FASTQ Format VCF Format Narrowpeak Unaligned Reads Gappedpeak Bed Bedgraph Alignment to Reference Compress to Binary Format
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Discovery and Annotation Using LCM-454 Transcriptome Sequencing Scott J
Downloaded from genome.cshlp.org on September 23, 2021 - Published by Cold Spring Harbor Laboratory Press Methods Gene discovery and annotation using LCM-454 transcriptome sequencing Scott J. Emrich,1,2,6 W. Brad Barbazuk,3,6 Li Li,4 and Patrick S. Schnable1,4,5,7 1Bioinformatics and Computational Biology Graduate Program, Iowa State University, Ames, Iowa 50010, USA; 2Department of Electrical and Computer Engineering, Iowa State University, Ames, Iowa 50010, USA; 3Donald Danforth Plant Science Center, St. Louis, Missouri 63132, USA; 4Interdepartmental Plant Physiology Graduate Major and Department of Genetics, Development, and Cell Biology, Iowa State University, Ames, Iowa 50010, USA; 5Department of Agronomy and Center for Plant Genomics, Iowa State University, Ames, Iowa 50010, USA 454 DNA sequencing technology achieves significant throughput relative to traditional approaches. More than 261,000 ESTs were generated by 454 Life Sciences from cDNA isolated using laser capture microdissection (LCM) from the developmentally important shoot apical meristem (SAM) of maize (Zea mays L.). This single sequencing run annotated >25,000 maize genomic sequences and also captured ∼400 expressed transcripts for which homologous sequences have not yet been identified in other species. Approximately 70% of the ESTs generated in this study had not been captured during a previous EST project conducted using a cDNA library constructed from hand-dissected apex tissue that is highly enriched for SAMs. In addition, at least 30% of the 454-ESTs do not align to any of the ∼648,000 extant maize ESTs using conservative alignment criteria. These results indicate that the combination of LCM and the deep sequencing possible with 454 technology enriches for SAM transcripts not present in current EST collections. -

The Variant Call Format Specification Vcfv4.3 and Bcfv2.2
The Variant Call Format Specification VCFv4.3 and BCFv2.2 27 Jul 2021 The master version of this document can be found at https://github.com/samtools/hts-specs. This printing is version 1715701 from that repository, last modified on the date shown above. 1 Contents 1 The VCF specification 4 1.1 An example . .4 1.2 Character encoding, non-printable characters and characters with special meaning . .4 1.3 Data types . .4 1.4 Meta-information lines . .5 1.4.1 File format . .5 1.4.2 Information field format . .5 1.4.3 Filter field format . .5 1.4.4 Individual format field format . .6 1.4.5 Alternative allele field format . .6 1.4.6 Assembly field format . .6 1.4.7 Contig field format . .6 1.4.8 Sample field format . .7 1.4.9 Pedigree field format . .7 1.5 Header line syntax . .7 1.6 Data lines . .7 1.6.1 Fixed fields . .7 1.6.2 Genotype fields . .9 2 Understanding the VCF format and the haplotype representation 11 2.1 VCF tag naming conventions . 12 3 INFO keys used for structural variants 12 4 FORMAT keys used for structural variants 13 5 Representing variation in VCF records 13 5.1 Creating VCF entries for SNPs and small indels . 13 5.1.1 Example 1 . 13 5.1.2 Example 2 . 14 5.1.3 Example 3 . 14 5.2 Decoding VCF entries for SNPs and small indels . 14 5.2.1 SNP VCF record . 14 5.2.2 Insertion VCF record . -

File Formats Exercises
Introduction to high-throughput sequencing file formats – advanced exercises Daniel Vodák (Bioinformatics Core Facility, The Norwegian Radium Hospital) ( [email protected] ) All example files are located in directory “/data/file_formats/data”. You can set your current directory to that path (“cd /data/file_formats/data”) Exercises: a) FASTA format File “Uniprot_Mammalia.fasta” contains a collection of mammalian protein sequences obtained from the UniProt database ( http://www.uniprot.org/uniprot ). 1) Count the number of lines in the file. 2) Count the number of header lines (i.e. the number of sequences) in the file. 3) Count the number of header lines with “Homo” in the organism/species name. 4) Count the number of header lines with “Homo sapiens” in the organism/species name. 5) Display the header lines which have “Homo” in the organism/species names, but only such that do not have “Homo sapiens” there. b) FASTQ format File NKTCL_31T_1M_R1.fq contains a reduced collection of tumor read sequences obtained from The Sequence Read Archive ( http://www.ncbi.nlm.nih.gov/sra ). 1) Count the number of lines in the file. 2) Count the number of lines beginning with the “at” symbol (“@”). 3) How many reads are there in the file? c) SAM format File NKTCL_31T_1M.sam contains alignments of pair-end reads stored in files NKTCL_31T_1M_R1.fq and NKTCL_31T_1M_R2.fq. 1) Which program was used for the alignment? 2) How many header lines are there in the file? 3) How many non-header lines are there in the file? 4) What do the non-header lines represent? 5) Reads from how many template sequences have been used in the alignment process? c) BED format File NKTCL_31T_1M.bed holds information about alignment locations stored in file NKTCL_31T_1M.sam. -

To Find Information About Arabidopsis Genes Leonore Reiser1, Shabari
UNIT 1.11 Using The Arabidopsis Information Resource (TAIR) to Find Information About Arabidopsis Genes Leonore Reiser1, Shabari Subramaniam1, Donghui Li1, and Eva Huala1 1Phoenix Bioinformatics, Redwood City, CA USA ABSTRACT The Arabidopsis Information Resource (TAIR; http://arabidopsis.org) is a comprehensive Web resource of Arabidopsis biology for plant scientists. TAIR curates and integrates information about genes, proteins, gene function, orthologs gene expression, mutant phenotypes, biological materials such as clones and seed stocks, genetic markers, genetic and physical maps, genome organization, images of mutant plants, protein sub-cellular localizations, publications, and the research community. The various data types are extensively interconnected and can be accessed through a variety of Web-based search and display tools. This unit primarily focuses on some basic methods for searching, browsing, visualizing, and analyzing information about Arabidopsis genes and genome, Additionally we describe how members of the community can share data using TAIR’s Online Annotation Submission Tool (TOAST), in order to make their published research more accessible and visible. Keywords: Arabidopsis ● databases ● bioinformatics ● data mining ● genomics INTRODUCTION The Arabidopsis Information Resource (TAIR; http://arabidopsis.org) is a comprehensive Web resource for the biology of Arabidopsis thaliana (Huala et al., 2001; Garcia-Hernandez et al., 2002; Rhee et al., 2003; Weems et al., 2004; Swarbreck et al., 2008, Lamesch, et al., 2010, Berardini et al., 2016). The TAIR database contains information about genes, proteins, gene expression, mutant phenotypes, germplasms, clones, genetic markers, genetic and physical maps, genome organization, publications, and the research community. In addition, seed and DNA stocks from the Arabidopsis Biological Resource Center (ABRC; Scholl et al., 2003) are integrated with genomic data, and can be ordered through TAIR. -

A Semantic Standard for Describing the Location of Nucleotide and Protein Feature Annotation Jerven T
Bolleman et al. Journal of Biomedical Semantics (2016) 7:39 DOI 10.1186/s13326-016-0067-z RESEARCH Open Access FALDO: a semantic standard for describing the location of nucleotide and protein feature annotation Jerven T. Bolleman1*, Christopher J. Mungall2, Francesco Strozzi3, Joachim Baran4, Michel Dumontier5, Raoul J. P. Bonnal6, Robert Buels7, Robert Hoehndorf8, Takatomo Fujisawa9, Toshiaki Katayama10 and Peter J. A. Cock11 Abstract Background: Nucleotide and protein sequence feature annotations are essential to understand biology on the genomic, transcriptomic, and proteomic level. Using Semantic Web technologies to query biological annotations, there was no standard that described this potentially complex location information as subject-predicate-object triples. Description: We have developed an ontology, the Feature Annotation Location Description Ontology (FALDO), to describe the positions of annotated features on linear and circular sequences. FALDO can be used to describe nucleotide features in sequence records, protein annotations, and glycan binding sites, among other features in coordinate systems of the aforementioned “omics” areas. Using the same data format to represent sequence positions that are independent of file formats allows us to integrate sequence data from multiple sources and data types. The genome browser JBrowse is used to demonstrate accessing multiple SPARQL endpoints to display genomic feature annotations, as well as protein annotations from UniProt mapped to genomic locations. Conclusions: Our ontology allows -

EMBL-EBI Powerpoint Presentation
Processing data from high-throughput sequencing experiments Simon Anders Use-cases for HTS · de-novo sequencing and assembly of small genomes · transcriptome analysis (RNA-Seq, sRNA-Seq, ...) • identifying transcripted regions • expression profiling · Resequencing to find genetic polymorphisms: • SNPs, micro-indels • CNVs · ChIP-Seq, nucleosome positions, etc. · DNA methylation studies (after bisulfite treatment) · environmental sampling (metagenomics) · reading bar codes Use cases for HTS: Bioinformatics challenges Established procedures may not be suitable. New algorithms are required for · assembly · alignment · statistical tests (counting statistics) · visualization · segmentation · ... Where does Bioconductor come in? Several steps: · Processing of the images and determining of the read sequencest • typically done by core facility with software from the manufacturer of the sequencing machine · Aligning the reads to a reference genome (or assembling the reads into a new genome) • Done with community-developed stand-alone tools. · Downstream statistical analyis. • Write your own scripts with the help of Bioconductor infrastructure. Solexa standard workflow SolexaPipeline · "Firecrest": Identifying clusters ⇨ typically 15..20 mio good clusters per lane · "Bustard": Base calling ⇨ sequence for each cluster, with Phred-like scores · "Eland": Aligning to reference Firecrest output Large tab-separated text files with one row per identified cluster, specifying · lane index and tile index · x and y coordinates of cluster on tile · for each -

The Encodedb Portal: Simplified Access to ENCODE Consortium Data Laura L
Downloaded from genome.cshlp.org on September 30, 2021 - Published by Cold Spring Harbor Laboratory Press Resource The ENCODEdb portal: Simplified access to ENCODE Consortium data Laura L. Elnitski, Prachi Shah, R. Travis Moreland, Lowell Umayam, Tyra G. Wolfsberg, and Andreas D. Baxevanis1 Genome Technology Branch, National Human Genome Research Institute, National Institutes of Health, Bethesda, Maryland 20892, USA The Encyclopedia of DNA Elements (ENCODE) project aims to identify and characterize all functional elements in a representative chromosomal sample comprising 1% of the human genome. Data generated by members of The ENCODE Project Consortium are housed in a number of public databases, such as the UCSC Genome Browser, NCBI’s Gene Expression Omnibus (GEO), and EBI’s ArrayExpress. As such, it is often difficult for biologists to gather all of the ENCODE data from a particular genomic region of interest and integrate them with relevant information found in other public databases. The ENCODEdb portal was developed to address this problem. ENCODEdb provides a unified, single point-of-access to data generated by the ENCODE Consortium, as well as to data from other source databases that lie within ENCODE regions; this provides the user a complete view of all known data in a particular region of interest. ENCODEdb Genomic Context searches allow for the retrieval of information on functional elements annotated within ENCODE regions, including mRNA, EST, and STS sequences; single nucleotide polymorphisms, and UniGene clusters. Information is also retrieved from GEO, OMIM, and major genome sequence browsers. ENCODEdb Consortium Data searches allow users to perform compound queries on array-based ENCODE data available both from GEO and from the UCSC Genome Browser. -
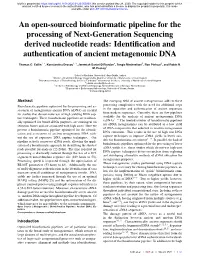
An Open-Sourced Bioinformatic Pipeline for the Processing of Next-Generation Sequencing Derived Nucleotide Reads
bioRxiv preprint doi: https://doi.org/10.1101/2020.04.20.050369; this version posted May 28, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. An open-sourced bioinformatic pipeline for the processing of Next-Generation Sequencing derived nucleotide reads: Identification and authentication of ancient metagenomic DNA Thomas C. Collin1, *, Konstantina Drosou2, 3, Jeremiah Daniel O’Riordan4, Tengiz Meshveliani5, Ron Pinhasi6, and Robin N. M. Feeney1 1School of Medicine, University College Dublin, Ireland 2Division of Cell Matrix Biology Regenerative Medicine, University of Manchester, United Kingdom 3Manchester Institute of Biotechnology, School of Earth and Environmental Sciences, University of Manchester, United Kingdom [email protected] 5Institute of Paleobiology and Paleoanthropology, National Museum of Georgia, Tbilisi, Georgia 6Department of Evolutionary Anthropology, University of Vienna, Austria *Corresponding Author Abstract The emerging field of ancient metagenomics adds to these Bioinformatic pipelines optimised for the processing and as- processing complexities with the need for additional steps sessment of metagenomic ancient DNA (aDNA) are needed in the separation and authentication of ancient sequences from modern sequences. Currently, there are few pipelines for studies that do not make use of high yielding DNA cap- available for the analysis of ancient metagenomic DNA ture techniques. These bioinformatic pipelines are tradition- 1 4 ally optimised for broad aDNA purposes, are contingent on (aDNA) ≠ The limited number of bioinformatic pipelines selection biases and are associated with high costs. -

A Combined RAD-Seq and WGS Approach Reveals the Genomic
www.nature.com/scientificreports OPEN A combined RAD‑Seq and WGS approach reveals the genomic basis of yellow color variation in bumble bee Bombus terrestris Sarthok Rasique Rahman1,2, Jonathan Cnaani3, Lisa N. Kinch4, Nick V. Grishin4 & Heather M. Hines1,5* Bumble bees exhibit exceptional diversity in their segmental body coloration largely as a result of mimicry. In this study we sought to discover genes involved in this variation through studying a lab‑generated mutant in bumble bee Bombus terrestris, in which the typical black coloration of the pleuron, scutellum, and frst metasomal tergite is replaced by yellow, a color variant also found in sister lineages to B. terrestris. Utilizing a combination of RAD‑Seq and whole‑genome re‑sequencing, we localized the color‑generating variant to a single SNP in the protein‑coding sequence of transcription factor cut. This mutation generates an amino acid change that modifes the conformation of a coiled‑coil structure outside DNA‑binding domains. We found that all sequenced Hymenoptera, including sister lineages, possess the non‑mutant allele, indicating diferent mechanisms are involved in the same color transition in nature. Cut is important for multiple facets of development, yet this mutation generated no noticeable external phenotypic efects outside of setal characteristics. Reproductive capacity was reduced, however, as queens were less likely to mate and produce female ofspring, exhibiting behavior similar to that of workers. Our research implicates a novel developmental player in pigmentation, and potentially caste, thus contributing to a better understanding of the evolution of diversity in both of these processes. Understanding the genetic architecture underlying phenotypic diversifcation has been a long-standing goal of evolutionary biology. -

Sequence Alignment/Map Format Specification
Sequence Alignment/Map Format Specification The SAM/BAM Format Specification Working Group 3 Jun 2021 The master version of this document can be found at https://github.com/samtools/hts-specs. This printing is version 53752fa from that repository, last modified on the date shown above. 1 The SAM Format Specification SAM stands for Sequence Alignment/Map format. It is a TAB-delimited text format consisting of a header section, which is optional, and an alignment section. If present, the header must be prior to the alignments. Header lines start with `@', while alignment lines do not. Each alignment line has 11 mandatory fields for essential alignment information such as mapping position, and variable number of optional fields for flexible or aligner specific information. This specification is for version 1.6 of the SAM and BAM formats. Each SAM and BAMfilemay optionally specify the version being used via the @HD VN tag. For full version history see Appendix B. Unless explicitly specified elsewhere, all fields are encoded using 7-bit US-ASCII 1 in using the POSIX / C locale. Regular expressions listed use the POSIX / IEEE Std 1003.1 extended syntax. 1.1 An example Suppose we have the following alignment with bases in lowercase clipped from the alignment. Read r001/1 and r001/2 constitute a read pair; r003 is a chimeric read; r004 represents a split alignment. Coor 12345678901234 5678901234567890123456789012345 ref AGCATGTTAGATAA**GATAGCTGTGCTAGTAGGCAGTCAGCGCCAT +r001/1 TTAGATAAAGGATA*CTG +r002 aaaAGATAA*GGATA +r003 gcctaAGCTAA +r004 ATAGCT..............TCAGC -r003 ttagctTAGGC -r001/2 CAGCGGCAT The corresponding SAM format is:2 1Charset ANSI X3.4-1968 as defined in RFC1345. -

Orthofiller: Utilising Data from Multiple Species to Improve the Completeness of Genome Annotations Michael P
Dunne and Kelly BMC Genomics (2017) 18:390 DOI 10.1186/s12864-017-3771-x SOFTWARE Open Access OrthoFiller: utilising data from multiple species to improve the completeness of genome annotations Michael P. Dunne and Steven Kelly* Abstract Backround: Complete and accurate annotation of sequenced genomes is of paramount importance to their utility and analysis. Differences in gene prediction pipelines mean that genome annotations for a species can differ considerably in the quality and quantity of their predicted genes. Furthermore, genes that are present in genome sequences sometimes fail to be detected by computational gene prediction methods. Erroneously unannotated genes can lead to oversights and inaccurate assertions in biological investigations, especially for smaller-scale genome projects, which rely heavily on computational prediction. Results: Here we present OrthoFiller, a tool designed to address the problem of finding and adding such missing genes to genome annotations. OrthoFiller leverages information from multiple related species to identify those genes whose existence can be verified through comparison with known gene families, but which have not been predicted. By simulating missing gene annotations in real sequence datasets from both plants and fungi we demonstrate the accuracy and utility of OrthoFiller for finding missing genes and improving genome annotations. Furthermore, we show that applying OrthoFiller to existing “complete” genome annotations can identify and correct substantial numbers of erroneously missing genes in these two sets of species. Conclusions: We show that significant improvements in the completeness of genome annotations can be made by leveraging information from multiple species. Keywords: Genome annotation, Gene prediction, Orthology, Orthogroup Background of several effective algorithms for identifying genes in Genome sequences have become fundamental to many de novo sequenced genomes [3]. -

Infravec2 Open Research Data Management Plan
INFRAVEC2 OPEN RESEARCH DATA MANAGEMENT PLAN Authors: Andrea Crisanti, Gareth Maslen, Andy Yates, Paul Kersey, Alain Kohl, Clelia Supparo, Ken Vernick Date: 10th July 2020 Version: 3.0 Overview Infravec2 will align to Open Research Data, as follows: Data Types and Standards Infravec2 will generate a variety of data types, including molecular data types: genome sequence and assembly, structural annotation (gene models, repeats, other functional regions) and functional annotation (protein function assignment), variation data, and transcriptome data; arbovirus and malaria experimental infection data, linked to archived samples; and microbiome data (Operational Taxonomic Units), including natural virome composition. All data will be released according to the appropriate standards and formats for each data type. For example, DNA sequence will be released in FASTA format; variant calls in Variant Call Format; sequence alignments in BAM (Binary Alignment Map) and CRAM (Compressed Read Alignment Map) formats, etc. We will strongly encourage the organisation of linked data sets as Track Hubs, a mechanism for publishing a set of linked genomic data that aids data discovery, sharing, and selection for subsequent analysis. We will develop internal standards within the consortium to define minimal metadata that will accompany all data sets, following the template of the Minimal Information Standards for Biological and Biomedical Investigations (http://www.dcc.ac.uk/resources/metadata-standards/mibbi-minimum-information-biological- and-biomedical-investigations). Data Exploitation, Accessibility, Curation and Preservation All molecular data for which existing public data repositories exist will be submitted to such repositories on or before the publication of written manuscripts, with early release of data (i.e.