The Coalescent Process
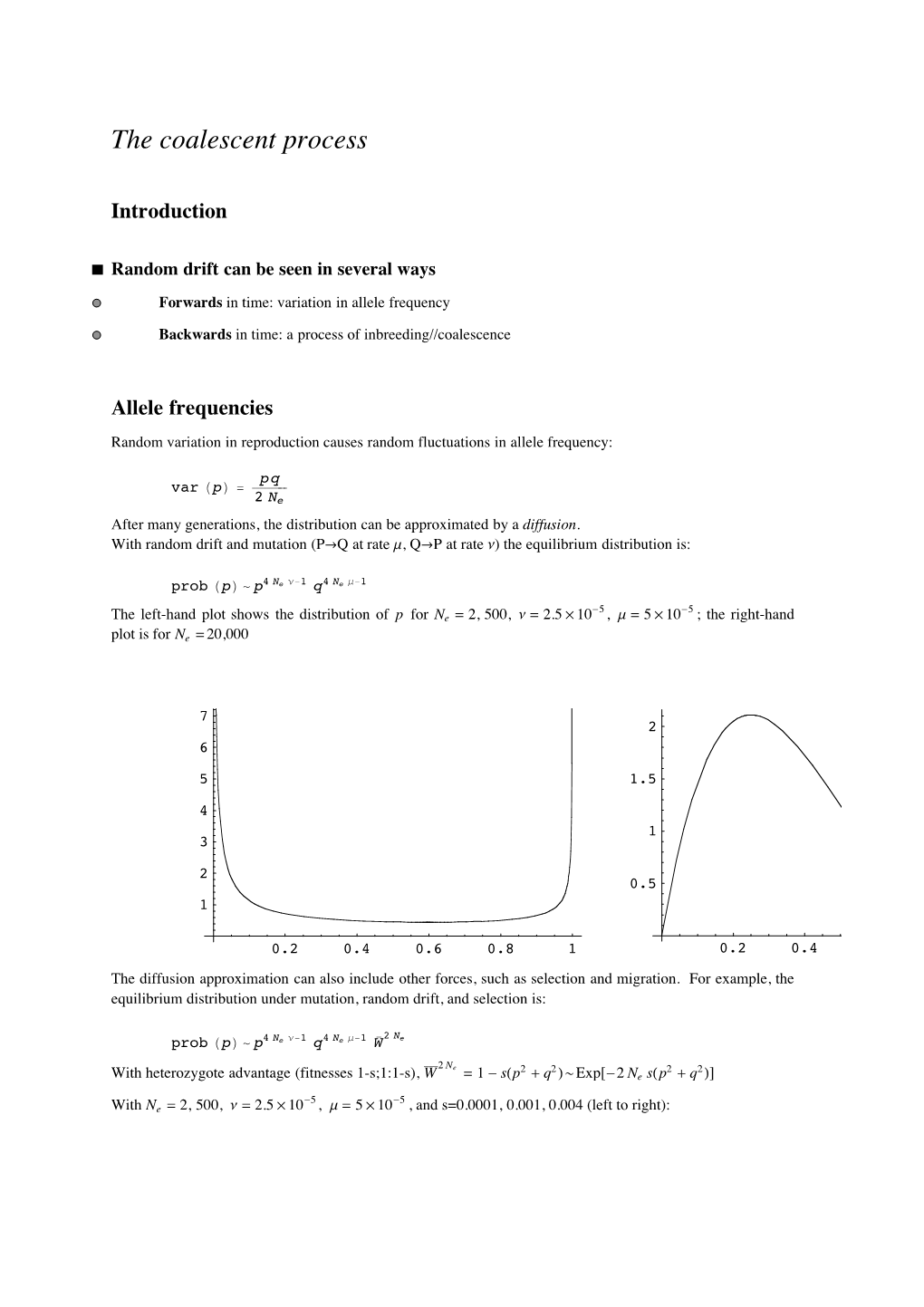
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Natural Selection and the Distribution of Identity-By-Descent in the Human Genome
Copyright Ó 2010 by the Genetics Society of America DOI: 10.1534/genetics.110.113977 Natural Selection and the Distribution of Identity-by-Descent in the Human Genome Anders Albrechtsen,*,1,2 Ida Moltke†,1 and Rasmus Nielsen‡ *Department of Biostatistics, University of Copenhagen, Copenhagen, 1014, Denmark, †Center for Bioinformatics, Copenhagen, 2200, Denmark and ‡Department of Integrative Biology and Statistics, University of California, Berkeley, California 94720 Manuscript received April 30, 2010 Accepted for publication June 14, 2010 ABSTRACT There has recently been considerable interest in detecting natural selection in the human genome. Selection will usually tend to increase identity-by-descent (IBD) among individuals in a population, and many methods for detecting recent and ongoing positive selection indirectly take advantage of this. In this article we show that excess IBD sharing is a general property of natural selection and we show that this fact makes it possible to detect several types of selection including a type that is otherwise difficult to detect: selection acting on standing genetic variation. Motivated by this, we use a recently developed method for identifying IBD sharing among individuals from genome-wide data to scan populations from the new HapMap phase 3 project for regions with excess IBD sharing in order to identify regions in the human genome that have been under strong, very recent selection. The HLA region is by far the region showing the most extreme signal, suggesting that much of the strong recent selection acting on the human genome has been immune related and acting on HLA loci. As equilibrium overdominance does not tend to increase IBD, we argue that this type of selection cannot explain our observations. -

Coalescent Likelihood Methods
Coalescent Likelihood Methods Mary K. Kuhner Genome Sciences University of Washington Seattle WA Outline 1. Introduction to coalescent theory 2. Practical example 3. Genealogy samplers 4. Break 5. Survey of samplers 6. Evolutionary forces 7. Practical considerations Population genetics can help us to find answers We are interested in questions like { How big is this population? { Are these populations isolated? How common is migration? { How fast have they been growing or shrinking? { What is the recombination rate across this region? { Is this locus under selection? All of these questions require comparison of many individuals. Coalescent-based studies How many gray whales were there prior to whaling? When was the common ancestor of HIV lines in a Libyan hospital? Is the highland/lowland distinction in Andean ducks recent or ancient? Did humans wipe out the Beringian bison population? What proportion of HIV virions in a patient actually contribute to the breeding pool? What is the direction of gene flow between European rabbit populations? Basics: Wright-Fisher population model All individuals release many gametes and new individuals for the next generation are formed randomly from these. Wright-Fisher population model Population size N is constant through time. Each individual gets replaced every generation. Next generation is drawn randomly from a large gamete pool. Only genetic drift affects the allele frequencies. Other population models Other population models can often be equated to Wright-Fisher The N parameter becomes the effective -

Natural Selection and Coalescent Theory
Natural Selection and Coalescent Theory John Wakeley Harvard University INTRODUCTION The story of population genetics begins with the publication of Darwin’s Origin of Species and the tension which followed concerning the nature of inheritance. Today, workers in this field aim to understand the forces that produce and maintain genetic variation within and between species. For this we use the most direct kind of genetic data: DNA sequences, even entire genomes. Our “Great Obsession” with explaining genetic variation (Gillespie, 2004a) can be traced back to Darwin’s recognition that natural selection can occur only if individuals of a species vary, and this variation is heritable. Darwin might have been surprised that the importance of natural selection in shaping variation at the molecular level would be de-emphasized, beginning in the late 1960s, by scientists who readily accepted the fact and importance of his theory (Kimura, 1983). The motivation behind this chapter is the possible demise of this Neutral Theory of Molecular Evolution, which a growing number of population geneticists feel must follow recent observations of genetic variation within and between species. One hundred fifty years after the publication of the Origin, we are struggling to fully incorporate natural selection into the modern, genealogical models of population genetics. The main goal of this chapter is to present the mathematical models that have been used to describe the effects of positive selective sweeps on genetic variation, as mediated by gene genealogies, or coalescent trees. Background material, comprised of population genetic theory and simulation results, is provided in order to facilitate an understanding of these models. -

Population Genetics: Wright Fisher Model and Coalescent Process
POPULATION GENETICS: WRIGHT FISHER MODEL AND COALESCENT PROCESS by Hailong Cui and Wangshu Zhang Superviser: Prof. Quentin Berger A Final Project Report Presented In Partial Fulfillment of the Requirements for Math 505b April 2014 Acknowledgments We want to thank Prof. Quentin Berger for introducing to us the Wright Fisher model in the lecture, which inspired us to choose Population Genetics for our project topic. The resources Prof. Berger provided us have been excellent learning mate- rials and his feedback has helped us greatly to create this report. We also like to acknowledge that the research papers (in the reference) are integral parts of this process. They have motivated us to learn more about models beyond the class, and granted us confidence that these probabilistic models can actually be used for real applications. ii Contents Acknowledgments ii Abstract iv 1 Introduction to Population Genetics 1 2 Wright Fisher Model 2 2.1 Random drift . 2 2.2 Genealogy of the Wright Fisher model . 4 3 Coalescent Process 8 3.1 Kingman’s Approximation . 8 4 Applications 11 Reference List 12 iii Abstract In this project on Population Genetics, we aim to introduce models that lay the foundation to study more complicated models further. Specifically we will discuss Wright Fisher model as well as Coalescent process. The reason these are of our inter- est is not just the mathematical elegance of these models. With the availability of massive amount of sequencing data, we actually can use these models (or advanced models incorporating variable population size, mutation effect etc, which are how- ever out of the scope of this project) to solve and answer real questions in molecular biology. -

Identity by Descent in Pedigrees and Populations; Methods for Genome-Wide Linkage and Association
Identity by descent in pedigrees and populations Overview - 1 Identity by descent in pedigrees and populations; methods for genome-wide linkage and association. UNE Short Course: Feb 14-18, 2011 Dr Elizabeth A Thompson UNE-Short Course Feb 2011 Identity by descent in pedigrees and populations Overview - 2 Timetable Monday 9:00-10:30am 1: Introduction and Overview. 11:00am-12:30pm 2: Identity by Descent; relationships and relatedness 1:30-3:00pm 3: Genetic variation and allelic association. 3:30-5:00pm 4: Allelic association and population structure. Tuesday 9:00-10:30am 5: Genetic associations for a quantitative trait 11:00am-12:30pm 6: Hidden Markov models; HMM 1:30-3:00pm 7: Haplotype blocks and the coalescent. 3:30-5:00pm 8: LD mapping via coalescent ancestry. Wednesday a.m. 9:00-10:30am 9: The EM algorithm 11:00am-12:30pm 10: MCMC and Bayesian sampling Dr Elizabeth A Thompson UNE-Short Course Feb 2011 Identity by descent in pedigrees and populations Overview - 3 Wednesday p.m. 1:30-3:00pm 11: Association mapping in structured populations 3:30-5:00pm 12: Association mapping in admixed populations Thursday 9:00-10:30am 13: Inferring ibd segments; two chromosomes. 11:00am-12:30pm 14: BEAGLE: Haplotype and ibd imputation. 1:30-3:00pm 15: ibd between two individuals. 3:30-5:00pm 16: ibd among multiple chromosomes. Friday 9:00-10:30am 17: Pedigrees in populations. 11:00am-12:30pm 18: Lod scores within and between pedigrees. 1:30-3:00pm 19: Wrap-up and questions. Bibliography Software notes and links. -

Prediction of Identity by Descent Probabilities from Marker-Haplotypes Theo Meuwissen, Mike Goddard
Prediction of identity by descent probabilities from marker-haplotypes Theo Meuwissen, Mike Goddard To cite this version: Theo Meuwissen, Mike Goddard. Prediction of identity by descent probabilities from marker-haplotypes. Genetics Selection Evolution, BioMed Central, 2001, 33 (6), pp.605-634. 10.1051/gse:2001134. hal-00894392 HAL Id: hal-00894392 https://hal.archives-ouvertes.fr/hal-00894392 Submitted on 1 Jan 2001 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Genet. Sel. Evol. 33 (2001) 605–634 605 © INRA, EDP Sciences, 2001 Original article Prediction of identity by descent probabilities from marker-haplotypes a; b Theo H.E. MEUWISSEN ∗, Mike E. GODDARD a Research Institute of Animal Science and Health, Box 65, 8200 AB Lelystad, The Netherlands b Institute of Land and Food Resources, University of Melbourne, Parkville Victorian Institute of Animal Science, Attwood, Victoria, Australia (Received 13 February 2001; accepted 11 June 2001) Abstract – The prediction of identity by descent (IBD) probabilities is essential for all methods that map quantitative trait loci (QTL). The IBD probabilities may be predicted from marker genotypes and/or pedigree information. Here, a method is presented that predicts IBD prob- abilities at a given chromosomal location given data on a haplotype of markers spanning that position. -

Frontiers in Coalescent Theory: Pedigrees, Identity-By-Descent, and Sequentially Markov Coalescent Models
Frontiers in Coalescent Theory: Pedigrees, Identity-by-Descent, and Sequentially Markov Coalescent Models The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Wilton, Peter R. 2016. Frontiers in Coalescent Theory: Pedigrees, Identity-by-Descent, and Sequentially Markov Coalescent Models. Doctoral dissertation, Harvard University, Graduate School of Arts & Sciences. Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:33493608 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA Frontiers in Coalescent Theory: Pedigrees, Identity-by-descent, and Sequentially Markov Coalescent Models a dissertation presented by Peter Richard Wilton to The Department of Organismic and Evolutionary Biology in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the subject of Biology Harvard University Cambridge, Massachusetts May 2016 ©2016 – Peter Richard Wilton all rights reserved. Thesis advisor: Professor John Wakeley Peter Richard Wilton Frontiers in Coalescent Theory: Pedigrees, Identity-by-descent, and Sequentially Markov Coalescent Models Abstract The coalescent is a stochastic process that describes the genetic ancestry of individuals sampled from a population. It is one of the main tools of theoretical population genetics and has been used as the basis of many sophisticated methods of inferring the demo- graphic history of a population from a genetic sample. This dissertation is presented in four chapters, each developing coalescent theory to some degree. -

Identity-By-Descent Detection Across 487,409 British Samples Reveals Fine Scale Population Structure and Ultra-Rare Variant Asso
ARTICLE https://doi.org/10.1038/s41467-020-19588-x OPEN Identity-by-descent detection across 487,409 British samples reveals fine scale population structure and ultra-rare variant associations ✉ Juba Nait Saada 1 , Georgios Kalantzis 1, Derek Shyr 2, Fergus Cooper 3, Martin Robinson 3, ✉ Alexander Gusev 4,5,7 & Pier Francesco Palamara 1,6,7 1234567890():,; Detection of Identical-By-Descent (IBD) segments provides a fundamental measure of genetic relatedness and plays a key role in a wide range of analyses. We develop FastSMC, an IBD detection algorithm that combines a fast heuristic search with accurate coalescent-based likelihood calculations. FastSMC enables biobank-scale detection and dating of IBD segments within several thousands of years in the past. We apply FastSMC to 487,409 UK Biobank samples and detect ~214 billion IBD segments transmitted by shared ancestors within the past 1500 years, obtaining a fine-grained picture of genetic relatedness in the UK. Sharing of common ancestors strongly correlates with geographic distance, enabling the use of genomic data to localize a sample’s birth coordinates with a median error of 45 km. We seek evidence of recent positive selection by identifying loci with unusually strong shared ancestry and detect 12 genome-wide significant signals. We devise an IBD-based test for association between phenotype and ultra-rare loss-of-function variation, identifying 29 association sig- nals in 7 blood-related traits. 1 Department of Statistics, University of Oxford, Oxford, UK. 2 Department of Biostatistics, Harvard T.H. Chan School of Public Health, Boston, MA 02115, USA. 3 Department of Computer Science, University of Oxford, Oxford, UK. -

Variation in Meiosis, Across Genomes, and in Populations
REVIEW Identity by Descent: Variation in Meiosis, Across Genomes, and in Populations Elizabeth A. Thompson1 Department of Statistics, University of Washington, Seattle, Washington 98195-4322 ABSTRACT Gene identity by descent (IBD) is a fundamental concept that underlies genetically mediated similarities among relatives. Gene IBD is traced through ancestral meioses and is defined relative to founders of a pedigree, or to some time point or mutational origin in the coalescent of a set of extant genes in a population. The random process underlying changes in the patterns of IBD across the genome is recombination, so the natural context for defining IBD is the ancestral recombination graph (ARG), which specifies the complete ancestry of a collection of chromosomes. The ARG determines both the sequence of coalescent ancestries across the chromosome and the extant segments of DNA descending unbroken by recombination from their most recent common ancestor (MRCA). DNA segments IBD from a recent common ancestor have high probability of being of the same allelic type. Non-IBD DNA is modeled as of independent allelic type, but the population frame of reference for defining allelic independence can vary. Whether of IBD, allelic similarity, or phenotypic covariance, comparisons may be made to other genomic regions of the same gametes, or to the same genomic regions in other sets of gametes or diploid individuals. In this review, I present IBD as the framework connecting evolutionary and coalescent theory with the analysis of genetic data observed on individuals. I focus on the high variance of the processes that determine IBD, its changes across the genome, and its impact on observable data. -

Rapid Estimation of SNP Heritability Using Predictive Process Approximation in Large Scale Cohort Studies
bioRxiv preprint doi: https://doi.org/10.1101/2021.05.12.443931; this version posted May 14, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Rapid Estimation of SNP Heritability using Predictive Process approximation in Large scale Cohort Studies Souvik Seal1, Abhirup Datta2, and Saonli Basu3 1Department of Biostatistics and Informatics, University of Colorado Anschutz Medical Campus 2Department of Biostatistics, Johns Hopkins Bloomberg School of Public Health 3Division of Biostatistics, University of Minnesota Twin Cities Abstract With the advent of high throughput genetic data, there have been attempts to estimate heritability from genome-wide SNP data on a cohort of distantly related individuals using linear mixed model (LMM). Fitting such an LMM in a large scale cohort study, however, is tremendously challenging due to its high dimensional linear algebraic operations. In this paper, we propose a new method named PredLMM approximating the aforementioned LMM moti- vated by the concepts of genetic coalescence and gaussian predictive process. PredLMM has substantially better computational complexity than most of the existing LMM based methods and thus, provides a fast alternative for estimating heritability in large scale cohort studies. Theoretically, we show that under a model of genetic coalescence, the limiting form of our approximation is the celebrated predictive process approximation of large gaussian process likelihoods that has well-established accuracy standards. We illustrate our approach with ex- tensive simulation studies and use it to estimate the heritability of multiple quantitative traits from the UK Biobank cohort. -

The Coalescent Model
The Coalescent Model Florian Weber Karlsruhe Institute of Technology (KIT) Abstract. The coalescent model is the result of thorough analysis of the Wright-Fisher model. It analyzes the frequency and distribution of coalescence events in the evolutionary history. These events approximate an exponential distribution with the highest rate in the recent past. The number of surviving lineages in a generation also increases the frequency quadratically. Since the number of surviving lineages always decreases as one goes further into the past, this actually strengthens the effect of the exponential distribution. This model is however incomplete in that it assumes a constant population- size which is often not the case. Further analysis demonstrates that the likelihood of a coalescence event is inversely proportional to the population- size. In order to incorporate this observation into the model, it is possible to scale the time used by the model in a non-linear way to the actual time. With that extension the coalescent model still has limitations, but it is possible to remove most of those with similar changes. This allows to incorporate things like selection, separated groups and heterozygous organisms. 1 Motivation When we want to consider how allele-frequencies change in an environment with non-overlapping generations, random mating and no selection, there are a couple of models to choose from: The Hardy-Weinberg model and the more advanced Wright-Fisher model are probably the most well known ones. Both have serious disadvantages though: While the applicability of Hardy- Weinberg for real-world-populations is questionable at best, the Wright-Fisher model is computationally expensive and cannot perform calculations from the present into the past. -

Identity-By-Descent Analyses for Measuring Population Dynamics and Selection in Recombining Pathogens
bioRxiv preprint doi: https://doi.org/10.1101/088039; this version posted March 14, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Identity-by-descent analyses for measuring population dynamics 2 and selection in recombining pathogens 3 4 Lyndal Henden1,2, Stuart Lee3,4, Ivo Mueller1,2, Alyssa Barry1,2, Melanie Bahlo1,2,* 5 6 1. Population Health and Immunity Division, The Walter and Eliza Hall Institute of Medical 7 Research, Parkville VIC, Australia 8 2. Department of Medical Biology, University of Melbourne, Parkville VIC, Australia 9 3. Department of Econometrics and Business Statistics, Monash University, Clayton VIC, 10 Australia 11 4. Molecular Medicine Division, The Walter and Eliza Hall Institute of Medical Research, 12 Parkville VIC, Australia 13 14 * Corresponding author 15 E-mail: [email protected] (MB) 16 17 18 19 20 21 22 23 24 25 1 bioRxiv preprint doi: https://doi.org/10.1101/088039; this version posted March 14, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 26 Abstract 27 28 Identification of genomic regions that are identical by descent (IBD) has proven useful for human 29 genetic studies where analyses have led to the discovery of familial relatedness and fine-mapping 30 of disease critical regions.