Grid-Cell Modules Remain Coordinated When Neural Activity Is Dissociated from External Sensory Cues
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Phase Relations of Purkinje Cells in the Rabbit Flocculus During Compensatory Eye Movements
JOURNALOFNEUROPHYSIOLOGY Vol. 74, No. 5, November 1995. Printed in U.S.A. Phase Relations of Purkinje Cells in the Rabbit Flocculus During Compensatory Eye Movements C. I. DE ZEEUW, D. R. WYLIE, J. S. STAHL, AND J. I. SIMPSON Department of Physiology and Neuroscience, New York University Medical Center, New York 1OOM; and Department of Anatomy, Erasmus University Rotterdam, 3000 DR Rotterdam, Postbus 1738, The Netherlands SUMMARY AND CONCLUSIONS 17 cases (14%) showed CS modulation. In the majority (15 of 1. Purkinje cells in the rabbit flocculus that respond best to 17) of these cases, the CS activity increased with contralateral rotation about the vertical axis (VA) project to flocculus-receiving head rotation; these modulations occurred predominantly at the neurons (FRNs) in the medial vestibular nucleus. During sinusoi- higher stimulus velocities. dal rotation, the phase of FRNs leads that of medial vestibular 7. On the basis of the finding that FRNs of the medial vestibular nucleus neurons not receiving floccular inhibition (non-FRNs) . If nucleus lead non-FRNs, we predicted that floccular VA Purkinje the FRN phase lead is produced by signals from the ~~OCCU~US,then cells would in turn lead FRNs. This prediction is confirmed in the the Purkinje cells should functionally lead the FRNs. In the present present study. The data are therefore consistent with the hypothesis study we recorded from VA Purkinje cells in the flocculi of awake, that the phase-leading characteristics of FRN modulation could pigmented rabbits during compensatory eye movements to deter- come about by summation of VA Purkinje cell activity with that mine whether Purkinje cells have the appropriate firing rate phases of cells whose phase would otherwise be identical to that of non- to explain the phase-leading characteristics of the FRNs. -

Dark Data and Its Future Prospects
International Journal of Engineering Technology Science and Research IJETSR www.ijetsr.com ISSN 2394 – 3386 Volume 5, Issue 1 January 2018 Dark Data and Its Future Prospects Sarang Saxena St. Joseph’s Degree & Pg College ABSTRACT Dark data is data which is acquired through various computer network operations but not used in any manner to derive insights or for decision making. The ability of an organisation to collect data can exceed the throughput at which it can analyse the data. In some cases the organisation may not even be aware that the data is being collected. It’s data that is ever present, unknown and unmanaged. For example, a company may collect data on how users use its products, internal statistics about software development processes, and website visits. However, a large portion of the collected data is never even analysed. In recent times, there are certain businesses which have realised the importance of such data and are working towards finding techniques, methods, processes and developing software so that such data is properly utilized Through this paper I’m trying to find the scope and importance of dark data, its importance in the future, its implications on the smaller firms and organisations in need of relevant and accurate data which is difficult to find but is hoarded by giant firms who do not intend to reveal such information as a part of their practice or have no idea that the data even exists, the amount of data generatedby organisations and the percentage of which is actually utilised, the businesses formed around mining dark data and analysing it, the effects of dark data, the dangers and limitations of dark data. -

Dark Data As the New Challenge for Big Data Science and the Introduction of the Scientific Data Officer
Delft University of Technology Dark Data as the New Challenge for Big Data Science and the Introduction of the Scientific Data Officer Schembera, Björn; Duran, Juan Manuel DOI 10.1007/s13347-019-00346-x Publication date 2019 Document Version Final published version Published in Philosophy & Technology Citation (APA) Schembera, B., & Duran, J. M. (2019). Dark Data as the New Challenge for Big Data Science and the Introduction of the Scientific Data Officer. Philosophy & Technology, 33(1), 93-115. https://doi.org/10.1007/s13347-019-00346-x Important note To cite this publication, please use the final published version (if applicable). Please check the document version above. Copyright Other than for strictly personal use, it is not permitted to download, forward or distribute the text or part of it, without the consent of the author(s) and/or copyright holder(s), unless the work is under an open content license such as Creative Commons. Takedown policy Please contact us and provide details if you believe this document breaches copyrights. We will remove access to the work immediately and investigate your claim. This work is downloaded from Delft University of Technology. For technical reasons the number of authors shown on this cover page is limited to a maximum of 10. Philosophy & Technology https://doi.org/10.1007/s13347-019-00346-x RESEARCH ARTICLE Dark Data as the New Challenge for Big Data Science and the Introduction of the Scientific Data Officer Bjorn¨ Schembera1 · Juan M. Duran´ 2 Received: 25 June 2018 / Accepted: 1 March 2019 / © The Author(s) 2019 Abstract Many studies in big data focus on the uses of data available to researchers, leaving without treatment data that is on the servers but of which researchers are unaware. -

An Intelligent Navigator for Large-Scale Dark Structured Data
CognitiveDB: An Intelligent Navigator for Large-scale Dark Structured Data Michael Gubanov, Manju Priya, Maksim Podkorytov Department of Computer Science University of Texas at San Antonio {mikhail.gubanov, maksim.podkorytov}@utsa.edu, [email protected] ABSTRACT Consider a data scientist who is trying to predict hard Volume and Variety of Big data [Stonebraker, 2012] are sig- cider sales in the regions of interest to be able to restock nificant impediments for anyone who wants to quickly un- accordingly for the next season. Since consumption of hard derstand what information is inside a large-scale dataset. cider is correlated with weather, access to the complete weather Such data is often informally referred to as 'dark' due to data per region would improve prediction accuracy. How- the difficulties of understanding its contents. For example, ever, such a dataset is not available internally, hence, the there are millions of structured tables available on the Web, data scientist is considering enriching existing data with de- but finding a specific table or summarizing all Web tables tailed weather data from an external data source (e.g. the to understand what information is available is infeasible or Web). However, the cost to do ETL (Extract-Transform- very difficult because of the scale involved. Load) from a large-scale external data source like the Web Here we present and demonstrate CognitiveDB, an intel- in order to integrate weather data is expected to be high, ligent cognitive data management system that can quickly therefore the data scientist never acquires the missing data summarize the contents of an unexplored large-scale struc- pieces and does not take weather into account. -

An Epidemiology of Big Data
Syracuse University SURFACE Dissertations - ALL SURFACE May 2014 An Epidemiology of Big Data John Mark Young Syracuse University Follow this and additional works at: https://surface.syr.edu/etd Part of the Social and Behavioral Sciences Commons Recommended Citation Young, John Mark, "An Epidemiology of Big Data" (2014). Dissertations - ALL. 105. https://surface.syr.edu/etd/105 This Dissertation is brought to you for free and open access by the SURFACE at SURFACE. It has been accepted for inclusion in Dissertations - ALL by an authorized administrator of SURFACE. For more information, please contact [email protected]. ABSTRACT Federal legislation designed to transform the U.S. healthcare system and the emergence of mobile technology are among the common drivers that have contributed to a data explosion, with industry analysts and stakeholders proclaiming this decade the big data decade in healthcare (Horowitz, 2012). But a precise definition of big data is hazy (Dumbill, 2013). Instead, the healthcare industry mainly relies on metaphors, buzzwords, and slogans that fail to provide information about big data’s content, value, or purposes for existence (Burns, 2011). Bollier and Firestone (2010) even suggests “big data does not really exist in healthcare” (p. 29). While federal policymakers and other healthcare stakeholders struggle with the adoption of Meaningful Use Standards, International Classification of Diseases-10 (ICD-10), and electronic health record interoperability standards, big data in healthcare remains a widely misunderstood -

Neuronal Couplings Between Retinal Ganglion Cells Inferred by Efficient Inverse Statistical Physics Methods
Supporting Information Appendix for Neuronal couplings between retinal ganglion cells inferred by efficient inverse statistical physics methods S. Cocco, S. Leibler, R. Monasson Table of contents: 1. Inference of couplings and of their accuracy within the inverse Ising model (page 2) 2. Limitations of the Ising model: higher-order correlations and couplings (page 7) 3. Algorithm for the Inverse Integrate-and-Fire Problem (page 11) 4. On Cross-correlograms: Analysis of Data, Relationship with Couplings, and Simulations (page 14) 5. Correspondence between Integrate-and-Fire and Ising Inverse Models (page 23) 6. Spatial features of the inferred couplings (page 28) 7. On states and the large-N limit in the inverse Ising model (page 36) 8. Bibliography and footnotes (page 39) Supporting Information Appendix, Section 1: Inference of couplings and of their accuracy within the inverse Ising model I. INFERENCE AND MINIMIZATION OF THE ISING ENTROPY A multi-electrode recording provides the firing times of N recorded cells during a time interval of duration T . In the Ising inverse approach the recording interval is divided into time windows (time-bins) of width ∆t and the data are encoded in T/∆t configurations s = (s1,s2,...,sN ) of the N binary variables si, (i = 1,...,N) called spins (by τ analogy with magnetic systems described by the Ising model). The value of each spin variable is: si = 1, if the cell i is active in the time-bin τ (τ = 1,...,B = T/∆t), si = 0 otherwise. Let pi be the probability that the cell i is active in a given time-bin, and pij be the joint probability that the cells i and j are both active in the same bin. -
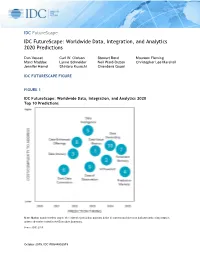
IDC Futurescape: Worldwide Data, Integration, and Analytics 2020 Predictions
IDC FutureScape IDC FutureScape: Worldwide Data, Integration, and Analytics 2020 Predictions Dan Vesset Carl W. Olofson Stewart Bond Maureen Fleming Marci Maddox Lynne Schneider Neil Ward-Dutton Christopher Lee Marshall Jennifer Hamel Shintaro Kusachi Chandana Gopal IDC FUTURESCAPE FIGURE FIGURE 1 IDC FutureScape: Worldwide Data, Integration, and Analytics 2020 Top 10 Predictions Note: Marker number refers only to the order the prediction appears in the document and does not indicate rank or importance, unless otherwise noted in the Executive Summary. Source: IDC, 2019 October 2019, IDC #US44802519 EXECUTIVE SUMMARY When almost 100 CEOs were asked by IDC in a study in August 2019 about the importance of various strategic areas to their organization over the next five years, fully 80% of respondents (second only to focus on digital trust) mentioned data (or more specifically, using data in advanced decision models to affect performance and competitive advantage). And these executives are not simply paying homage to the current trendy topics. According to IDC's Worldwide Big Data and Analytics Tracker and Spending Guide, enterprises worldwide spent $169 billion on BDA software, hardware, and services in 2018. However, most enterprises don't know if they are getting value or how much value they are getting out of data and analytics. Most are not tapping into dark data that is created by sits unused nor are most enterprises even attempting to get a return on their data asset externally. Most have subpar data literacy and incomplete data intelligence. Many are planning under the assumptions of huge potential benefits of artificial intelligence (AI), but lack foundational data, integration, and analytics capabilities that are prerequisites for moving along the AI-based automation evolution path. -

Population-Based Linkage of Big Data in Dental Research
International Journal of Environmental Research and Public Health Review Population-Based Linkage of Big Data in Dental Research Tim Joda 1,*, Tuomas Waltimo 2, Christiane Pauli-Magnus 3, Nicole Probst-Hensch 4 and Nicola U. Zitzmann 1 1 Department of Reconstructive Dentistry, University Center for Dental Medicine Basel, 4056 Basel, Switzerland; [email protected] 2 Department of Oral Health & Medicine Dentistry, University Center for Dental Medicine Basel, 4056 Basel, Switzerland; [email protected] 3 Department of Clinical Research & Clinical Trial Unit, Faculty of Medicine, University of Basel, 4031 Basel, Switzerland; [email protected] 4 Department of Epidemiology & Public Health, Swiss Tropical & Public Health Institute Basel, University of Basel, 4051 Basel, Switzerland; [email protected] * Correspondence: [email protected]; Tel.: +4161-267-2631 Received: 19 October 2018; Accepted: 23 October 2018; Published: 25 October 2018 Abstract: Population-based linkage of patient-level information opens new strategies for dental research to identify unknown correlations of diseases, prognostic factors, novel treatment concepts and evaluate healthcare systems. As clinical trials have become more complex and inefficient, register-based controlled (clinical) trials (RC(C)T) are a promising approach in dental research. RC(C)Ts provide comprehensive information on hard-to-reach populations, allow observations with minimal loss to follow-up, but require large sample sizes with generating high level of external validity. Collecting data is only valuable if this is done systematically according to harmonized and inter-linkable standards involving a universally accepted general patient consent. Secure data anonymization is crucial, but potential re-identification of individuals poses several challenges. -

Industrial Statistics
Industrial Statistics Ronald Christensen and Aparna V. Huzurbazar Department of Mathematics and Statistics University of New Mexico Copyright c 2002, 2021 v To Walt and Ed Contents Preface xiii 0.1 Standards xiii 0.2 Six-Sigma xiii 0.3 Lean xiv 0.4 This Book xiv 0.4.1 Computing xv 1 Introduction1 1.1 Four Principles for Quality1 1.1.1 Institute and Maintain Leadership for Quality Improvement2 1.1.2 Create Cooperation3 1.1.3 Train, Retrain, and Educate5 1.1.4 Insist on Action5 1.2 Some Technical Matters5 1.3 The Shewhart Cycle: PDSA6 1.4 Benchmarking7 1.5 Exercises7 2 Basic Tools 9 2.1 Data Collection9 2.2 Pareto and Other Charts 11 2.3 Histograms 13 2.3.1 Stem and Leaf Displays 16 2.3.2 Dot Plots 18 2.4 Box Plots 19 2.5 Cause and Effect Diagrams 19 2.6 Flow Charts 20 3 Probability 23 3.1 Introduction 23 3.2 Something about counting 25 3.3 Working with Random Variables 26 3.3.1 Probability Distributions, Expected Values 27 3.3.2 Independence, Covariance, Correlation 28 3.3.3 Expected values and variances for sample means 30 3.4 Binomial Distribution 32 3.5 Poisson distribution 34 3.6 Normal Distribution 34 vii viii CONTENTS 4 Control Charts 37 4.1 Individuals Chart 38 4.2 Means Charts and Dispersion Charts 40 4.2.1 Process Capability 46 4.2.1.1 Six-Sigma 47 4.3 Attribute Charts 48 4.4 Control Chart Summary 53 4.5 Average Run Lengths 54 4.6 Discussion 55 4.7 Testing Mean Shifts from a Target 56 4.7.1 Exponentially Weighted Moving Average Charts 56 4.7.1.1 Derivations of results 58 4.7.2 CUSUM charts 60 4.8 Computing 61 4.8.1 Minitab 61 -

Anomalies and Error Sources
NICMOS 4 Anomalies and Error Sources In This Chapter... NICMOS Dark Current and Bias / 4-4 Bars / 4-19 Detector Nonlinearity Issues / 4-20 Flatfielding / 4-22 Pixel Defects and Bad Imaging Regions / 4-24 Effects of Overexposure / 4-28 Cosmic Rays of Unusual Size / 4-34 Scattered Earthlight / 4-36 The previous chapter described the basic stages of NICMOS pipeline processing. As with any instrument, however, high quality data reduction does not end with the standard pipeline processing. NICMOS data are subject to a variety of anomalies, artifacts, and instabilities which complicate the task of data reduction and analysis. Most of these can be handled with careful post-facto recalibration and processing, which usually yield excellent, scientific grade data reductions. Careful NICMOS data processing usually requires a certain amount of "hands-on" interaction from the user, who must inspect for data anomalies and treat them accordingly during the reduction procedures. This chapter describes the most common problems affecting NICMOS data at the level of frame-by-frame processing. In some cases, recognizing and treating problems with NICMOS data requires a moderately in-depth understanding of the details of instrumental behavior; problems with dark and bias subtraction are a good example. Where appropriate, this chapter offers a fairly detailed discussion of the relevant workings of the instrument, but the reader should consult the NICMOS Instrument Handbook for further details. Each section of this chapter deals with a different aspect of NICMOS data processing, roughly following the order of the processing steps in the standard STSDAS pipeline. Various potential problems are described and illustrated. -

Visual Analytics on Biomedical Dark Data
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 28 October 2020 Visual Analytics on Biomedical Dark Data Shashwat Aggarwal a, Ramesh Singh b a University of Delhi a National Informatics Center 1. INTRODUCTION In today’s data centralized world, the practice of data visualization has become an indispensable tool in numerous domains such as Research, Marketing, Journalism, Biology etc. Data visualization is the art of efficiently organizing and presenting data in a graphically appealing format. It speeds up the process of decision making and pattern recognition, thereby enabling decision makers to make informed decisions. With the rise in technology, the data has been exploding exponentially, and the world’s scientific knowledge is accessible with ease. There is an enormous amount of data available in the form of scientific articles, government reports, natural language, and images that in total contributes to around 80% of overall data generated as shown in an excerpt from The Digital Universe [1] in Figure 1. However, most of the data lack structure and cannot be easily categorized and imported into regular databases. This type of data is often termed as Dark Data. Data visualization techniques proffer a potential solution to overcome the problem of handling and analyzing overwhelming amounts of such information. It enables the decision maker to look at data differently and more imaginatively. It promotes creative data exploration by allowing quick comprehension of information, the discovery of emerging trends, identification of relationships and patterns etc. Figure 1. Worldwide growth of corporate data categorized by structured and unstructured/dark data over the past decade. 1 © 2020 by the author(s). -

Data Science by Analyticbridge
Data Science by AnalyticBridge Vincent Granville, Ph.D. Founder, Data Wizard, Managing Partner www.AnalyticBridge.com - www.DataScienceCentral.com Download the most recent version at http://bit.ly/oB0zxn This version was released on 01/03/2013 (123 pages – Gartner’s contribution updated) Previous release was on 06/05/2012 (123 pages) Published by AnalyticBridge. Connect with the Author: . AnalyticBridge – www.analyticbridge.com/profile/VincentGranville . LinkedIn – www.linkedin.com/in/vincentg . Facebook – www.facebook.com/analyticbridge . Twitter – www.twitter.com/analyticbridge . GooglePlus – https://plus.google.com/116460988472927384512 . Quora – www.quora.com/Vincent-Granville 1 Content Introduction Part I - Data Science Recipes 1. New random number generator: simple, strong and fast 2. Lifetime value of an e-mail blast: much longer than you think 3. Two great ideas to create a much better search engine 4. Identifying the number of clusters: finally a solution 5. Online advertising: a solution to optimize ad relevancy 6. Example of architecture for AaaS (Analytics as a Service) 7. Why and how to build a data dictionary for big data sets 8. Hidden decision trees: a modern scoring methodology 9. Scorecards: Logistic, Ridge and Logic Regression 10. Iterative Algorithm for Linear Regression 11. Approximate Solutions to Linear Regression Problems 12. Theorems for Traders 13. Preserving metric and score consistency over time and across clients 14. Advertising: reach and frequency mathematical formulas 15. Real Life Example of Text Mining to Detect Fraudulent Buyers 16. Discount optimization problem in retail analytics 17. Sales forecasts: how to improve accuracy while simplifying models? 18. How could Amazon increase sales by redefining relevancy? 19.