Identification of Long-Lived Synaptic Proteins by Proteomic Analysis Of
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Histone Isoform H2A1H Promotes Attainment of Distinct Physiological
Bhattacharya et al. Epigenetics & Chromatin (2017) 10:48 DOI 10.1186/s13072-017-0155-z Epigenetics & Chromatin RESEARCH Open Access Histone isoform H2A1H promotes attainment of distinct physiological states by altering chromatin dynamics Saikat Bhattacharya1,4,6, Divya Reddy1,4, Vinod Jani5†, Nikhil Gadewal3†, Sanket Shah1,4, Raja Reddy2,4, Kakoli Bose2,4, Uddhavesh Sonavane5, Rajendra Joshi5 and Sanjay Gupta1,4* Abstract Background: The distinct functional efects of the replication-dependent histone H2A isoforms have been dem- onstrated; however, the mechanistic basis of the non-redundancy remains unclear. Here, we have investigated the specifc functional contribution of the histone H2A isoform H2A1H, which difers from another isoform H2A2A3 in the identity of only three amino acids. Results: H2A1H exhibits varied expression levels in diferent normal tissues and human cancer cell lines (H2A1C in humans). It also promotes cell proliferation in a context-dependent manner when exogenously overexpressed. To uncover the molecular basis of the non-redundancy, equilibrium unfolding of recombinant H2A1H-H2B dimer was performed. We found that the M51L alteration at the H2A–H2B dimer interface decreases the temperature of melting of H2A1H-H2B by ~ 3 °C as compared to the H2A2A3-H2B dimer. This diference in the dimer stability is also refected in the chromatin dynamics as H2A1H-containing nucleosomes are more stable owing to M51L and K99R substitu- tions. Molecular dynamic simulations suggest that these substitutions increase the number of hydrogen bonds and hydrophobic interactions of H2A1H, enabling it to form more stable nucleosomes. Conclusion: We show that the M51L and K99R substitutions, besides altering the stability of histone–histone and histone–DNA complexes, have the most prominent efect on cell proliferation, suggesting that the nucleosome sta- bility is intimately linked with the physiological efects observed. -

AP2A2 Antibody Cat
AP2A2 Antibody Cat. No.: 63-513 AP2A2 Antibody Formalin-fixed and paraffin-embedded human Flow cytometric analysis of HepG2 cells using AP2A2 hepatocarcinoma with AP2A2 Antibody , which was Antibody (bottom histogram) compared to a negative peroxidase-conjugated to the secondary antibody, control cell (top histogram). FITC-conjugated goat-anti- followed by DAB staining. rabbit secondary antibodies were used for the analysis. Specifications HOST SPECIES: Rabbit SPECIES REACTIVITY: Human This AP2A2 antibody is generated from rabbits immunized with a KLH conjugated IMMUNOGEN: synthetic peptide between 610-637 amino acids from the Central region of human AP2A2. TESTED APPLICATIONS: Flow, IHC-P, WB For WB starting dilution is: 1:1000 APPLICATIONS: For IHC-P starting dilution is: 1:10~50 For FACS starting dilution is: 1:10~50 September 25, 2021 1 https://www.prosci-inc.com/ap2a2-antibody-63-513.html PREDICTED MOLECULAR 104 kDa WEIGHT: Properties This antibody is purified through a protein A column, followed by peptide affinity PURIFICATION: purification. CLONALITY: Polyclonal ISOTYPE: Rabbit Ig CONJUGATE: Unconjugated PHYSICAL STATE: Liquid BUFFER: Supplied in PBS with 0.09% (W/V) sodium azide. CONCENTRATION: batch dependent Store at 4˚C for three months and -20˚C, stable for up to one year. As with all antibodies STORAGE CONDITIONS: care should be taken to avoid repeated freeze thaw cycles. Antibodies should not be exposed to prolonged high temperatures. Additional Info OFFICIAL SYMBOL: AP2A2 AP-2 complex subunit alpha-2, 100 kDa coated vesicle -
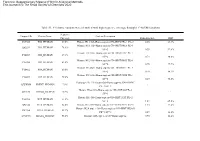
Table S1. 49 Histone Variants Were Identified with High Sequence Coverage Through LC-MS/MS Analysis Electronic Supplementary
Electronic Supplementary Material (ESI) for Analytical Methods. This journal is © The Royal Society of Chemistry 2020 Table S1. 49 histone variants were identified with high sequence coverage through LC-MS/MS analysis Sequence Uniprot IDs Protein Name Protein Description Coverage Ratio E2+/E2- RSD P07305 H10_HUMAN 67.5% Histone H1.0 OS=Homo sapiens GN=H1F0 PE=1 SV=3 4.85 23.3% Histone H1.1 OS=Homo sapiens GN=HIST1H1A PE=1 Q02539 H11_HUMAN 74.4% SV=3 0.35 92.6% Histone H1.2 OS=Homo sapiens GN=HIST1H1C PE=1 P16403 H12_HUMAN 67.1% SV=2 0.73 80.6% Histone H1.3 OS=Homo sapiens GN=HIST1H1D PE=1 P16402 H13_HUMAN 63.8% SV=2 0.75 77.7% Histone H1.4 OS=Homo sapiens GN=HIST1H1E PE=1 P10412 H14_HUMAN 69.0% SV=2 0.70 80.3% Histone H1.5 OS=Homo sapiens GN=HIST1H1B PE=1 P16401 H15_HUMAN 79.6% SV=3 0.29 98.3% Testis-specific H1 histone OS=Homo sapiens GN=H1FNT Q75WM6 H1FNT_HUMAN 7.8% \ \ PE=2 SV=3 Histone H1oo OS=Homo sapiens GN=H1FOO PE=2 Q8IZA3 H1FOO_HUMAN 5.2% \ \ SV=1 Histone H1t OS=Homo sapiens GN=HIST1H1T PE=2 P22492 H1T_HUMAN 31.4% SV=4 1.42 65.0% Q92522 H1X_HUMAN 82.6% Histone H1x OS=Homo sapiens GN=H1FX PE=1 SV=1 1.15 33.2% Histone H2A type 1 OS=Homo sapiens GN=HIST1H2AG P0C0S8 H2A1_HUMAN 99.2% PE=1 SV=2 0.57 26.8% Q96QV6 H2A1A_HUMAN 58.0% Histone H2A type 1-A OS=Homo sapiens 0.90 11.2% GN=HIST1H2AA PE=1 SV=3 Histone H2A type 1-B/E OS=Homo sapiens P04908 H2A1B_HUMAN 99.2% GN=HIST1H2AB PE=1 SV=2 0.92 30.2% Histone H2A type 1-C OS=Homo sapiens Q93077 H2A1C_HUMAN 100.0% GN=HIST1H2AC PE=1 SV=3 0.76 27.6% Histone H2A type 1-D OS=Homo sapiens P20671 -

UNIVERSITY of CALIFORNIA, SAN DIEGO Functional Analysis of Sall4
UNIVERSITY OF CALIFORNIA, SAN DIEGO Functional analysis of Sall4 in modulating embryonic stem cell fate A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Molecular Pathology by Pei Jen A. Lee Committee in charge: Professor Steven Briggs, Chair Professor Geoff Rosenfeld, Co-Chair Professor Alexander Hoffmann Professor Randall Johnson Professor Mark Mercola 2009 Copyright Pei Jen A. Lee, 2009 All rights reserved. The dissertation of Pei Jen A. Lee is approved, and it is acceptable in quality and form for publication on microfilm and electronically: ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ Co-Chair ______________________________________________________________ Chair University of California, San Diego 2009 iii Dedicated to my parents, my brother ,and my husband for their love and support iv Table of Contents Signature Page……………………………………………………………………….…iii Dedication…...…………………………………………………………………………..iv Table of Contents……………………………………………………………………….v List of Figures…………………………………………………………………………...vi List of Tables………………………………………………….………………………...ix Curriculum vitae…………………………………………………………………………x Acknowledgement………………………………………………….……….……..…...xi Abstract………………………………………………………………..…………….....xiii Chapter 1 Introduction ..…………………………………………………………………………….1 Chapter 2 Materials and Methods……………………………………………………………..…12 -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Author's Accepted Manuscript
Author’s Accepted Manuscript Synapse-specific stabilization of plasticity processes: The synaptic tagging and capture hypothesis revisited ten years later Angel Barco, Mikel Lopez de Armentia, Juan M. Alarcon PII: S0149-7634(08)00008-0 DOI: doi:10.1016/j.neubiorev.2008.01.002 Reference: NBR 1021 www.elsevier.com/locate/neubiorev To appear in: Neuroscience and Biobehavioral Reviews Received date: 30 October 2007 Revised date: 28 December 2007 Accepted date: 7 January 2008 Cite this article as: Angel Barco, Mikel Lopez de Armentia and Juan M. Alarcon, Synapse-specific stabilization of plasticity processes: The synaptic tagging and capture hypothesis revisited ten years later, Neuroscience and Biobehavioral Reviews (2008), doi:10.1016/j.neubiorev.2008.01.002 This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting galley proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain. The STC hypothesis revisited Synapse-specific stabilization of plasticity processes: The synaptic tagging and capture hypothesis revisited ten years later Angel Barco1*, Mikel Lopez de Armentia1 and Juan M. Alarcon2 1Instituto de Neurociencias de Alicante. (Universidad Miguel Hernández-Consejo Superior de Investigaciones Científicas). Campus de Sant Joan. Apt. 18. Sant Joan d’Alacant. 03550. Alicante, Spain. 2SUNY Downstate Medical Center. 450 Clarkson Ave., Box 25, Brooklyn, NY 11203. -

Download Download
Supplementary Figure S1. Results of flow cytometry analysis, performed to estimate CD34 positivity, after immunomagnetic separation in two different experiments. As monoclonal antibody for labeling the sample, the fluorescein isothiocyanate (FITC)- conjugated mouse anti-human CD34 MoAb (Mylteni) was used. Briefly, cell samples were incubated in the presence of the indicated MoAbs, at the proper dilution, in PBS containing 5% FCS and 1% Fc receptor (FcR) blocking reagent (Miltenyi) for 30 min at 4 C. Cells were then washed twice, resuspended with PBS and analyzed by a Coulter Epics XL (Coulter Electronics Inc., Hialeah, FL, USA) flow cytometer. only use Non-commercial 1 Supplementary Table S1. Complete list of the datasets used in this study and their sources. GEO Total samples Geo selected GEO accession of used Platform Reference series in series samples samples GSM142565 GSM142566 GSM142567 GSM142568 GSE6146 HG-U133A 14 8 - GSM142569 GSM142571 GSM142572 GSM142574 GSM51391 GSM51392 GSE2666 HG-U133A 36 4 1 GSM51393 GSM51394 only GSM321583 GSE12803 HG-U133A 20 3 GSM321584 2 GSM321585 use Promyelocytes_1 Promyelocytes_2 Promyelocytes_3 Promyelocytes_4 HG-U133A 8 8 3 GSE64282 Promyelocytes_5 Promyelocytes_6 Promyelocytes_7 Promyelocytes_8 Non-commercial 2 Supplementary Table S2. Chromosomal regions up-regulated in CD34+ samples as identified by the LAP procedure with the two-class statistics coded in the PREDA R package and an FDR threshold of 0.5. Functional enrichment analysis has been performed using DAVID (http://david.abcc.ncifcrf.gov/) -

Supplemental Information
Supplemental information Dissection of the genomic structure of the miR-183/96/182 gene. Previously, we showed that the miR-183/96/182 cluster is an intergenic miRNA cluster, located in a ~60-kb interval between the genes encoding nuclear respiratory factor-1 (Nrf1) and ubiquitin-conjugating enzyme E2H (Ube2h) on mouse chr6qA3.3 (1). To start to uncover the genomic structure of the miR- 183/96/182 gene, we first studied genomic features around miR-183/96/182 in the UCSC genome browser (http://genome.UCSC.edu/), and identified two CpG islands 3.4-6.5 kb 5’ of pre-miR-183, the most 5’ miRNA of the cluster (Fig. 1A; Fig. S1 and Seq. S1). A cDNA clone, AK044220, located at 3.2-4.6 kb 5’ to pre-miR-183, encompasses the second CpG island (Fig. 1A; Fig. S1). We hypothesized that this cDNA clone was derived from 5’ exon(s) of the primary transcript of the miR-183/96/182 gene, as CpG islands are often associated with promoters (2). Supporting this hypothesis, multiple expressed sequences detected by gene-trap clones, including clone D016D06 (3, 4), were co-localized with the cDNA clone AK044220 (Fig. 1A; Fig. S1). Clone D016D06, deposited by the German GeneTrap Consortium (GGTC) (http://tikus.gsf.de) (3, 4), was derived from insertion of a retroviral construct, rFlpROSAβgeo in 129S2 ES cells (Fig. 1A and C). The rFlpROSAβgeo construct carries a promoterless reporter gene, the β−geo cassette - an in-frame fusion of the β-galactosidase and neomycin resistance (Neor) gene (5), with a splicing acceptor (SA) immediately upstream, and a polyA signal downstream of the β−geo cassette (Fig. -

Systematic Identification of Genes Regulating Synaptic Remodeling In
Genes Genet. Syst. (2020) 95, p. 101–110 Genes required for synaptic remodeling 101 Systematic identification of genes regulating synaptic remodeling in the Drosophila visual system Tomohiro Araki, Jiro Osaka, Yuya Kato, Mai Shimozono, Hinata Kawamura, Riku Iwanaga, Satoko Hakeda-Suzuki and Takashi Suzuki* Graduate School of Life Science and Technology, Tokyo Institute of Technology, Yokohama, Kanagawa 226-8501, Japan (Received 17 December 2019, accepted 21 January 2020; J-STAGE Advance published date: 4 June 2020) In many animals, neural activity contributes to the adaptive refinement of synaptic properties, such as firing frequency and the number of synapses, for learning, memorizing and adapting for survival. However, the molecular mech- anisms underlying such activity-dependent synaptic remodeling remain largely unknown. In the synapses of Drosophila melanogaster, the presynaptic active zone (AZ) forms a T-shaped presynaptic density comprising AZ proteins, including Bruchpilot (Brp). In a previous study, we found that the signal from a fusion pro- tein molecular marker consisting of Brp and mCherry becomes diffuse under con- tinuous light over three days (LL), reflecting disassembly of the AZ, while remaining punctate under continuous darkness. To identify the molecular players control- ling this synaptic remodeling, we used the fusion protein molecular marker and performed RNAi screening against 208 neuron-related transmembrane genes that are highly expressed in the Drosophila visual system. Second analyses using the STaR (synaptic tagging with recombination) technique, which showed a decrease in synapse number under the LL condition, and subsequent mutant and overexpres- sion analysis confirmed that five genes are involved in the activity-dependent AZ disassembly. -

Identification of Novel Chemotherapeutic Strategies For
www.nature.com/scientificreports OPEN Identification of novel chemotherapeutic strategies for metastatic uveal melanoma Received: 17 November 2016 Paolo Fagone1, Rosario Caltabiano2, Andrea Russo3, Gabriella Lupo1, Accepted: 09 February 2017 Carmelina Daniela Anfuso1, Maria Sofia Basile1, Antonio Longo3, Ferdinando Nicoletti1, Published: 17 March 2017 Rocco De Pasquale4, Massimo Libra1 & Michele Reibaldi3 Melanoma of the uveal tract accounts for approximately 5% of all melanomas and represents the most common primary intraocular malignancy. Despite improvements in diagnosis and more effective local therapies for primary cancer, the rate of metastatic death has not changed in the past forty years. In the present study, we made use of bioinformatics to analyze the data obtained from three public available microarray datasets on uveal melanoma in an attempt to identify novel putative chemotherapeutic options for the liver metastatic disease. We have first carried out a meta-analysis of publicly available whole-genome datasets, that included data from 132 patients, comparing metastatic vs. non metastatic uveal melanomas, in order to identify the most relevant genes characterizing the spreading of tumor to the liver. Subsequently, the L1000CDS2 web-based utility was used to predict small molecules and drugs targeting the metastatic uveal melanoma gene signature. The most promising drugs were found to be Cinnarizine, an anti-histaminic drug used for motion sickness, Digitoxigenin, a precursor of cardiac glycosides, and Clofazimine, a fat-soluble iminophenazine used in leprosy. In vitro and in vivo validation studies will be needed to confirm the efficacy of these molecules for the prevention and treatment of metastatic uveal melanoma. Uveal melanoma is the most common primary intraocular cancer, and after the skin, the uveal tract is the second most common location for melanoma1. -

Chemical Modulation of Mitochondria–Endoplasmic Reticulum Contact Sites
cells Review Chemical Modulation of Mitochondria–Endoplasmic Reticulum Contact Sites 1 1, 1 1 Ana Paula Magalhães Rebelo , Federica Dal Bello y , Tomas Knedlik , Natasha Kaar , Fabio Volpin 1, Sang Hun Shin 1 and Marta Giacomello 1,2,* 1 Department of Biology, University of Padua, Via U. Bassi 58/B, 35121 Padua, Italy; [email protected] (A.P.M.R.); [email protected] (F.D.B.); [email protected] (T.K.); [email protected] (N.K.); [email protected] (F.V.); [email protected] (S.H.S.) 2 Department of Biomedical Sciences, University of Padua, Via U. Bassi 58/B, 35121 Padua, Italy * Correspondence: [email protected]; Tel.: +39-049-827-6300 Current Affiliation: Molecular Angiogenesis Laboratory, GIGA Research, University of Liège, B34, y Avenue de l’Hôpital, 1, 4000 Liège, Belgium. Received: 8 June 2020; Accepted: 2 July 2020; Published: 7 July 2020 Abstract: Contact sites between mitochondria and endoplasmic reticulum (ER) are points in which the two organelles are in close proximity. Due to their structural and functional complexity, their exploitation as pharmacological targets has never been considered so far. Notwithstanding, the number of compounds described to target proteins residing at these interfaces either directly or indirectly is rising. Here we provide original insight into mitochondria–ER contact sites (MERCs), with a comprehensive overview of the current MERCs pharmacology. Importantly, we discuss the considerable potential of MERCs to become a druggable target for the development of novel therapeutic strategies. Keywords: mitochondria–endoplasmic reticulum contact sites; mitochondria-associated membranes; pharmacology; drug targets; synthetic and biological compounds; neurodegeneration; diabetes; cancer 1. -

Arcachon and Cholinergic Transmission Victor P. Whittaker*
J Physiology (Paris) (1998) 92, 53-57 © Elsevier, Paris Arcachon and cholinergic transmission Victor P. Whittaker* Max-Planck-lnstitut fiir biophysikalische Chemic, D-37070 GOttingen, Germany Abstract -- The cholinergic nature of transmission at the electromotor synapse of Torpedo marmorata was established at Arcachon in 1939 by Feldberg, Fessard and Nachmansohn (J. Physiol. (Lond.) 97 (1939/1940) 3P-4P) soon after transmission at the neuromuscular junction had been shown to be cholinergic. In 1964, after a quarter of a century of neglect, workers in Cambridge, then in Paris, Gtittingen and elsewhere, began to use this system, 500-1000 times richer in cholinergic synapses than muscle, for intensive studies of cholinergic transmission at the cellular and molecular level. (@Elsevier, Paris) Resum6 -- Arcachon et la transmission cholinergique. La nature cholinergique de la transmission ~ la synapse 61ectromotrice de la torpille a 6t6 6tablie ~ Arcachon en 1939 par Feldberg, Fessard et Nachmansohn (J. Physiol. (Lond.) 97 (1939/1940) 3P-4P), peu aprrs la drcouverte du caractrre cholinergique de la transmission h la jonction neuromusculaire. Aprrs une prriode assez longue de d6su6tude, les chercheurs de Cambridge, Paris, Grttingen et d'ailleurs ont commencr, vers 1964, h utiliser ce systrme, 500-1000 fois plus riche en terminaisons cholinergiques que le muscle, pour 6tudier intensivement la transmission cholinergique au niveau cellulaire et molr- culaire. (@Elsevier, Paris) Torpedo marmorata I electric organ / electromotor synapse / cholinergic transmission 1. Introduction However, in terms of availability, amount of tissue per specimen and density of synaptic material, the The electric ray, Torpedo marmorata, is fairly Torpedinidae, especially T. marmorata (Eastern At- common in the Bay of Biscay (Baie de Gascoigne): lantic), T.