoxygenase ALOX5 NM_000698.2 5-LO, 5-LOX, 5LPG, LOG5 Oxidoreductase Activity arachidonate 5-lipoxygenase Extracellular Region,Extracellular Region Part,Extracellular Space,Cell Cell Signaling,Cell AREG NM_001657.2 AR, CRDGF, SDGF Proliferation Go 0008283,Receptor Binding,Growth Factor Activity amphiregulin Cytoplasm,Carboxylic Acid Metabolic Process,Glutamine Family Amino Acid Metabolic Process,Amino Acid Metabolic -

An Investigation of the Properties of Mixtures of Starches with Mono-Myristic Acid Triglyceride As Potential Selectively Digestible Films
An Investigation of the Properties of Mixtures of Starches with Mono-Myristic Acid Triglyceride as Potential Selectively Digestible Films. A dissertation submitted for the degree of Doctor of Philosophy by Christiane Charlotte Fertig at The School of Pharmacy Faculty of Medicine University of London October 2002 ProQuest Number: 10104897 All rights reserved INFORMATION TO ALL USERS The quality of this reproduction is dependent upon the quality of the copy submitted. In the unlikely event that the author did not send a complete manuscript and there are missing pages, these will be noted. Also, if material had to be removed, a note will indicate the deletion. uest. ProQuest 10104897 Published by ProQuest LLC(2016). Copyright of the Dissertation is held by the Author. All rights reserved. This work is protected against unauthorized copying under Title 17, United States Code. Microform Edition © ProQuest LLC. ProQuest LLC 789 East Eisenhower Parkway P.O. Box 1346 Ann Arbor, Ml 48106-1346 Abstract Abstract The aim of this work was the production of melt-films based on a mixture of unprocessed starch and a mono-myristic acid triglyceride for use as a coating on colon delivery formulations. The starches were embedded in the triglyceride-matrix during a melting process and drug release across the films was obtained by the digestion of the starch fraction through colonic a-amylases. During the first part of this study, the physicochemical properties of 7 starches, whose amylose content ranged from 0 % to nearly 100 %, were investigated and sub sequently correlated by statistical analysis. The second part comprised the identification of the optimum film formula with respect to film-forming temperature, starch content and film thickness. -

Automotive Manufacturers' Cost/Revenue, Financial
- HP ' "434 n no. DOT-TSC-flHTSA-79-21 HS 803-657 M'O. I OOT- TSC- jJ I VHTS4 AUTOMOTIVE MANUFACTURERS' 79-21 COST/REVENUE. FINANCIAL AND RISK ANALYSIS: PROJECTED IMPACT OF AUTOMOBILE MANUFACTURING ON THE PLASTICS INDUSTRY Robert Kaiser H.H. Aerospace Design Company Inc. Civil Air Terminal Bedford MA 01370 of TR4a, AUGUST 1979 FINAL REPORT DOCUMENT IS AVAILABLE TO THE U.S. PUBLIC THROUGH THE NATIONAL TECHNICAL INFORMATION SERVICE, SPRINGFIELD, VIRGINIA 22161 Prepared for y.S. DEPARTMENT OF TRANSPORTATION, Nation^^ Highway Traffic Safety Administration Office of Research and Development Washington DC 20590 , , f Technical Report Documentation Page 1 . Report No. 2. Government Accession No. 3. Recipient's Cotolog No. HS-803-657 4. Titl« ond Subtitle 5. Report Date ('AUTOMOTIVE MANUFACTURERS' COST/REVENUE, August 1979 . FINANCIAL AND RISK ANALYSIS: PROJECTED IMPACT OF 6. Performing Orgonizotion Code AUTOMOBILE MANUFACTURING ON THE PLASTICS INDUSTRY 8. Performing Orgonizotion Report No. 7. Author^ s) Robert. Kaiser DOT-TSC-NHTSA-79-21 9. Performing Orgonizotion Name and Address 10. Work Unit No. (TRAIS) HS H.H, Aerospace Company, Inc.* 927/R9404 civil Air Terminal _ 11. Controct or Gront No. Bedford, MA 01736 DOT-TSC-1333 13. Type of Report ond Period Covered 12. Sponsoring Agency Nome and Address U.S. Department of Transportation Final Report National Highway Traffic Safety Administration June 1977 to July 1978 Office of Research and Development 14. Sponsoring Agency Code Washington DC 20590 15. Sopplementory Mores Department of Transportation “Under contract to: Research and Special Programs Administration Transportation Systems Center Kendall Square, Cambridge MA 02142 16. -

Targeting Undruggable Oncoprotein Epitopes with Protein Catalyzed Capture Agents
Targeting Undruggable Oncoprotein Epitopes with Protein Catalyzed Capture Agents Thesis by Ryan Kenneth Henning In Partial Fulfillment of the Requirements for the degree of Doctor of Philosophy CALIFORNIA INSTITUTE OF TECHNOLOGY Pasadena, California 2017 (Defended June 13, 2016) ii 2016 Ryan Kenneth Henning ORCID: 0000-0002-3783-2455 iii ACKNOWLEDGEMENTS I would like to start by thanking my advisor, Prof. Jim Heath, for taking me into his group. I am extremely grateful for the opportunities I had to work on such amazing projects throughout my graduate career. Jim, you are phenomenal at identifying interesting problems and addressing them in new and creative ways. It is truly inspiring to have been in such a vibrant scientific atmosphere and I am certain that I am leaving as better scientist than when I arrived. And of course thanks for being such a great provider! I must also thank the members of my thesis committee. I could not have asked for a better committee chair than Prof. David Tirrell. You were always extremely thoughtful and supportive during my committee meetings. I also thank you for the encouragement and support when I attempted to use BONCAT early on in my project. That taught me not to shy away from trying to combine different technologies or do new things. I thank Prof. Ray Deshaies for teaching me about PROTACs. I still think that is one of the coolest concepts in science today and it wouldn’t exist if not for you. I was also inspired during all of our discussions to see how much you truly love science. -

Preparation of Poly Alpha-1,3-Glucan Esters
(19) *EP003317304B1* (11) EP 3 317 304 B1 (12) EUROPEAN PATENT SPECIFICATION (45) Date of publication and mention (51) Int Cl.: of the grant of the patent: C08B 37/00 (2006.01) 03.06.2020 Bulletin 2020/23 (86) International application number: (21) Application number: 16734840.8 PCT/US2016/038909 (22) Date of filing: 23.06.2016 (87) International publication number: WO 2017/003808 (05.01.2017 Gazette 2017/01) (54) PREPARATION OF POLY ALPHA-1,3-GLUCAN ESTERS USING CYCLIC ORGANIC ANHYDRIDES HERSTELLUNG VON POLY-ALPHA-1,3-GLUCAN-ESTERN UNTER VERWENDUNG VON CYCLISCHEN ORGANISCHEN ANHYDRIDEN PRÉPARATION DE POLYALPHA-1,3-GLUCANE-ESTERS À L’AIDE D’ANHYDRIDES ORGANIQUES CYCLIQUES (84) Designated Contracting States: • LENGES, Christian Peter AL AT BE BG CH CY CZ DE DK EE ES FI FR GB Wilmington, Delaware 19810 (US) GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO PL PT RO RS SE SI SK SM TR (74) Representative: Dehns St. Bride’s House (30) Priority: 30.06.2015 US 201562186570 P 10 Salisbury Square London EC4Y 8JD (GB) (43) Date of publication of application: 09.05.2018 Bulletin 2018/19 (56) References cited: US-A1- 2014 187 766 (73) Proprietor: DuPont Industrial Biosciences USA, LLC • NASROLLAHI ZAHRA ET AL: "Functionalized Wilmington, Delaware 19805 (US) nanoscale [beta]-1,3-glucan to improve Her2+ breast cancer therapy: In vitro and in vivo study", (72) Inventors: JOURNAL OF CONTROLLED RELEASE, vol. 202, • PAULLIN, Jayme L. 15 January 2015 (2015-01-15), pages 49-56, Claymont, Delaware 19703 (US) XP029219872, ISSN: 0168-3659, DOI: • NAMBIAR, Rakesh 10.1016/J.JCONREL.2015.01.014 West Chester, Pennsylvania 19382 (US) Note: Within nine months of the publication of the mention of the grant of the European patent in the European Patent Bulletin, any person may give notice to the European Patent Office of opposition to that patent, in accordance with the Implementing Regulations. -

New Mesh Headings for 2018 Listed in Alphabetical Order with Heading, Scope Note, Annotation (AN), and Tree Locations
New MeSH Headings for 2018 Listed in alphabetical order with Heading, Scope Note, Annotation (AN), and Tree Locations 2-Hydroxypropyl-beta-cyclodextrin AAA Domain Derivative of beta-cyclodextrin that is used as an excipient for An approximately 250 amino acid domain common to AAA steroid drugs and as a lipid chelator. ATPases and AAA Proteins. It consists of a highly conserved N- terminal P-Loop ATPase subdomain with an alpha-beta-alpha Tree locations: conformation, and a less-conserved C-terminal subdomain with an beta-Cyclodextrins D04.345.103.333.500 all alpha conformation. The N-terminal subdomain includes Walker D09.301.915.400.375.333.500 A and Walker B motifs which function in ATP binding and D09.698.365.855.400.375.333.500 hydrolysis. Tree locations: Amino Acid Motifs G02.111.570.820.709.275.500.913 AAA Proteins Abuse-Deterrent Formulations A large, highly conserved and functionally diverse superfamily of Drug formulations or delivery systems intended to discourage the NTPases and nucleotide-binding proteins that are characterized by a abuse of CONTROLLED SUBSTANCES. These may include conserved 200 to 250 amino acid nucleotide-binding and catalytic physical barriers to prevent chewing or crushing the drug; chemical domain, the AAA+ module. They assemble into hexameric ring barriers that prevent extraction of psychoactive ingredients; agonist- complexes that function in the energy-dependent remodeling of antagonist combinations to reduce euphoria associated with abuse; macromolecules. Members include ATPASES ASSOCIATED aversion, where controlled substances are combined with others that WITH DIVERSE CELLULAR ACTIVITIES. will produce an unpleasant effect if the patient manipulates the dosage form or exceeds the recommended dose; delivery systems Tree locations: that are resistant to abuse such as implants; or combinations of these Acid Anhydride Hydrolases D08.811.277.040.013 methods. -
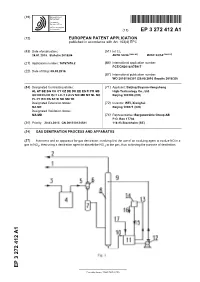
Gas Denitration Process and Apparatus
(19) TZZ¥ _ _T (11) EP 3 272 412 A1 (12) EUROPEAN PATENT APPLICATION published in accordance with Art. 153(4) EPC (43) Date of publication: (51) Int Cl.: 24.01.2018 Bulletin 2018/04 B01D 53/56 (2006.01) B01D 53/14 (2006.01) (21) Application number: 16767678.2 (86) International application number: PCT/CN2016/075917 (22) Date of filing: 09.03.2016 (87) International publication number: WO 2016/150301 (29.09.2016 Gazette 2016/39) (84) Designated Contracting States: (71) Applicant: Beijing Boyuan-Hengsheng AL AT BE BG CH CY CZ DE DK EE ES FI FR GB High-Technology Co., Ltd. GR HR HU IE IS IT LI LT LU LV MC MK MT NL NO Beijing 100190 (CN) PL PT RO RS SE SI SK SM TR Designated Extension States: (72) Inventor: WEI, Xionghui BA ME Beijing 100871 (CN) Designated Validation States: MA MD (74) Representative: Bergenstråhle Group AB P.O. Box 17704 (30) Priority: 20.03.2015 CN 201510124581 118 93 Stockholm (SE) (54) GAS DENITRATION PROCESS AND APPARATUS (57) A process and an apparatus for gas denitration, involving first the use of an oxidizing agent to oxidize NO in a gas to NO2, then using a denitration agent to absorb the NO 2 in the gas, thus achieving the purpose of denitration. EP 3 272 412 A1 Printed by Jouve, 75001 PARIS (FR) EP 3 272 412 A1 Description Technical Field 5 [0001] The present invention relates to the industrial field of gas denitration, and particularly to a process and an apparatus for the denitration of flue gases or various combustion exhaust (waste) gases. -

WO 2015/084763 A2 11 June 2015 (11.06.2015) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date WO 2015/084763 A2 11 June 2015 (11.06.2015) P O P C T (51) International Patent Classification: (74) Common Representative: MERCK SHARP & DOHME C07D 401/06 (2006.01) CORP.; 126 East Lincoln Avenue, Rahway, New Jersey 07065-0907 (US). (21) International Application Number: PCT/US20 14/068008 (81) Designated States (unless otherwise indicated, for every kind of national protection available): AE, AG, AL, AM, (22) International Filing Date: AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, 2 December 2014 (02.12.2014) BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, (25) Filing Language: English DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IR, IS, JP, KE, KG, KN, KP, KR, (26) Publication Language: English KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, MG, (30) Priority Data: MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, 61/91 1,686 4 December 2013 (04. 12.2013) PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, SC, 62/017,542 26 June 2014 (26.06.2014) SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (71) Applicant: MERCK SHARP & DOHME CORP. [US/US]; 126 East Lincoln Avenue, Rahway, New Jersey (84) Designated States (unless otherwise indicated, for every 07065-0907 (US). -

Understanding Physiology of Diseases and Cell Lines Using Omics Based Approaches
UNDERSTANDING PHYSIOLOGY OF DISEASES AND CELL LINES USING OMICS BASED APPROACHES by Amit Kumar A dissertation submitted to Johns Hopkins University in conformity with the requirements for the degree of Doctor of Philosophy Baltimore, Maryland May, 2015 © 2015 Amit Kumar All Rights Reserved Abstract This thesis focuses on understanding physiology of diseases and cell lines using OMICS based approaches such as microarrays based gene expression analysis and mass spectrometry based proteins analysis. It includes extensive work on functionally characterizing mass spectrometry based proteomics data for identifying secreted proteins using bioinformatics tools. This dissertation also includes work on using omics based techniques coupled with bioinformatics tools to elucidate pathophysiology of diseases such as Type 2 Diabetes (T2D). Although the well-known characteristic of T2D is hyperglycemia, there are multiple other metabolic abnormalities that occur in T2D, including insulin resistance and dyslipidemia. In order to attain a greater understanding of the alterations in metabolic tissues associated with T2D, microarray analysis of gene expression in metabolic tissues from a mouse model of pre-diabetes and T2D to understand the metabolic abnormalities that may contribute to T2D was performed. This study also uncovered the novel genes and pathways regulated by the insulin sensitizing agent (CL-316,243) to identify key pathways and target genes in metabolic tissues that can reverse the diabetic phenotype. Specifically, he found significant decreases in the expression of mitochondrial and peroxisomal fatty acid oxidation genes in the skeletal muscle and adipose tissue of adult MKR mice, and in the liver of pre-diabetic MKR mice, compared to healthy mice. In addition, this study also explained the lower free fatty acid levels in MKR mice after treatment with CL-316,243 and provided biomarker genes such as ACAA1 and HSD17b4. -
WO 2017/003808 Al 5 January 2017 (05.01.2017) P O P C T
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization I International Bureau (10) International Publication Number (43) International Publication Date WO 2017/003808 Al 5 January 2017 (05.01.2017) P O P C T (51) International Patent Classification: (81) Designated States (unless otherwise indicated, for every C 37/00 (2006.01) kind of national protection available): AE, AG, AL, AM, AO, AT, AU, AZ, BA, BB, BG, BH, BN, BR, BW, BY, (21) International Application Number: BZ, CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, PCT/US20 16/038909 DO, DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, (22) International Filing Date: HN, HR, HU, ID, IL, IN, IR, IS, JP, KE, KG, KN, KP, KR, 23 June 2016 (23.06.2016) KZ, LA, LC, LK, LR, LS, LU, LY, MA, MD, ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, NO, NZ, OM, (25) Filing Language: English PA, PE, PG, PH, PL, PT, QA, RO, RS, RU, RW, SA, SC, (26) Publication Language: English SD, SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (30) Priority Data: 62/186,570 30 June 2015 (30.06.2015) US (84) Designated States (unless otherwise indicated, for every kind of regional protection available): ARIPO (BW, GH, (71) Applicant: E. I. DU PONT DE NEMOURS AND COM¬ GM, KE, LR, LS, MW, MZ, NA, RW, SD, SL, ST, SZ, PANY [US/US]; Chestnut Run Plaza, 974 Centre Road, TZ, UG, ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, RU, P.O.