Expression Profiling of Macrophages Reveals Multiple Populations With
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Serum Scd163 As a Biomarker of Adipose Tissue Inflammation in Obstructive Sleep Apnoea Patients: Limits and Perspectives
AGORA | CORRESPONDENCE Serum sCD163 as a biomarker of adipose tissue inflammation in obstructive sleep apnoea patients: limits and perspectives To the Editor: Recently, MURPHY et al. [1] demonstrated, through a challenging animal, cellular and human translational approach, that intermittent hypoxia induces a pro-inflammatory activation of adipose tissue macrophages (ATMs), promoting insulin resistance. They confirmed that intermittent hypoxia lowers insulin-mediated glucose uptake in 3T3-L1 adipocytes, and evidenced the interrelationship between inflammation and insulin resistance in the adipose tissue of mice exposed to intermittent hypoxia. They then measured the concentration of serum soluble CD163 (sCD163), an assumed pro-inflammatory biomarker reflecting macrophage activation, in obstructive sleep apnoea (OSA) patients classified according to their BMI and apnoea/hypopnoea index (AHI). This is the first time serum sCD163 has been evaluated in OSA patients, and its association with AHI is promising. With the aim of enhancing its relevance in further studies, we wish to discuss the following points. 1) Activation of adipose tissue macrophages (ATMs) towards M1 phenotype in OSA patients. MURPHY et al. [1] made the assumption that circulating sCD163 levels in OSA patients reflect the intermittent hypoxia-dependent polarisation of ATMs towards a M1 phenotype. As appropriately cited, KRAČMEROVÁ et al. [2] showed that circulating sCD163 concentrations are associated with both macrophage content in human adipose tissue, and insulin sensitivity. However, the assumption that serum sCD163 reflects such a polarisation based on the ADAM-17-dependent cleavage, as hypothesised by ETZERODT et al. [3], is unconvincing since only shown in HEK293 cells. Human ATMs are characterised by a high expression of CD163 which, in contrast, is weakly expressed in differentiated M1 macrophages [4]. -

ENSG Gene Encodes Effector TCR Pathway Costimulation Inhibitory/Exhaustion Synapse/Adhesion Chemokines/Receptors
ENSG Gene Encodes Effector TCR pathway Costimulation Inhibitory/exhaustion Synapse/adhesion Chemokines/receptors ENSG00000111537 IFNG IFNg x ENSG00000109471 IL2 IL-2 x ENSG00000232810 TNF TNFa x ENSG00000271503 CCL5 CCL5 x x ENSG00000139187 KLRG1 Klrg1 x ENSG00000117560 FASLG Fas ligand x ENSG00000121858 TNFSF10 TRAIL x ENSG00000134545 KLRC1 Klrc1 / NKG2A x ENSG00000213809 KLRK1 Klrk1 / NKG2D x ENSG00000188389 PDCD1 PD-1 x x ENSG00000117281 CD160 CD160 x x ENSG00000134460 IL2RA IL-2 receptor x subunit alpha ENSG00000110324 IL10RA IL-10 receptor x subunit alpha ENSG00000115604 IL18R1 IL-18 receptor 1 x ENSG00000115607 IL18RAP IL-18 receptor x accessory protein ENSG00000081985 IL12RB2 IL-12 receptor x beta 2 ENSG00000186810 CXCR3 CXCR3 x x ENSG00000005844 ITGAL CD11a x ENSG00000160255 ITGB2 CD18; Integrin x x beta-2 ENSG00000156886 ITGAD CD11d x ENSG00000140678 ITGAX; CD11c x x Integrin alpha-X ENSG00000115232 ITGA4 CD49d; Integrin x x alpha-4 ENSG00000169896 ITGAM CD11b; Integrin x x alpha-M ENSG00000138378 STAT4 Stat4 x ENSG00000115415 STAT1 Stat1 x ENSG00000170581 STAT2 Stat2 x ENSG00000126561 STAT5a Stat5a x ENSG00000162434 JAK1 Jak1 x ENSG00000100453 GZMB Granzyme B x ENSG00000145649 GZMA Granzyme A x ENSG00000180644 PRF1 Perforin 1 x ENSG00000115523 GNLY Granulysin x ENSG00000100450 GZMH Granzyme H x ENSG00000113088 GZMK Granzyme K x ENSG00000057657 PRDM1 Blimp-1 x ENSG00000073861 TBX21 T-bet x ENSG00000115738 ID2 ID2 x ENSG00000176083 ZNF683 Hobit x ENSG00000137265 IRF4 Interferon x regulatory factor 4 ENSG00000140968 IRF8 Interferon -

Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse
Welcome to More Choice CD Marker Handbook For more information, please visit: Human bdbiosciences.com/eu/go/humancdmarkers Mouse bdbiosciences.com/eu/go/mousecdmarkers Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse CD3 CD3 CD (cluster of differentiation) molecules are cell surface markers T Cell CD4 CD4 useful for the identification and characterization of leukocytes. The CD CD8 CD8 nomenclature was developed and is maintained through the HLDA (Human Leukocyte Differentiation Antigens) workshop started in 1982. CD45R/B220 CD19 CD19 The goal is to provide standardization of monoclonal antibodies to B Cell CD20 CD22 (B cell activation marker) human antigens across laboratories. To characterize or “workshop” the antibodies, multiple laboratories carry out blind analyses of antibodies. These results independently validate antibody specificity. CD11c CD11c Dendritic Cell CD123 CD123 While the CD nomenclature has been developed for use with human antigens, it is applied to corresponding mouse antigens as well as antigens from other species. However, the mouse and other species NK Cell CD56 CD335 (NKp46) antibodies are not tested by HLDA. Human CD markers were reviewed by the HLDA. New CD markers Stem Cell/ CD34 CD34 were established at the HLDA9 meeting held in Barcelona in 2010. For Precursor hematopoetic stem cell only hematopoetic stem cell only additional information and CD markers please visit www.hcdm.org. Macrophage/ CD14 CD11b/ Mac-1 Monocyte CD33 Ly-71 (F4/80) CD66b Granulocyte CD66b Gr-1/Ly6G Ly6C CD41 CD41 CD61 (Integrin b3) CD61 Platelet CD9 CD62 CD62P (activated platelets) CD235a CD235a Erythrocyte Ter-119 CD146 MECA-32 CD106 CD146 Endothelial Cell CD31 CD62E (activated endothelial cells) Epithelial Cell CD236 CD326 (EPCAM1) For Research Use Only. -

Basic Helix-Loop-Helix Transcription Factors DEC1 and DEC2 Regulate the Paclitaxel-Induced Apoptotic Pathway of MCF-7 Human Breast Cancer Cells
INTERNATIONAL JOURNAL OF MoleCular MEDICine 27: 491-495, 2011 Basic helix-loop-helix transcription factors DEC1 and DEC2 regulate the paclitaxel-induced apoptotic pathway of MCF-7 human breast cancer cells YUNYAN WU1,2, FUYUKI SATO1, UJJAL KUMAR BHAWAL3, TAKESHI KAWAMOTO4, KATSUMI FUJIMOTO4, MITSUHIDE NOSHIRO4, SATOKO MOROHASHI1, YUKIO KATO4 and HIROSHI KIJIMA1 1Department of Pathology and Bioscience, Hirosaki University Graduate School of Medicine, Hirosaki 036-8562, Japan; 2Department of Pathology, College of Basic Medical Sciences, China Medical University, Shenyang 110001, P.R. China; 3Department of Oral and Maxillofacial Surgery and High-Tech Research Center, Kanagawa Dental College, Yokosuka 238-8580; 4Department of Dental and Medical Biochemistry, Hiroshima University Graduate School of Biomedical Science, Hiroshima 734-8553, Japan Received November 17, 2010; Accepted January 10, 2011 DOI: 10.3892/ijmm.2011.617 Abstract. Differentiated embryonic chondrocyte gene Introduction (DEC) 1 (BHLHE40/Stra13/Sharp2) and DEC2 (BHLHE41/ Sharp1) are basic helix-loop-helix (bHLH) transcription Paclitaxel is an anti-tumor drug that is used against a wide factors that are associated with the regulation of apoptosis, cell variety of solid tumors (1,2) and affects the expression of proliferation and circadian rhythms, as well as malignancy Bcl-2, Bcl-xL and phosphorylated-c-Jun NH2-terminal kinase in various cancers. However, the roles of DEC1 and DEC2 (JNK), and the activation of caspases and poly (ADP-ribose) expression in breast cancer are poorly understood. In this polymerase PARP (3-6), resulting in the induction of apop- study, we sought to examine the roles of DEC1 and DEC2 tosis by p53-dependent or -independent mechanisms (7-9). -

Characterization of a Novel Mouse Model with Genetic Deletion of CD177
Protein Cell 2015, 6(2):117–126 DOI 10.1007/s13238-014-0109-1 Protein & Cell RESEARCH ARTICLE Characterization of a novel mouse model with genetic deletion of CD177 Qing Xie1,2, Julia Klesney-Tait3, Kathy Keck3, Corey Parlet2, Nicholas Borcherding2, Ryan Kolb2, Wei Li2, & Lorraine Tygrett2, Thomas Waldschmidt2, Alicia Olivier2, Songhai Chen4, Guang-Hui Liu5,6, Xiangrui Li1 , Weizhou Zhang2& 1 College of Veterinary Medicine, Nanjing Agricultural University, Nanjing 210095, China 2 Department of Pathology, Holden Comprehensive Cancer Center, Carver College of Medicine/University of Iowa, Iowa, IA 52242, USA 3 Department of Internal Medicine, Carver College of Medicine/University of Iowa, Iowa, IA 52242, USA 4 Department of Pharmacology, Carver College of Medicine/University of Iowa, Iowa, IA 52242, USA Cell 5 National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China & 6 Beijing Institute for Brain Disorders, Beijing 100069, China & Correspondence: [email protected] (X. Li), [email protected] (W. Zhang) Received September 1, 2014 Accepted September 25, 2014 Protein ABSTRACT neutrophil counts in inflammatory skin caused by S. aureus. Mechanistically we found that CD177 deletion in Neutrophils play an essential role in the innate immune mouse neutrophils has no significant impact in CXCL1/ response to infection. Neutrophils migrate from the KC- or fMLP-induced migration, but led to significant cell vasculature into the tissue in response to infection. death. Herein we established a novel genetic mouse Recently, a neutrophil cell surface receptor, CD177, was model to study the role of CD177 and found that CD177 shown to help mediate neutrophil migration across the plays an important role in neutrophils. -

The Mineralocorticoid Receptor Leads to Increased Expression of EGFR
www.nature.com/scientificreports OPEN The mineralocorticoid receptor leads to increased expression of EGFR and T‑type calcium channels that support HL‑1 cell hypertrophy Katharina Stroedecke1,2, Sandra Meinel1,2, Fritz Markwardt1, Udo Kloeckner1, Nicole Straetz1, Katja Quarch1, Barbara Schreier1, Michael Kopf1, Michael Gekle1 & Claudia Grossmann1* The EGF receptor (EGFR) has been extensively studied in tumor biology and recently a role in cardiovascular pathophysiology was suggested. The mineralocorticoid receptor (MR) is an important efector of the renin–angiotensin–aldosterone‑system and elicits pathophysiological efects in the cardiovascular system; however, the underlying molecular mechanisms are unclear. Our aim was to investigate the importance of EGFR for MR‑mediated cardiovascular pathophysiology because MR is known to induce EGFR expression. We identifed a SNP within the EGFR promoter that modulates MR‑induced EGFR expression. In RNA‑sequencing and qPCR experiments in heart tissue of EGFR KO and WT mice, changes in EGFR abundance led to diferential expression of cardiac ion channels, especially of the T‑type calcium channel CACNA1H. Accordingly, CACNA1H expression was increased in WT mice after in vivo MR activation by aldosterone but not in respective EGFR KO mice. Aldosterone‑ and EGF‑responsiveness of CACNA1H expression was confrmed in HL‑1 cells by Western blot and by measuring peak current density of T‑type calcium channels. Aldosterone‑induced CACNA1H protein expression could be abrogated by the EGFR inhibitor AG1478. Furthermore, inhibition of T‑type calcium channels with mibefradil or ML218 reduced diameter, volume and BNP levels in HL‑1 cells. In conclusion the MR regulates EGFR and CACNA1H expression, which has an efect on HL‑1 cell diameter, and the extent of this regulation seems to depend on the SNP‑216 (G/T) genotype. -

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -

Downregulation of Dipeptidyl Peptidase 4 Accelerates Progression to Castration-Resistant Prostate Cancer Joshua W
Published OnlineFirst September 21, 2018; DOI: 10.1158/0008-5472.CAN-18-0687 Cancer Priority Report Research Downregulation of Dipeptidyl Peptidase 4 Accelerates Progression to Castration-Resistant Prostate Cancer Joshua W. Russo1,CeGao2, Swati S. Bhasin2, Olga S. Voznesensky1, Carla Calagua3, Seiji Arai1,4, Peter S. Nelson5, Bruce Montgomery6, Elahe A. Mostaghel5, Eva Corey6, Mary-Ellen Taplin7, Huihui Ye3, Manoj Bhasin2, and Steven P. Balk1 Abstract The standard treatment for metastatic prostate cancer, cleaves dipeptides from multiple growth factors, resulting in androgen deprivation therapy (ADT), is designed to suppress their increased degradation. DPP4 mRNA and protein were androgen receptor (AR) activity. However, men invariably also decreased in clinical CRPC cases, and inhibition of progress to castration-resistant prostate cancer (CRPC), and DPP4 with sitagliptin enhanced the growth of prostate AR reactivation contributes to progression in most cases. To cancer xenografts following castration. Significantly, DPP4 identify mechanisms that may drive CRPC, we examined a inhibitors are frequently used to treat type 2 diabetes as they VCaP prostate cancer xenograft model as tumors progressed increase insulin secretion. Together, these results implicate from initial androgen sensitivity prior to castration to castra- DPP4 as an AR-regulated tumor suppressor gene whose tion resistance and then on to relapse after combined therapy loss enhances growth factor activity and suggest that treat- with further AR-targeted drugs (abiraterone plus enzaluta- ment with DPP4 inhibitors may accelerate emergence of mide). AR activity persisted in castration-resistant and abir- resistance to ADT. aterone/enzalutamide–resistant xenografts and was associated with increased expression of the AR gene and the AR-V7 splice Significance: These findings identify DPP4 as an AR-stim- variant. -
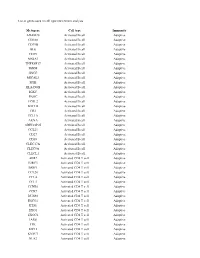
List of Genes Used in Cell Type Enrichment Analysis
List of genes used in cell type enrichment analysis Metagene Cell type Immunity ADAM28 Activated B cell Adaptive CD180 Activated B cell Adaptive CD79B Activated B cell Adaptive BLK Activated B cell Adaptive CD19 Activated B cell Adaptive MS4A1 Activated B cell Adaptive TNFRSF17 Activated B cell Adaptive IGHM Activated B cell Adaptive GNG7 Activated B cell Adaptive MICAL3 Activated B cell Adaptive SPIB Activated B cell Adaptive HLA-DOB Activated B cell Adaptive IGKC Activated B cell Adaptive PNOC Activated B cell Adaptive FCRL2 Activated B cell Adaptive BACH2 Activated B cell Adaptive CR2 Activated B cell Adaptive TCL1A Activated B cell Adaptive AKNA Activated B cell Adaptive ARHGAP25 Activated B cell Adaptive CCL21 Activated B cell Adaptive CD27 Activated B cell Adaptive CD38 Activated B cell Adaptive CLEC17A Activated B cell Adaptive CLEC9A Activated B cell Adaptive CLECL1 Activated B cell Adaptive AIM2 Activated CD4 T cell Adaptive BIRC3 Activated CD4 T cell Adaptive BRIP1 Activated CD4 T cell Adaptive CCL20 Activated CD4 T cell Adaptive CCL4 Activated CD4 T cell Adaptive CCL5 Activated CD4 T cell Adaptive CCNB1 Activated CD4 T cell Adaptive CCR7 Activated CD4 T cell Adaptive DUSP2 Activated CD4 T cell Adaptive ESCO2 Activated CD4 T cell Adaptive ETS1 Activated CD4 T cell Adaptive EXO1 Activated CD4 T cell Adaptive EXOC6 Activated CD4 T cell Adaptive IARS Activated CD4 T cell Adaptive ITK Activated CD4 T cell Adaptive KIF11 Activated CD4 T cell Adaptive KNTC1 Activated CD4 T cell Adaptive NUF2 Activated CD4 T cell Adaptive PRC1 Activated -

Finding Drug Targeting Mechanisms with Genetic Evidence for Parkinson’S Disease
bioRxiv preprint doi: https://doi.org/10.1101/2020.07.24.208975; this version posted July 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Finding drug targeting mechanisms with genetic evidence for Parkinson’s disease Catherine S. Storm1,*, Demis A. Kia1, Mona Almramhi1, Sara Bandres-Ciga2, Chris Finan3, Aroon D. Hingorani3,4,5, International Parkinson’s Disease Genomics Consortium (IPDGC), Nicholas W. Wood1,6,* 1 Department of Clinical and Movement Neurosciences, University College London Queen Square Institute of Neurology, London, WC1N 3BG, United Kingdom 2 Molecular Genetics Section, Laboratory of Neurogenetics, National Institute on Aging, National Institutes of Health, Bethesda, MD 20892, United States of America 3 Institute of Cardiovascular Science, Faculty of Population Health, University College London, London WC1E 6BT, United Kingdom 4 University College London British Heart Foundation Research Accelerator Centre, New Delhi, India 5 Health Data Research UK, 222 Euston Road, London, United Kingdom 6 Lead Contact * Correspondence: [email protected] (CSS), [email protected] (NWW) Summary Parkinson’s disease (PD) is a neurodegenerative movement disorder that currently has no disease-modifying treatment, partly owing to inefficiencies in drug target identification and validation using human evidence. Here, we use Mendelian randomization to investigate more than 3000 genes that encode druggable proteins, seeking to predict their efficacy as drug targets for PD. We use expression and protein quantitative trait loci for druggable genes to mimic exposure to medications, and we examine the causal effect on PD risk (in two large case-control cohorts), PD age at onset and progression. -

The Title of the Dissertation
UNIVERSITY OF CALIFORNIA SAN DIEGO Novel network-based integrated analyses of multi-omics data reveal new insights into CD8+ T cell differentiation and mouse embryogenesis A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Bioinformatics and Systems Biology by Kai Zhang Committee in charge: Professor Wei Wang, Chair Professor Pavel Arkadjevich Pevzner, Co-Chair Professor Vineet Bafna Professor Cornelis Murre Professor Bing Ren 2018 Copyright Kai Zhang, 2018 All rights reserved. The dissertation of Kai Zhang is approved, and it is accept- able in quality and form for publication on microfilm and electronically: Co-Chair Chair University of California San Diego 2018 iii EPIGRAPH The only true wisdom is in knowing you know nothing. —Socrates iv TABLE OF CONTENTS Signature Page ....................................... iii Epigraph ........................................... iv Table of Contents ...................................... v List of Figures ........................................ viii List of Tables ........................................ ix Acknowledgements ..................................... x Vita ............................................. xi Abstract of the Dissertation ................................. xii Chapter 1 General introduction ............................ 1 1.1 The applications of graph theory in bioinformatics ......... 1 1.2 Leveraging graphs to conduct integrated analyses .......... 4 1.3 References .............................. 6 Chapter 2 Systematic -

The Role of Tumor Necrosis Factor-Like Weak Inducer of Apoptosis in Atherosclerosis Via Its Two Different Receptors (Review)
EXPERIMENTAL AND THERAPEUTIC MEDICINE 14: 891-897, 2017 The role of tumor necrosis factor-like weak inducer of apoptosis in atherosclerosis via its two different receptors (Review) HENGDAO LIU1,2, DAN LIN3, HONG XIANG1, WEI CHEN1, SHAOLI ZHAO1,4, HUI PENG1, JIE YANG1, PAN CHEN1, SHUHUA CHEN1 and HONGWEI LU1,2 1Center for Experimental Medical Research; 2Department of Cardiology, The Third Xiangya Hospital of Central South University, Changsha, Hunan 410013; 3Qingdao Center for Disease Control and Prevention, Qingdao, Shandong 266033; 4Department of Endocrinology, The Third Xiangya Hospital of Central South University, Changsha, Hunan 410013, P.R. China Received May 3, 2016; Accepted March 31, 2017 DOI: 10.3892/etm.2017.4600 Abstract. At present, it is commonly accepted that athero- 1. Introduction sclerosis is a chronic inflammatory disease characterized by disorder of the arterial wall. As one of the inflammatory cyto- Atherosclerosis remains a major global challenge (1). The kines of the tumor necrosis factor superfamily, tumor necrosis World Health Organization statistics data suggest that, by factor-like weak inducer of apoptosis (TWEAK) participates 2020, atherosclerosis will be the most common cause of death in the formation and progression of atherosclerosis. TWEAK, and morbidity in both developed and developing countries (2). when binding to its initial receptor, fibroblast growth factor Atherosclerosis has long been studied as a chronic inflamma- inducible molecule 14 (Fn14), exerts adverse biological func- tory disorder of the arterial wall, which involves numerous tions in atherosclerosis, including dysfunction of endothelial cytokines. Accumulating data suggests that one of the cyto- cells, phenotypic change of smooth muscle cells and inflam- kines, tumor necrosis factor-like weak inducer of apoptosis matory responses of monocytes/macrophages.