EXOC1 Plays an Integral Role in Spermatogonia Pseudopod
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Identification of Differentially Expressed Genes of Primary
Cell Research (2004); 14(6):507-512 ARTICLE http://www.cell-research.com Identification of differentially expressed genes of primary spermatocyte against round spermatid isolated from human testis using the laser capture microdissection technique Gang LIANG1,4, Xiao Dong ZHANG1, Lu Jing WANG1, Yu Shen SHA2, Jian Chao ZHANG2, Shi Ying MIAO1, Shu Dong ZONG2, Lin Fang WANG1,*, S.S. KOIDE3 1National Laboratory Medical Molecular Biology, Institute of Basic Medical Sciences, Chinese Academy of Medical Sciences, Peking Union Medical College, 5 Dong Dan San Tiao, 100005 Beijing, China 2National Research Institute for Family Planning, WHO Collaboration Center for Research in Human Reproduction, Beijing, 12 Da Hui Si, 100081 Beijing, China 3Center for Biomedical Research, Population Council, 1230 York Avenue, New York, NY 10021, USA 4Chinese National Human Genome Center, Beijing, 3-707 North Yong Chang Road BDA, Beijing 100176, China ABSTRACT The method of laser capture microdissection (LCM) combined with suppressive subtractive hybridization (SSH) was developed to isolate specific germ cells from human testis sections and to identify the genes expressed during differen- tiation and development. In the present study, over 10,000 primary spermatocytes and round spermatid cells were successfully isolated by LCM. Using the cDNAs from primary spermatocytes and round spermatids, SSH cDNAs library of primary spermatocyte-specific was constructed. The average insert size of the cDNA isolated from 75 randomly picked white clones was 500 bp, ranging from 250 bp to 1.7 kb. Using the dot-blot method, a total of 421 clones were examined, resulting in the identification of 390 positive clones emitting strong signals. -

Protein Interaction Network of Alternatively Spliced Isoforms from Brain Links Genetic Risk Factors for Autism
ARTICLE Received 24 Aug 2013 | Accepted 14 Mar 2014 | Published 11 Apr 2014 DOI: 10.1038/ncomms4650 OPEN Protein interaction network of alternatively spliced isoforms from brain links genetic risk factors for autism Roser Corominas1,*, Xinping Yang2,3,*, Guan Ning Lin1,*, Shuli Kang1,*, Yun Shen2,3, Lila Ghamsari2,3,w, Martin Broly2,3, Maria Rodriguez2,3, Stanley Tam2,3, Shelly A. Trigg2,3,w, Changyu Fan2,3, Song Yi2,3, Murat Tasan4, Irma Lemmens5, Xingyan Kuang6, Nan Zhao6, Dheeraj Malhotra7, Jacob J. Michaelson7,w, Vladimir Vacic8, Michael A. Calderwood2,3, Frederick P. Roth2,3,4, Jan Tavernier5, Steve Horvath9, Kourosh Salehi-Ashtiani2,3,w, Dmitry Korkin6, Jonathan Sebat7, David E. Hill2,3, Tong Hao2,3, Marc Vidal2,3 & Lilia M. Iakoucheva1 Increased risk for autism spectrum disorders (ASD) is attributed to hundreds of genetic loci. The convergence of ASD variants have been investigated using various approaches, including protein interactions extracted from the published literature. However, these datasets are frequently incomplete, carry biases and are limited to interactions of a single splicing isoform, which may not be expressed in the disease-relevant tissue. Here we introduce a new interactome mapping approach by experimentally identifying interactions between brain-expressed alternatively spliced variants of ASD risk factors. The Autism Spliceform Interaction Network reveals that almost half of the detected interactions and about 30% of the newly identified interacting partners represent contribution from splicing variants, emphasizing the importance of isoform networks. Isoform interactions greatly contribute to establishing direct physical connections between proteins from the de novo autism CNVs. Our findings demonstrate the critical role of spliceform networks for translating genetic knowledge into a better understanding of human diseases. -

5850.Full.Pdf
Soluble NSF Attachment Protein Receptors (SNAREs) in RBL-2H3 Mast Cells: Functional Role of Syntaxin 4 in Exocytosis and Identification of a Vesicle-Associated This information is current as Membrane Protein 8-Containing Secretory of September 25, 2021. Compartment Fabienne Paumet, Joëlle Le Mao, Sophie Martin, Thierry Galli, Bernard David, Ulrich Blank and Michèle Roa Downloaded from J Immunol 2000; 164:5850-5857; ; doi: 10.4049/jimmunol.164.11.5850 http://www.jimmunol.org/content/164/11/5850 http://www.jimmunol.org/ References This article cites 60 articles, 36 of which you can access for free at: http://www.jimmunol.org/content/164/11/5850.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision by guest on September 25, 2021 • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2000 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. Soluble NSF Attachment Protein Receptors (SNAREs) in RBL-2H3 Mast Cells: Functional Role of Syntaxin 4 in Exocytosis and Identification of a Vesicle-Associated Membrane Protein 8-Containing Secretory Compartment1 Fabienne Paumet,* Joe¨lle Le Mao,* Sophie Martin,* Thierry Galli,† Bernard David,* Ulrich Blank,* and Miche`le Roa2* Mast cells upon stimulation through high affinity IgE receptors massively release inflammatory mediators by the fusion of spe- cialized secretory granules (related to lysosomes) with the plasma membrane. -

Transcriptomic Analysis of the Aquaporin (AQP) Gene Family
Pancreatology 19 (2019) 436e442 Contents lists available at ScienceDirect Pancreatology journal homepage: www.elsevier.com/locate/pan Transcriptomic analysis of the Aquaporin (AQP) gene family interactome identifies a molecular panel of four prognostic markers in patients with pancreatic ductal adenocarcinoma Dimitrios E. Magouliotis a, b, Vasiliki S. Tasiopoulou c, Konstantinos Dimas d, * Nikos Sakellaridis d, Konstantina A. Svokos e, Alexis A. Svokos f, Dimitris Zacharoulis b, a Division of Surgery and Interventional Science, Faculty of Medical Sciences, UCL, London, UK b Department of Surgery, University of Thessaly, Biopolis, Larissa, Greece c Faculty of Medicine, School of Health Sciences, University of Thessaly, Biopolis, Larissa, Greece d Department of Pharmacology, Faculty of Medicine, School of Health Sciences, University of Thessaly, Biopolis, Larissa, Greece e The Warren Alpert Medical School of Brown University, Providence, RI, USA f Riverside Regional Medical Center, Newport News, VA, USA article info abstract Article history: Background: This study aimed to assess the differential gene expression of aquaporin (AQP) gene family Received 14 October 2018 interactome in pancreatic ductal adenocarcinoma (PDAC) using data mining techniques to identify novel Received in revised form candidate genes intervening in the pathogenicity of PDAC. 29 January 2019 Method: Transcriptome data mining techniques were used in order to construct the interactome of the Accepted 9 February 2019 AQP gene family and to determine which genes members are differentially expressed in PDAC as Available online 11 February 2019 compared to controls. The same techniques were used in order to evaluate the potential prognostic role of the differentially expressed genes. Keywords: PDAC Results: Transcriptome microarray data of four GEO datasets were incorporated, including 142 primary Aquaporin tumor samples and 104 normal pancreatic tissue samples. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Male Reproductive System
MALE REPRODUCTIVE SYSTEM DR RAJARSHI ASH M.B.B.S.(CAL); D.O.(EYE) ; M.D.-PGT(2ND YEAR) DEPARTMENT OF PHYSIOLOGY CALCUTTA NATIONAL MEDICAL COLLEGE PARTS OF MALE REPRODUCTIVE SYSTEM A. Gonads – Two ovoid testes present in scrotal sac, out side the abdominal cavity B. Accessory sex organs - epididymis, vas deferens, seminal vesicles, ejaculatory ducts, prostate gland and bulbo-urethral glands C. External genitalia – penis and scrotum ANATOMY OF MALE INTERNAL GENITALIA AND ACCESSORY SEX ORGANS SEMINIFEROUS TUBULE Two principal cell types in seminiferous tubule Sertoli cell Germ cell INTERACTION BETWEEN SERTOLI CELLS AND SPERM BLOOD- TESTIS BARRIER • Blood – testis barrier protects germ cells in seminiferous tubules from harmful elements in blood. • The blood- testis barrier prevents entry of antigenic substances from the developing germ cells into circulation. • High local concentration of androgen, inositol, glutamic acid, aspartic acid can be maintained in the lumen of seminiferous tubule without difficulty. • Blood- testis barrier maintains higher osmolality of luminal content of seminiferous tubules. FUNCTIONS OF SERTOLI CELLS 1.Germ cell development 2.Phagocytosis 3.Nourishment and growth of spermatids 4.Formation of tubular fluid 5.Support spermiation 6.FSH and testosterone sensitivity 7.Endocrine functions of sertoli cells i)Inhibin ii)Activin iii)Follistatin iv)MIS v)Estrogen 8.Sertoli cell secretes ‘Androgen binding protein’(ABP) and H-Y antigen. 9.Sertoli cell contributes formation of blood testis barrier. LEYDIG CELL • Leydig cells are present near the capillaries in the interstitial space between seminiferous tubules. • They are rich in mitochondria & endoplasmic reticulum. • Leydig cells secrete testosterone,DHEA & Androstenedione. • The activity of leydig cell is different in different phases of life. -

Distinct Genetic Alterations in Colorectal Cancer
Distinct Genetic Alterations in Colorectal Cancer Hassan Ashktorab1*, Alejandro A. Scha¨ffer2, Mohammad Daremipouran3, Duane T. Smoot3, Edward Lee3, Hassan Brim3 1 Department of Medicine and Cancer Center, Howard University, College of Medicine, Washington, DC, United States of America, 2 National Center for Biotechnology Information, National Institutes of Health (NIH), Department of Health and Human Services (DHHS), Bethesda, Maryland, United States of America, 3 Department of Pathology, Howard University, College of Medicine, Washington, DC, United States of America Abstract Background: Colon cancer (CRC) development often includes chromosomal instability (CIN) leading to amplifications and deletions of large DNA segments. Epidemiological, clinical, and cytogenetic studies showed that there are considerable differences between CRC tumors from African Americans (AAs) and Caucasian patients. In this study, we determined genomic copy number aberrations in sporadic CRC tumors from AAs, in order to investigate possible explanations for the observed disparities. Methodology/Principal Findings: We applied genome-wide array comparative genome hybridization (aCGH) using a 105k chip to identify copy number aberrations in samples from 15 AAs. In addition, we did a population comparative analysis with aCGH data in Caucasians as well as with a widely publicized list of colon cancer genes (CAN genes). There was an average of 20 aberrations per patient with more amplifications than deletions. Analysis of DNA copy number of frequently altered chromosomes revealed that deletions occurred primarily in chromosomes 4, 8 and 18. Chromosomal duplications occurred in more than 50% of cases on chromosomes 7, 8, 13, 20 and X. The CIN profile showed some differences when compared to Caucasian alterations. Conclusions/Significance: Chromosome X amplification in male patients and chromosomes 4, 8 and 18 deletions were prominent aberrations in AAs. -

Proteomic Analysis of HIV-1 Nef Cellular Binding Partners Reveals a Role for Exocyst Complex Proteins in Mediating Enhancement of Intercellular Nanotube Formation
Proteomic analysis of HIV-1 Nef cellular binding partners reveals a role for exocyst complex proteins in mediating enhancement of intercellular nanotube formation The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Mukerji, Joya, Kevin C Olivieri, Vikas Misra, Kristin A Agopian, and Dana Gabuzda. 2012. Proteomic analysis of hiv-1 nef cellular binding partners reveals a role for exocyst complex proteins in mediating enhancement of intercellular nanotube formation. Retrovirology 9: 33. Published Version doi:10.1186/1742-4690-9-33 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:10445557 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA Mukerji et al. Retrovirology 2012, 9:33 http://www.retrovirology.com/content/9/1/33 RESEARCH Open Access Proteomic analysis of HIV-1 Nef cellular binding partners reveals a role for exocyst complex proteins in mediating enhancement of intercellular nanotube formation Joya Mukerji1,2, Kevin C Olivieri1, Vikas Misra1, Kristin A Agopian1,2 and Dana Gabuzda1,2,3* Abstract Background: HIV-1 Nef protein contributes to pathogenesis via multiple functions that include enhancement of viral replication and infectivity, alteration of intracellular trafficking, and modulation of cellular signaling pathways. Nef stimulates formation of tunneling nanotubes and virological synapses, and is transferred to bystander cells via these intercellular contacts and secreted microvesicles. Nef associates with and activates Pak2, a kinase that regulates T-cell signaling and actin cytoskeleton dynamics, but how Nef promotes nanotube formation is unknown. -

A Compendium of Co-Regulated Protein Complexes in Breast Cancer Reveals Collateral Loss Events
bioRxiv preprint doi: https://doi.org/10.1101/155333; this version posted June 26, 2017. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. A compendium of co-regulated protein complexes in breast cancer reveals collateral loss events Colm J. Ryan*1, Susan Kennedy1, Ilirjana Bajrami2, David Matallanas1, Christopher J. Lord2 1Systems Biology Ireland, School of Medicine, University College Dublin, Dublin 4, Ireland 2The Breast Cancer Now Toby Robins Breast Cancer Research Centre and CRUK Gene Function Laboratory, The Institute of Cancer Research, London, SW3 6JB, United Kingdom. * Correspondence: [email protected] Summary Protein complexes are responsible for the bulk of activities within the cell, but how their behavior and composition varies across tumors remains poorly understood. By combining proteomic profiles of breast tumors with a large-scale protein-protein interaction network, we have identified a set of 258 high-confidence protein complexes whose subunits have highly correlated protein abundance across tumor samples. We used this set to identify complexes that are reproducibly under- or over- expressed in specific breast cancer subtypes. We found that mutation or deletion of one subunit of a complex was often associated with a collateral reduction in protein expression of additional complex members. This collateral loss phenomenon was evident from proteomic, but not transcriptomic, profiles suggesting post- transcriptional control. Mutation of the tumor suppressor E-cadherin (CDH1) was associated with a collateral loss of members of the adherens junction complex, an effect we validated using an engineered model of E-cadherin loss. -

Microtubule Regulation in Cystic Fibrosis Pathophysiology
MICROTUBULE REGULATION IN CYSTIC FIBROSIS PATHOPHYSIOLOGY By: SHARON MARIE RYMUT Submitted in partial fulfillment of the requirements For the degree of Doctor of Philosophy Dissertation Advisor: Dr. Thomas J Kelley Department of Pharmacology CASE WESTERN RESERVE UNIVERSITY August 2015 CASE WESTERN RESERVE UNIVERSITY SCHOOL OF GRADUATE STUDIES We hereby approve the thesis/ dissertation of Sharon Marie Rymut candidate for the Doctor of Philosophy degree* Dissertation Advisor: Thomas J Kelley Committee Chair: Paul N MacDonald Committee Member: Ruth E Siegel Committee Member: Craig A Hodges Committee Member: Danny Manor Committee Member: Rebecca J Darrah Date of Defense: April 29, 2015 * We also certify that written approval has been obtained for any proprietary material contained therein. ii Dedication There are five chapters in this dissertation. To Mom, Dad, Joe, Marie and Susan, I dedicate one chapter to each of you. You can fight about which chapter you want later. iii Table of Contents Table of Contents................................................................................................................iv List of Tables .................................................................................................................... vii List of Figures .................................................................................................................. viii Acknowledgements ............................................................................................................. x List of Abbreviations ....................................................................................................... -
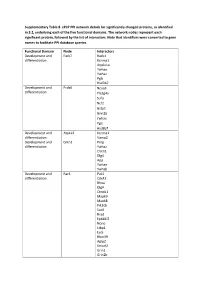
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4 -

Epigenome-Wide Association Study on Diffusing Capacity of the Lung: Replication and Meta- Analysis
EWAS of pulmonary diffusion 1 http://hdl.handle.net/1765/112075 Epigenome-wide association study on diffusing capacity Epigenome-wideof the lung: replication association and study on diffusingmeta-analysis capacity 3.4 of the lung: replication and meta- analysis 2 Erasmus Medical Center Rotterdam ABSTRACT Background: Epigenetics may play an important role in pathogenesis of lung diseases. However, little is known about the epigenetic factors that influence impaired gas exchange at the lungs. Aim: To identify the epigenetic signatures of the diffusing capacity of the lung measured by carbon monoxide uptake. Methods: Epigenome association study (EWAS) was performed on diffusing capacity, measured by carbon monoxide uptake (DLCO) and per alveolar volume (DLCO/VA) using the single-breath technique in 2,674 individuals from two population-based cohort studies, the Rotterdam Study (the discovery panel) and the Framingham Heart Study (the replication panel). We assessed the clinical relevance of our findings by investigating the identified sites in whole blood and lung tissue specific gene expression. Finally, we performed an exploratory epigenome-wide analysis, combining both cohorts. Results: We identified and replicated two CpG sites (cg05575921 and cg05951221) that were significantly associated with DLCO/VA and one (cg05575921) suggestively associated with DLCO. Furthermore, we found a positive association between AHRR (cg05575921) hypomethylation and gene expression of EXOC3 in whole blood. We confirmed that the expression of EXOC3 is positively associated with DLCO/VA and DLCO in lung tissue. Finally, several genome-wide associations were identified after meta-analysing the data of both cohorts. Conclusions: We report on epigenome-wide associations with diffusing capacity in the general population.