Estimating Drivers of Cell State Transitions Using Gene Regulatory Network Models
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Identification of GA-Binding Protein Transcription Factor Alpha Subunit
International Journal of Molecular Sciences Article Identification of GA-Binding Protein Transcription Factor Alpha Subunit (GABPA) as a Novel Bookmarking Factor Shunya Goto 1, Masashi Takahashi 1, Narumi Yasutsune 1, Sumiki Inayama 1, Dai Kato 2, Masashi Fukuoka 3, Shu-ichiro Kashiwaba 1 and Yasufumi Murakami 1,2,* 1 Department of Biological Science and Technology, Faculty of Industrial Science and Technology, Tokyo University of Science, 6-3-1 Niijuku, Katsushika-ku, Tokyo 125-8585, Japan; [email protected] (S.G.); [email protected] (M.T.); [email protected] (N.Y.); [email protected] (S.I.); [email protected] (S.K.) 2 Order-MadeMedical Research Inc., 208Todai-Kashiwa VP, 5-4-19 Kashiwanoha, Kashiwa-shi, Chiba-ken 277-0882, Japan; [email protected] 3 Department of Molecular Pharmacology, National Institute of Neuroscience, National Center of Neurology and Psychiatry, Tokyo 187-8551, Japan; [email protected] * Correspondence: [email protected]; Tel.: +81-3-5876-1717 (ext. 1919); Fax: +81-3-5876-1470 Received: 7 February 2019; Accepted: 27 February 2019; Published: 4 March 2019 Abstract: Mitotic bookmarking constitutes a mechanism for transmitting transcriptional patterns through cell division. Bookmarking factors, comprising a subset of transcription factors (TFs), and multiple histone modifications retained in mitotic chromatin facilitate reactivation of transcription in the early G1 phase. However, the specific TFs that act as bookmarking factors remain largely unknown. Previously, we identified the “early G1 genes” and screened TFs that were predicted to bind to the upstream region of these genes, then identified GA-binding protein transcription factor alpha subunit (GABPA) and Sp1 transcription factor (SP1) as candidate bookmarking factors. -

GABPA Is a Master Regulator of Luminal Identity and Restrains Aggressive Diseases in Bladder Cancer
Cell Death & Differentiation (2020) 27:1862–1877 https://doi.org/10.1038/s41418-019-0466-7 ARTICLE GABPA is a master regulator of luminal identity and restrains aggressive diseases in bladder cancer 1,2,3 3,4 5 2,5 5 3 5 5 Yanxia Guo ● Xiaotian Yuan ● Kailin Li ● Mingkai Dai ● Lu Zhang ● Yujiao Wu ● Chao Sun ● Yuan Chen ● 5 6 3 1,2 1,2 3,7 Guanghui Cheng ● Cheng Liu ● Klas Strååt ● Feng Kong ● Shengtian Zhao ● Magnus Bjorkhölm ● Dawei Xu 3,7 Received: 3 June 2019 / Revised: 20 November 2019 / Accepted: 21 November 2019 / Published online: 4 December 2019 © The Author(s) 2019. This article is published with open access Abstract TERT promoter mutations occur in the majority of glioblastoma, bladder cancer (BC), and other malignancies while the ETS family transcription factors GABPA and its partner GABPB1 activate the mutant TERT promoter and telomerase in these tumors. GABPA depletion or the disruption of the GABPA/GABPB1 complex by knocking down GABPB1 was shown to inhibit telomerase, thereby eliminating the tumorigenic potential of glioblastoma cells. GABPA/B1 is thus suggested as a cancer therapeutic target. However, it is unclear about its role in BC. Here we unexpectedly observed that GABPA ablation 1234567890();,: 1234567890();,: inhibited TERT expression, but robustly increased proliferation, stem, and invasive phenotypes and cisplatin resistance in BC cells, while its overexpression exhibited opposite effects, and inhibited in vivo metastasizing in a xenograft transplant model. Mechanistically, GABPA directly activates the transcription of FoxA1 and GATA3, key transcription factors driving luminal differentiation of urothelial cells. Consistently, TCGA/GEO dataset analyses show that GABPA expression is correlated positively with luminal while negatively with basal signatures. -

Distinct Contributions of DNA Methylation and Histone Acetylation to the Genomic Occupancy of Transcription Factors
Downloaded from genome.cshlp.org on October 8, 2021 - Published by Cold Spring Harbor Laboratory Press Research Distinct contributions of DNA methylation and histone acetylation to the genomic occupancy of transcription factors Martin Cusack,1 Hamish W. King,2 Paolo Spingardi,1 Benedikt M. Kessler,3 Robert J. Klose,2 and Skirmantas Kriaucionis1 1Ludwig Institute for Cancer Research, University of Oxford, Oxford, OX3 7DQ, United Kingdom; 2Department of Biochemistry, University of Oxford, Oxford, OX1 3QU, United Kingdom; 3Target Discovery Institute, University of Oxford, Oxford, OX3 7FZ, United Kingdom Epigenetic modifications on chromatin play important roles in regulating gene expression. Although chromatin states are often governed by multilayered structure, how individual pathways contribute to gene expression remains poorly under- stood. For example, DNA methylation is known to regulate transcription factor binding but also to recruit methyl-CpG binding proteins that affect chromatin structure through the activity of histone deacetylase complexes (HDACs). Both of these mechanisms can potentially affect gene expression, but the importance of each, and whether these activities are inte- grated to achieve appropriate gene regulation, remains largely unknown. To address this important question, we measured gene expression, chromatin accessibility, and transcription factor occupancy in wild-type or DNA methylation-deficient mouse embryonic stem cells following HDAC inhibition. We observe widespread increases in chromatin accessibility at ret- rotransposons when HDACs are inhibited, and this is magnified when cells also lack DNA methylation. A subset of these elements has elevated binding of the YY1 and GABPA transcription factors and increased expression. The pronounced ad- ditive effect of HDAC inhibition in DNA methylation–deficient cells demonstrates that DNA methylation and histone deacetylation act largely independently to suppress transcription factor binding and gene expression. -

Ten Commandments for a Good Scientist
Unravelling the mechanism of differential biological responses induced by food-borne xeno- and phyto-estrogenic compounds Ana María Sotoca Covaleda Wageningen 2010 Thesis committee Thesis supervisors Prof. dr. ir. Ivonne M.C.M. Rietjens Professor of Toxicology Wageningen University Prof. dr. Albertinka J. Murk Personal chair at the sub-department of Toxicology Wageningen University Thesis co-supervisor Dr. ir. Jacques J.M. Vervoort Associate professor at the Laboratory of Biochemistry Wageningen University Other members Prof. dr. Michael R. Muller, Wageningen University Prof. dr. ir. Huub F.J. Savelkoul, Wageningen University Prof. dr. Everardus J. van Zoelen, Radboud University Nijmegen Dr. ir. Toine F.H. Bovee, RIKILT, Wageningen This research was conducted under the auspices of the Graduate School VLAG Unravelling the mechanism of differential biological responses induced by food-borne xeno- and phyto-estrogenic compounds Ana María Sotoca Covaleda Thesis submitted in fulfillment of the requirements for the degree of doctor at Wageningen University by the authority of the Rector Magnificus Prof. dr. M.J. Kropff, in the presence of the Thesis Committee appointed by the Academic Board to be defended in public on Tuesday 14 September 2010 at 4 p.m. in the Aula Unravelling the mechanism of differential biological responses induced by food-borne xeno- and phyto-estrogenic compounds. Ana María Sotoca Covaleda Thesis Wageningen University, Wageningen, The Netherlands, 2010, With references, and with summary in Dutch. ISBN: 978-90-8585-707-5 “Caminante no hay camino, se hace camino al andar. Al andar se hace camino, y al volver la vista atrás se ve la senda que nunca se ha de volver a pisar” - Antonio Machado – A mi madre. -

Accompanies CD8 T Cell Effector Function Global DNA Methylation
Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer, Benjamin G. Barwick, Benjamin A. Youngblood, Rafi Ahmed and Jeremy M. Boss This information is current as of October 1, 2021. J Immunol 2013; 191:3419-3429; Prepublished online 16 August 2013; doi: 10.4049/jimmunol.1301395 http://www.jimmunol.org/content/191/6/3419 Downloaded from Supplementary http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130139 Material 5.DC1 References This article cites 81 articles, 25 of which you can access for free at: http://www.jimmunol.org/content/191/6/3419.full#ref-list-1 http://www.jimmunol.org/ Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists by guest on October 1, 2021 • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Global DNA Methylation Remodeling Accompanies CD8 T Cell Effector Function Christopher D. Scharer,* Benjamin G. Barwick,* Benjamin A. Youngblood,*,† Rafi Ahmed,*,† and Jeremy M. -

Estrogen-Related Receptor Α (Errα) : a Novel Target in Type 2 Diabetes
Institutional Repository of the University of Basel University Library Schoenbeinstrasse 18-20 CH-4056 Basel, Switzerland http://edoc.unibas.ch/ Year: 2005 Estrogen-related receptor α (ERRα) : a novel target in type 2 diabetes Handschin, C. and Mootha, V. K. Posted at edoc, University of Basel Official URL: http://edoc.unibas.ch/dok/A5258721 Originally published as: Handschin, C. and Mootha, V. K.. (2005) Estrogen-related receptor α (ERRα) : a novel target in type 2 diabetes. Drug discovery today. Therapeutic strategies, Vol. 2, H. 2. S. 151-156. Estrogen-related receptor (ERR): a novel target in type 2 diabetes Christoph Handschin1,3 and Vamsi K. Mootha2 1Dana-Farber Cancer Institute and Department of Cell Biology, Harvard Medical School, One Jimmy Fund Way, Boston, MA 02115, USA 2Departments of Systems Biology and of Medicine, Harvard Medical School, Broad Institute of Harvard and MIT, Cambridge, MA 02139, USA Published in Drug Discovery Today: Therapeutic Strategies 2005 Volume 2, Issue 2, Pages 93-176. DOI: 10.1016/j.ddstr.2005.05.001 Copyright © Elsevier, Drug Discovery Today: Therapeutic Strategies - 1 - Estrogen-related receptor (ERR): a novel target in type 2 diabetes Christoph Handschin1,3 and Vamsi K. Mootha2 1Dana-Farber Cancer Institute and Department of Cell Biology, Harvard Medical School, One Jimmy Fund Way, Boston, MA 02115, USA 2Departments of Systems Biology and of Medicine, Harvard Medical School, Broad Institute of Harvard and MIT, Cambridge, MA 02139, USA 3Correspondence: Phone: +1 617 632 3305; Fax: +1 617 632 5363; Email: [email protected] (Christoph Handschin) - 2 - Abstract Recent studies have shown that reduced mitochondrial content and function in skeletal muscle are common features of type 2 diabetes. -

Supplemental Table 1. Primers and Probes for RT-Pcrs
Supplemental Table 1. Primers and probes for RT-PCRs Gene Direction Sequence Quantitative RT-PCR E2F1 Forward Primer 5’-GGA TTT CAC ACC TTT TCC TGG AT-3’ Reverse Primer 5’-CCT GGA AAC TGA CCA TCA GTA CCT-3’ Probe 5’-FAM-CGA GCT GGC CCA CTG CTC TCG-TAMRA-3' E2F2 Forward Primer 5'-TCC CAA TCC CCT CCA GAT C-3' Reverse Primer 5'-CAA GTT GTG CGA TGC CTG C-3' Probe 5' -FAM-TCC TTT TGG CCG GCA GCC G-TAMRA-3' E2F3a Forward Primer 5’-TTT AAA CCA TCT GAG AGG TAC TGA TGA-3’ Reverse Primer 5’-CGG CCC TCC GGC AA-3’ Probe 5’-FAM-CGC TTT CTC CTA GCT CCA GCC TTC G-TAMRA-3’ E2F3b Forward Primer 5’-TTT AAA CCA TCT GAG AGG TAC TGA TGA-3’ Reverse Primer 5’-CCC TTA CAG CAG CAG GCA A-3’ Probe 5’-FAM-CGC TTT CTC CTA GCT CCA GCC TTC G-TAMRA-3’ IRF-1 Forward Primer 5’-TTT GTA TCG GCC TGT GTG AAT G-3’ Reverse Primer 5’-AAG CAT GGC TGG GAC ATC A-3’ Probe 5’-FAM-CAG CTC CGG AAC AAA CAG GCA TCC TT-TAMRA-3' IRF-2 Forward Primer 5'-CGC CCC TCG GCA CTC T-3' Reverse Primer 5'-TCT TCC TAT GCA GAA AGC GAA AC-3' Probe 5'-FAM-TTC ATC GCT GGG CAC ACT ATC AGT-TAMRA-3' TBP Forward Primer 5’-CAC GAA CCA CGG CAC TGA TT-3’ Reverse Primer 5’-TTT TCT TGC TGC CAG TCT GGA C-3’ Probe 5’-FAM-TGT GCA CAG GAG CCA AGA GTG AAG A-BHQ1-3’ Primers and Probes for quantitative RT-PCRs were designed using the computer program “Primer Express” (Applied Biosystems, Foster City, CA, USA). -

TFARM: Transcription Factor Associatio Rule Miner
TFARM: Transcription Factor Associatio Rule Miner Liuba Nausicaa Martino [email protected] Alice Parodi Gaia Ceddia Piercesare Secchi Stefano Campaner Marco Masseroli May 19, 2021 Contents 1 Introduction ..............................1 2 Dataset ................................2 3 Extraction of the most relevant associations ...........4 4 Importance Index of a transcription factor .............7 4.1 Validation of the Importance Index formula ............ 14 5 Visualization tools .......................... 17 library(TFARM) 1 Introduction Looking for association rules between transcription factors in genomic regions of interest can be useful to find direct or indirect interactions among regulatory factors of DNA transcription. However, the results provided by the most recent algorithms for the search of association rules [1][2] alone are often not intelligible enough, since they only provide a list of association rules. A novel method is proposed for subsequent mining of these results to evaluate the contribution of the items in each association rule. The TFARM package allows us to identify and extract the most relevant association rules with a given target transcription factor and compute the Importance Index of a transcription factor (or a combination of some of them) in the extracted rules. Such an index is useful to associate a numerical value to the contribution of one or more transcription factors to the co-regulation with a given target transcription factor. TFARM: Transcription Factor Associatio Rule Miner 2 Dataset Association rules are extracted from a GRanges object in which metadata columns identify transcription factors and genomic coordinates are represented in the left-hand-side of the GRanges; thus, each row is a different genomic region. -

Duodenal Mucosal Mitochondrial Gene Expression Is Associated with Delayed Gastric Emptying in Diabetic Gastroenteropathy
Duodenal mucosal mitochondrial gene expression is associated with delayed gastric emptying in diabetic gastroenteropathy Susrutha Puthanmadhom Narayanan, … , Tamas Ordog, Adil E. Bharucha JCI Insight. 2021;6(2):e143596. https://doi.org/10.1172/jci.insight.143596. Research Article Endocrinology Gastroenterology Graphical abstract Find the latest version: https://jci.me/143596/pdf RESEARCH ARTICLE Duodenal mucosal mitochondrial gene expression is associated with delayed gastric emptying in diabetic gastroenteropathy Susrutha Puthanmadhom Narayanan,1 Daniel O’Brien,2 Mayank Sharma,1 Karl Miller,3 Peter Adams,3 João F. Passos,4 Alfonso Eirin,5 Tamas Ordog,4 and Adil E. Bharucha1 1Division of Gastroenterology and Hepatology, Department of Medicine, Mayo Clinic, Rochester, Minnesota, USA. 2Department of Biomedical Statistics and Informatics, Mayo Clinic, Rochester, Minnesota, USA. 3Sanford Burnham Prebys Medical Discovery Institute, San Diego, California, USA. 4Department of Physiology and Biomedical Engineering and 5Division of Nephrology & Hypertension Research, Department of Medicine, Mayo Clinic, Rochester, Minnesota, USA. Hindered by a limited understanding of the mechanisms responsible for diabetic gastroenteropathy (DGE), management is symptomatic. We investigated the duodenal mucosal expression of protein- coding genes and microRNAs (miRNA) in DGE and related them to clinical features. The diabetic phenotype, gastric emptying, mRNA, and miRNA expression and ultrastructure of duodenal mucosal biopsies were compared in 39 DGE patients and 21 controls. Among 3175 differentially expressed genes (FDR < 0.05), several mitochondrial DNA–encoded (mtDNA-encoded) genes (12 of 13 protein coding genes involved in oxidative phosphorylation [OXPHOS], both rRNAs and 9 of 22 transfer RNAs) were downregulated; conversely, nuclear DNA–encoded (nDNA-encoded) mitochondrial genes (OXPHOS) were upregulated in DGE. -

NOTCH1–RBPJ Complexes Drive Target Gene Expression Through Dynamic Interactions with Superenhancers
NOTCH1–RBPJ complexes drive target gene expression through dynamic interactions with superenhancers Hongfang Wanga,1, Chongzhi Zangb,1, Len Taingb, Kelly L. Arnettc, Yinling Joey Wonga, Warren S. Peard, Stephen C. Blacklowc, X. Shirley Liub,2, and Jon C. Astera,2 aDepartment of Pathology, Brigham and Women’s Hospital, Boston, MA 02115; bDepartment of Biostatistics and Computational Biology, Dana-Farber Cancer Institute, Harvard School of Public Health, Boston, MA 02215; cDepartment of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02215; and dDepartment of Pathology, Perelman School of Medicine, University of Pennsylvania, Philadelphia, PA 19104 Edited by Richard A. Flavell, Yale School of Medicine and Howard Hughes Medical Institute, New Haven, CT, and approved November 27, 2013 (received for review August 8, 2013) The main oncogenic driver in T-lymphoblastic leukemia is NOTCH1, and that the H3K27ac mark is characteristic of active enhancers which activates genes by forming chromatin-associated Notch and correlates with transcription activation (6–9). transcription complexes. Gamma-secretase-inhibitor treatment pre- In cancers such as T-LL, gain-of-function mutations in NOTCH1 vents NOTCH1 nuclear localization, but most genes with NOTCH1- cause excessive Notch activation and exaggerated expression of binding sites are insensitive to gamma-secretase inhibitors. Here, oncogenic target genes. To further elucidate how NOTCH1 reg- we demonstrate that fewer than 10% of NOTCH1-binding sites ulates the transcriptomes of T-LL cells, we recently used chromatin show dynamic changes in NOTCH1 occupancy when T-lympho- immunoprecipitation (ChIP)-Seq to identify RBPJ–NOTCH1- blastic leukemia cells are toggled between the Notch-on and -off binding sites genomewide in Notch-“addicted” murine and hu- states with gamma-secretase inhibiters. -
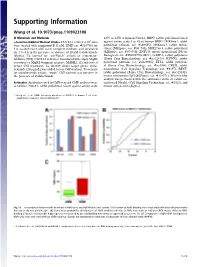
Supporting Information
Supporting Information Wang et al. 10.1073/pnas.1109023108 SI Materials and Methods 2278 to 2470 of human Notch1; RBPJ, rabbit polyclonal raised γ-Secretase-Inhibitor Washout Studies. CUTLL1 cells (2 × 105/mL) against amino acids 1 to 48 of human RBPJ; H3K4me1, rabbit were treated with compound E (1 μM, EMD cat. #565790) for polyclonal (Abcam, cat. #ab8895); H3K4me3, rabbit mono- 3 d, washed twice with warm complete medium, and incubated clonal (Millipore, cat. #04–745); H3K27me3, rabbit polyclonal for 2 to 4 h in the presence or absence of 20 μM cycloheximide (Millipore, cat. #07–449); ZNF143, mouse monoclonal (Novus (Sigma). To control for “off-Notch” effects of γ-secretase- Biologicals, cat. #H00007702-M01); GABPA, rabbit polyclonal inhibitor (GSI), CUTLL1 cells were transduced with empty MigRI (Santa Cruz Biotechnology, cat. #sc-22810); RUNX1, rabbit retrovirus or MigRI-dominant negative MAML1 (1) and sorted polyclonal (Abcam, cat. #ab23980); ETS1, rabbit polyclon- before GSI treatment. To identify direct target genes, cyclo- al (Santa Cruz Biotechnology, cat. #sc-350); CREB, rabbit heximide (20 μg/mL) was added before GSI washout. To control monoclonal (Cell Signaling Technology, cat. #9197); REST, for cycloheximide effects, “mock” GSI washout was perform in rabbit polyclonal (Santa Cruz Biotechnology, cat. #sc-25398); the presence of cycloheximide. mouse nonimmune IgG (Millipore, cat. #12–371). Western blot analysis was performed with the antibodies above or rabbit an- Antibodies. Antibodies used in ChIP-seq and ChIP analyses were tiactivated Notch1 (Cell Signaling Technology, cat. #2412), and as follows: Notch1, rabbit polyclonal raised against amino acids mouse anti–β-actin (Sigma). -

Sequence and Chromatin Determinants of Cell-Type Specific
Sequence and chromatin determinants of cell-type specific transcription factor binding: supplementary data Aaron Arvey1, Phaedra Agius1, William Stafford Noble2, and Christina Leslie1∗ 1Computational Biology Program, Memorial Sloan-Kettering Cancer Center, New York, NY 2Department of Genome Sciences, University of Washington, Seattle, WA March 15, 2012 Table S1: List of all TF ChIP-seq experiments analyzed in the study. File name Cell TF wgEncodeYaleChIPseqRawDataRep1Helas3Ap2alpha Helas3 AP2A1 wgEncodeYaleChIPseqRawDataRep2Helas3Ap2alpha Helas3 AP2A1 wgEncodeYaleChIPseqRawDataRep1Helas3Ap2gamma Helas3 TFAP2C wgEncodeYaleChIPseqRawDataRep2Helas3Ap2gamma Helas3 TFAP2C wgEncodeYaleChIPseqRawDataRep1K562Atf3 K562 ATF3 wgEncodeYaleChIPseqRawDataRep2K562Atf3 K562 ATF3 wgEncodeYaleChIPseqRawDataRep1Helas3Baf155Musigg Helas3 SMARCC1 wgEncodeYaleChIPseqRawDataRep2Helas3Baf155Musigg Helas3 SMARCC1 wgEncodeYaleChIPseqRawDataRep1Helas3Baf170Musigg Helas3 SMARCC2 wgEncodeHudsonalphaChipSeqRawDataRep1Gm12878Batf Gm12878 BATF wgEncodeHudsonalphaChipSeqRawDataRep2Gm12878Batf Gm12878 BATF wgEncodeHudsonalphaChipSeqRawDataRep1Gm12878Bcl11a Gm12878 BCL11A wgEncodeHudsonalphaChipSeqRawDataRep2Gm12878Bcl11a Gm12878 BCL11A wgEncodeHudsonalphaChipSeqRawDataRep1Gm12878Bcl3Pcr1xBcl3 Gm12878 BCL3 wgEncodeHudsonalphaChipSeqRawDataRep2Gm12878Bcl3Pcr1xBcl3 Gm12878 BCL3 wgEncodeYaleChIPseqRawDataRep1Helas3Bdp1 Helas3 BDP1 wgEncodeYaleChIPseqRawDataRep2Helas3Bdp1 Helas3 BDP1 wgEncodeYaleChIPseqRawDataRep1K562Bdp1 K562 BDP1 wgEncodeYaleChIPseqRawDataRep2K562Bdp1 K562