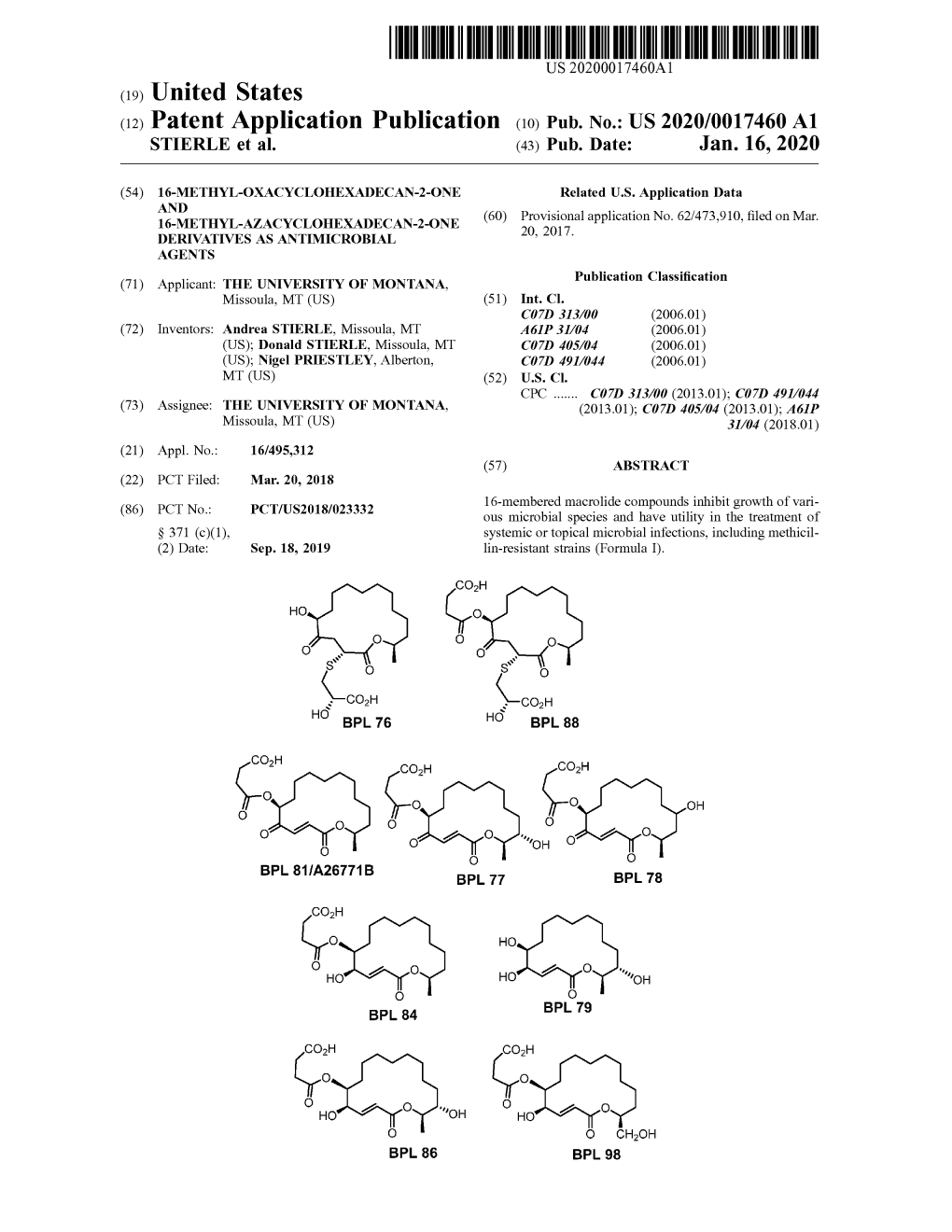
( 12 ) Patent Application Publication ( 10 ) Pub . No .: US 2020/0017460 A1
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Comparative Genomics of Staphylococcus Reveals Determinants of Speciation and Diversification of Antimicrobial Defense
1 Comparative Genomics of Staphylococcus Reveals Determinants of 2 Speciation and Diversification of Antimicrobial Defense. 3 4 5 Rosanna Coates-Brown1§, Josephine Moran1, Pisut Pongchaikul1¶, Alistair Darby1 and 6 Malcolm J. Horsburgh1* 7 8 9 10 11 12 1Institute of Integrative Biology, University of Liverpool, Liverpool, Merseyside, United 13 Kingdom. 14 15 § Present address: Genomic Diagnostic Laboratory, St Mary’s Hospital, Oxford Road, 16 Manchester, UK 17 ¶Present address: Faculty of Medicine Ramathibodi Hospital, Mahidol University, 270 18 Rama IV Road, Ratchathewi, Bangkok, 10400, Thailand 19 20 21 * Corresponding author: Institute of Integrative Biology, University of Liverpool, 22 Liverpool, L69 7ZB, United Kingdom. 23 Email: [email protected] 24 Tel: +44 1517954569 25 Fax +44 1517954410 26 Abstract 27 The bacterial genus Staphylococcus comprises diverse species with most being described 28 as colonizers of human and animal skin. A relational analysis of features that 29 discriminate its species and contribute to niche adaptation and survival remains to be fully 30 described. In this study, an interspecies, whole-genome comparative analysis of 21 31 Staphylococcus species was performed based on their orthologues. Three well-defined 32 multi-species groups were identified: group A (including aureus/epidermidis); group B 33 (including saprophyticus/xylosus) and group C (including pseudintermedius/delphini). 34 The machine learning algorithm Random Forest was applied to prioritise orthologues that 35 drive formation of the Staphylococcus species groups A-C. Orthologues driving 36 staphylococcal intrageneric diversity comprised regulatory, metabolic and antimicrobial 37 resistance proteins. Notably, the BraSR (NsaRS) two-component system (TCS) and its 38 associated BraDE transporters that regulate antimicrobial resistance showed limited 39 Distribution in the genus and their presence was most closely associated with a subset of 40 Staphylococcus species dominated by those that colonise human skin. -

Staphylococcus Edaphicus Sp
EVOLUTIONARY AND GENOMIC MICROBIOLOGY crossm Staphylococcus edaphicus sp. nov., Isolated in Antarctica, Harbors the mecC Gene and Genomic Islands with a Suspected Role in Adaptation to Extreme Environments Roman Pantu˚cˇek,a Ivo Sedlácˇek,b Adéla Indráková,a Veronika Vrbovská,a,b Ivana Mašlanˇová,a Vojteˇch Kovarˇovic,a Pavel Švec,b Stanislava Králová,b Lucie Krištofová,b Jana Kekláková,c Petr Petráš,c Jirˇí Doškarˇa aDivision of Genetics and Molecular Biology, Department of Experimental Biology, Faculty of Science, Masaryk University, Brno, Czech Republic bCzech Collection of Microorganisms, Department of Experimental Biology, Faculty of Science, Masaryk University, Brno, Czech Republic cReference Laboratory for Staphylococci, National Institute of Public Health, Prague, Czech Republic ABSTRACT Two Gram-stain-positive, coagulase-negative staphylococcal strains were isolated from abiotic sources comprising stone fragments and sandy soil in James Ross Island, Antarctica. Here, we describe properties of a novel species of the genus Staphylococcus that has a 16S rRNA gene sequence nearly identical to that of Staph- ylococcus saprophyticus. However, compared to S. saprophyticus and the next closest relatives, the new species demonstrates considerable phylogenetic distance at the whole-genome level, with an average nucleotide identity of Ͻ85% and inferred DNA-DNA hybridization of Ͻ30%. It forms a separate branch in the S. saprophyticus phylogenetic clade as confirmed by multilocus sequence analysis of six housekeep- ing genes, rpoB, hsp60, tuf, dnaJ, gap, and sod. Matrix-assisted laser desorption ion- ization–time of flight mass spectrometry (MALDI-TOF MS) and key biochemical charac- teristics allowed these bacteria to be distinguished from their nearest phylogenetic neighbors. In contrast to S. -

Intramammary Infections with Coagulase-Negative Staphylococcus Species
Printing of this thesis was financially supported by Printed by University Press, Zelzate ISBN number: 9789058642738 INTRAMAMMARY INFECTIONS WITH COAGULASE-NEGATIVE STAPHYLOCOCCUS SPECIES IN BOVINES - MOLECULAR DIAGNOSTICS AND EPIDEMIOLOGY - KARLIEN SUPRÉ 2011 PROMOTORS/PROMOTOREN Prof. dr. Sarne De Vliegher Faculteit Diergeneeskunde, UGent Prof. dr. Ruth N. Zadoks Royal (Dick) School of Veterinary Studies, University of Edinburgh; Moredun Research Institute, Penicuik, Schotland Prof. dr. Freddy Haesebrouck Faculteit Diergeneeskunde, UGent MEMBERS OF THE EXAMINATION COMMITTEE/LEDEN VAN DE EXAMENCOMMISSIE Prof. dr. dr. h. c. Aart de Kruif Voorzitter van de examencommissie Prof. dr. Mario Vaneechoutte Faculteit Geneeskunde en Gezondheidswetenschappen, UGent Dr. Margo Baele Directie Onderzoeksaangelegenheden, UGent Dr. Lic. Luc De Meulemeester MCC-Vlaanderen, Lier Prof. dr. Geert Opsomer Faculteit Diergeneeskunde, UGent Prof. dr. Marc Heyndrickx Instituut voor Landbouw en Visserijonderzoek (ILVO), Melle Dr. Suvi Taponen University of Helsinki, Finland Prof. dr. Ynte H. Schukken Cornell University, Ithaca, USA INTRAMAMMARY INFECTIONS WITH COAGULASE-NEGATIVE STAPHYLOCOCCUS SPECIES IN BOVINES - MOLECULAR DIAGNOSTICS AND EPIDEMIOLOGY - KARLIEN SUPRÉ Department of Reproduction, Obstetrics, and Herd Health Faculty of Veterinary Medicine, Ghent University Dissertation submitted in the fulfillment of the requirements for the degree of Doctor in Veterinary Sciences, Faculty of Veterinary Medicine, Ghent University INTRAMAMMAIRE INFECTIES MET COAGULASE-NEGATIEVE -

A Novel Approach to Eliminate Detection of Contaminating Staphylococcal Species Introduced During Clinical Testing
RESEARCH ARTICLE A novel approach to eliminate detection of contaminating Staphylococcal species introduced during clinical testing Wanyuan Ao, Adrianne Clifford, Maylene Corpuz, Robert Jenison* Great Basin Corporation, Salt Lake City, Utah, United States of America * [email protected] a1111111111 a1111111111 Abstract a1111111111 a1111111111 We describe here a strategy that can distinguish between Staphylococcus species truly a1111111111 present in a clinical sample from contaminating Staphylococcus species introduced during the testing process. Contaminating Staphylococcus species are present at low levels in PCR reagents and colonize lab personnel. To eliminate detection of contaminants, we describe an approach that utilizes addition of sufficient quantities of either non-target Staph- OPEN ACCESS ylococcal cells (Staphylococcus succinus or Staphylococcus muscae) or synthetic oligonu- Citation: Ao W, Clifford A, Corpuz M, Jenison R cleotide templates to helicase dependent isothermal amplification reactions to consume (2017) A novel approach to eliminate detection of Staphylococcus-specific tuf and mecA gene primers such that contaminating Staphylococ- contaminating Staphylococcal species introduced cus amplification is suppressed to below assay limits of detection. The suppressor template during clinical testing. PLoS ONE 12(2): e0171915. DNA is designed with perfect homology to the primers used in the assay but an internal doi:10.1371/journal.pone.0171915 sequence that is unrelated to the Staphylococcal species targeted for detection. Input Editor: Baochuan Lin, Defense Threat Reduction amount of the suppressor is determined by a mathematical model described herein and is Agency, UNITED STATES demonstrated to completely suppress contaminating levels of Staphylococcus while not Received: October 25, 2016 negatively impacting the appropriate clinical assay limit of detection. -

Table of Contents
An Investigation of Staphylococcus aureus and Related Species From Flood Affected and Other Environmental Sources A Thesis in Molecular Microbiology by Nadeesha Samanmalee Jayasundara BSc (Environmental Conservation & Management) School of Biomedical Science, Institute of Health & Biomedical Innovation Queensland University of Technology Brisbane, Australia Thesis submitted to Queensland University of Technology in fulfilment of the requirements for the degree of Masters of Applied Science (Research) May 2014 2 Abstract The genus Staphylococcus consists of 45 species and is widely distributed across environments such as skin and mucous membranes of humans and animals, as well as in soil, water and air. S. aureus and S. epidermidis are the most commonly associated species with human infections. Hence, most studies have focused on clinical and clinically sourced staphylococci. In addition, S. haemoliticus, S. intermidius, S. delphini, and S. saprophiticus are also considered potentially pathogenic members of the genus. Although staphylococci are distributed in various environments, there have been very few studies examining residential air as a reservoir of clinically significant pathogens, particularly Staphylococcus species. As a result, airborne transmission of staphylococci, and associated health risks, remains unclear. This study included not only residential air but also air samples from flood affected houses. Flood water can be considered as a potential carrier of pathogenic bacteria, because flood water can be affected by residential septic systems, municipal sanitary sewer systems, hospital waste, agricultural lands/operations and wastewater treatment plants. Even after the flood waters recede, microorganisms that are transported in water can remain in soil, in or on plant materials and on numerous other surfaces. Therefore, there is a great concern for use of previously flooded indoor and outdoor areas. -

Genetic Diversity of Methicillin Resistant Staphylococcus Aureus Strains in the Pretoria
Genetic diversity of methicillin resistant Staphylococcus aureus strains in the Pretoria region in South Africa ADEOLA MUJIDAT SALAWU Genetic diversity of methicillin resistant Staphylococcus aureus strains in the Pretoria region in South Africa by ADEOLA MUJIDAT SALAWU Submitted in partial fulfilment of the requirements for the degree MAGISTER SCIENTIAE MSc (Medical Microbiology) Department of Medical Microbiology Faculty of Health Sciences University of Pretoria Gauteng South Africa October 2013 Declaration I, the undersigned, declare that the dissertation hereby submitted to the University of Pretoria for the degree MSc (Medical Microbiology) and the work contained herein is my original work and has not previously, in its entirety or in part, been submitted to any university for a degree. I further declare that all sources cited are acknowledged by means of a list of references. Signed_________________this_________________day of_________________2014 Spending time with GOD is the key to our strength and success in all areas of life. Be sure that you never try to work GOD into your schedule, but always work your schedule around HIM Joyce Meyer Dedication To my dear husband (Babatunde Rotimi ): Thank you for your support, love and understanding ACKNOWLEDGEMENTS Firstly: I would like to extend my greatest gratitude to the almighty GOD, the creator of heaven and earth. My GOD, I will forever be grateful to You Secondly: I would like to sincerely thank the following individuals: Prof MM Ehlers (supervisor), Department of Medical Microbiology, University of Pretoria/NHLS, for her humility, endless motivation, patience, support and professional supervision in the successful completion of this research project. Prof, GOD bless you Dr MM Kock (co-supervisor), Department of Medical Microbiology, University of Pretoria/NHLS, for her guidance, understanding, support and molecular biology expertise regarding this research project. -

Posted 01/14
FINAL REPORT BioReD: Biomarkers and Tools for Reductive Dechlorination Site Assessment, Monitoring and Management SERDP Project ER-1586 November 2013 Frank Löffler Kirsti Ritalahti University of Tennessee Elizabeth Edwards University of Toronto Carmen Lebrón NAVFAC ESC Distribution Statement A This report was prepared under contract to the Department of Defense Strategic Environmental Research and Development Program (SERDP). The publication of this report does not indicate endorsement by the Department of Defense, nor should the contents be construed as reflecting the official policy or position of the Department of Defense. Reference herein to any specific commercial product, process, or service by trade name, trademark, manufacturer, or otherwise, does not necessarily constitute or imply its endorsement, recommendation, or favoring by the Department of Defense. Form Approved REPORT DOCUMENTATION PAGE OMB No. 0704-0188 Public reporting burden for this collection of information is estimated to average 1 hour per response, including the time for reviewing instructions, searching existing data sources, gathering and maintaining the data needed, and completing and reviewing this collection of information. Send comments regarding this burden estimate or any other aspect of this collection of information, including suggestions for reducing this burden to Department of Defense, Washington Headquarters Services, Directorate for Information Operations and Reports (0704-0188), 1215 Jefferson Davis Highway, Suite 1204, Arlington, VA 22202- 4302. Respondents should be aware that notwithstanding any other provision of law, no person shall be subject to any penalty for failing to comply with a collection of information if it does not display a currently valid OMB control number. PLEASE DO NOT RETURN YOUR FORM TO THE ABOVE ADDRESS. -

The Genera Staphylococcus and Macrococcus
Prokaryotes (2006) 4:5–75 DOI: 10.1007/0-387-30744-3_1 CHAPTER 1.2.1 ehT areneG succocolyhpatS dna succocorcMa The Genera Staphylococcus and Macrococcus FRIEDRICH GÖTZ, TAMMY BANNERMAN AND KARL-HEINZ SCHLEIFER Introduction zolidone (Baker, 1984). Comparative immu- nochemical studies of catalases (Schleifer, 1986), The name Staphylococcus (staphyle, bunch of DNA-DNA hybridization studies, DNA-rRNA grapes) was introduced by Ogston (1883) for the hybridization studies (Schleifer et al., 1979; Kilp- group micrococci causing inflammation and per et al., 1980), and comparative oligonucle- suppuration. He was the first to differentiate otide cataloguing of 16S rRNA (Ludwig et al., two kinds of pyogenic cocci: one arranged in 1981) clearly demonstrated the epigenetic and groups or masses was called “Staphylococcus” genetic difference of staphylococci and micro- and another arranged in chains was named cocci. Members of the genus Staphylococcus “Billroth’s Streptococcus.” A formal description form a coherent and well-defined group of of the genus Staphylococcus was provided by related species that is widely divergent from Rosenbach (1884). He divided the genus into the those of the genus Micrococcus. Until the early two species Staphylococcus aureus and S. albus. 1970s, the genus Staphylococcus consisted of Zopf (1885) placed the mass-forming staphylo- three species: the coagulase-positive species S. cocci and tetrad-forming micrococci in the genus aureus and the coagulase-negative species S. epi- Micrococcus. In 1886, the genus Staphylococcus dermidis and S. saprophyticus, but a deeper look was separated from Micrococcus by Flügge into the chemotaxonomic and genotypic proper- (1886). He differentiated the two genera mainly ties of staphylococci led to the description of on the basis of their action on gelatin and on many new staphylococcal species. -

1 SUPPLEMENTARY INFORMATION Captive Bottlenose Dolphins And
SUPPLEMENTARY INFORMATION Captive bottlenose dolphins and killer whales harbor a species-specific skin microbiota that varies among individuals Chiarello M., Villéger S., Bouvier C., Auguet JC., and Bouvier T. 1 Supplementary Information S1: Description of the two PCR protocols used in this study and comparison of bacterial composition on water samples Skin samples Water samples Kit Phusion High-Fidelity PuRe Taq Ready-To-Go PCR Beads Total vol. (µL) 20 25 DNA vol. (µL) 2 5 Initial denaturation 1 min 98°C 2 min 94°C PCR cycle 1 min 94°C; 40s 57.8°C; 30s 72°C 1 min 94°C; 40s 57.8°C; 30s 72°C Nb. of cycles 35 35 Final extension 10 min 72°C 10 min 72°C S1-Table 1: PCR reagents and conditions used for the two sample types studied. Skin DNA and water DNA were respectively amplified using the Phusion High-Fidelity DNA polymerase (Biolabs, Ipswich, USA) and PuRe Taq Ready-To-Go PCR Beads (Amersham Biosciences, Freiburg, Germany) following manufacturer’s instructions. 2 S1-Fig 1: Most abundant classes and families in planktonic communities analyzed using Phusion and Ready-To-Go kits. Both PCR types were performed on the same DNA extracted from animals’ surrounding water. Class-level bacterial composition was very similar between both PCR types. 3 S1-Fig 2: PCoAs based on Weighted Unifrac, showing planktonic communities analyzed using both PCR types. On (A) panel, all samples included in this study plus water replicates that could be amplified using Phusion kit. On (B) panel, only planktonic communities were displayed. -

Wall Teichoic Acid Structure Governs Horizontal Gene Transfer Between Major Bacterial Pathogens
ARTICLE Received 28 Jan 2013 | Accepted 22 Jul 2013 | Published 22 Aug 2013 DOI: 10.1038/ncomms3345 OPEN Wall teichoic acid structure governs horizontal gene transfer between major bacterial pathogens Volker Winstel1,2, Chunguang Liang3, Patricia Sanchez-Carballo4, Matthias Steglich5, Marta Munar3,w, Barbara M. Bro¨ker6, Jose R. Penade´s7, Ulrich Nu¨bel5, Otto Holst4, Thomas Dandekar3, Andreas Peschel1,2 & Guoqing Xia1,2 Mobile genetic elements (MGEs) encoding virulence and resistance genes are widespread in bacterial pathogens, but it has remained unclear how they occasionally jump to new host species. Staphylococcus aureus clones exchange MGEs such as S. aureus pathogenicity islands (SaPIs) with high frequency via helper phages. Here we report that the S. aureus ST395 lineage is refractory to horizontal gene transfer (HGT) with typical S. aureus but exchanges SaPIs with other species and genera including Staphylococcus epidermidis and Listeria mono- cytogenes. ST395 produces an unusual wall teichoic acid (WTA) resembling that of its HGT partner species. Notably, distantly related bacterial species and genera undergo efficient HGT with typical S. aureus upon ectopic expression of S. aureus WTA. Combined with genomic analyses, these results indicate that a ‘glycocode’ of WTA structures and WTA-binding helper phages permits HGT even across long phylogenetic distances thereby shaping the evolution of Gram-positive pathogens. 1 Cellular and Molecular Microbiology Division, Interfaculty Institute of Microbiology and Infection Medicine, University of Tu¨bingen, Elfriede-Aulhorn-Strae 6, 72076 Tu¨bingen, Germany. 2 German Center for Infection Research (DZIF), partner site Tu¨bingen, 72076 Tu¨bingen, Germany. 3 Bioinformatik, Biozentrum, University of Wu¨rzburg, Am Hubland, 97074 Wu¨rzburg, Germany. -

Two-Component Signal Transduction System Saers Positively Regulates Staphylococcus Epidermidis Glucose Metabolism
Hindawi Publishing Corporation e Scientific World Journal Volume 2014, Article ID 908121, 12 pages http://dx.doi.org/10.1155/2014/908121 Research Article Two-Component Signal Transduction System SaeRS Positively Regulates Staphylococcus epidermidis Glucose Metabolism Qiang Lou,1 Yijun Qi,1 Yuanfang Ma,1 and Di Qu2 1 Laboratory of Cellular and Molecular Immunology, Henan University, Kaifeng 475004, China 2 Key laboratory of Medical Molecular Virology of Ministry of Education and Ministry of Public Health, Institute of Medical Microbiology and Institutes of Biomedical Sciences, Shanghai Medical College of Fudan University, 138 Yixueyuan Road, Shanghai, 200032, China Correspondence should be addressed to Yuanfang Ma; [email protected] and Di Qu; [email protected] Received 30 August 2013; Accepted 21 November 2013; Published 23 January 2014 Academic Editors: D. Maiorano, H. Okamura, and M. Shiraishi Copyright © 2014 Qiang Lou et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Staphylococcus epidermidis, which is a causative pathogen of nosocomial infection, expresses its virulent traits such as biofilm and autolysis regulated by two-component signal transduction system SaeRS. In this study, we performed a proteomic analysis of differences in expression between the S. epidermidis 1457 wild-type and saeRS mutant to identify candidates regulated by saeRS using two-dimensional gel electrophoresis (2-DE) combined with matrix-assisted laser desorption/lonization mass spectrometry (MALDI-TOF-MS). Of 55 identified proteins that significantly differed in expression between the two strains, 15 were upregulated and 40 were downregulated. -

Comparative Genomic Analysis of the Genus Staphylococcus Including
Suzuki et al. BMC Genomics 2012, 13:38 http://www.biomedcentral.com/1471-2164/13/38 RESEARCH ARTICLE Open Access Comparative genomic analysis of the genus Staphylococcus including Staphylococcus aureus and its newly described sister species Staphylococcus simiae Haruo Suzuki1, Tristan Lefébure1,2, Paulina Pavinski Bitar1 and Michael J Stanhope1* Abstract Background: Staphylococcus belongs to the Gram-positive low G + C content group of the Firmicutes division of bacteria. Staphylococcus aureus is an important human and veterinary pathogen that causes a broad spectrum of diseases, and has developed important multidrug resistant forms such as methicillin-resistant S. aureus (MRSA). Staphylococcus simiae was isolated from South American squirrel monkeys in 2000, and is a coagulase-negative bacterium, closely related, and possibly the sister group, to S. aureus. Comparative genomic analyses of closely related bacteria with different phenotypes can provide information relevant to understanding adaptation to host environment and mechanisms of pathogenicity. Results: We determined a Roche/454 draft genome sequence for S. simiae and included it in comparative genomic analyses with 11 other Staphylococcus species including S. aureus. A genome based phylogeny of the genus confirms that S. simiae is the sister group to S. aureus and indicates that the most basal Staphylococcus lineage is Staphylococcus pseudintermedius, followed by Staphylococcus carnosus. Given the primary niche of these two latter taxa, compared to the other species in the genus, this phylogeny suggests that human adaptation evolved after the split of S. carnosus. The two coagulase-positive species (S. aureus and S. pseudintermedius) are not phylogenetically closest but share many virulence factors exclusively, suggesting that these genes were acquired by horizontal transfer.