Snapshot: Cytokines II Cristina M
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

IL17RB Antibody Cat
IL17RB Antibody Cat. No.: 62-448 IL17RB Antibody Formalin-fixed and paraffin-embedded human colon Flow cytometric analysis of HepG2 cells using IL17RB carcinoma reacted with IL17RB Antibody , which was Antibody (bottom histogram) compared to a negative peroxidase-conjugated to the secondary antibody, control cell (top histogram). FITC-conjugated goat-anti- followed by DAB staining. rabbit secondary antibodies were used for the analysis. Specifications HOST SPECIES: Rabbit SPECIES REACTIVITY: Human This IL17_ antibody is generated from rabbits immunized with a KLH conjugated synthetic IMMUNOGEN: peptide between 207-234 amino acids from the Central region of human IL17_. TESTED APPLICATIONS: Flow, IHC-P, WB For WB starting dilution is: 1:1000 APPLICATIONS: For IHC-P starting dilution is: 1:10~50 For FACS starting dilution is: 1:10~50 September 25, 2021 1 https://www.prosci-inc.com/il17rb-antibody-62-448.html PREDICTED MOLECULAR 56 kDa WEIGHT: Properties This antibody is prepared by Saturated Ammonium Sulfate (SAS) precipitation followed by PURIFICATION: dialysis CLONALITY: Polyclonal ISOTYPE: Rabbit Ig CONJUGATE: Unconjugated PHYSICAL STATE: Liquid BUFFER: Supplied in PBS with 0.09% (W/V) sodium azide. CONCENTRATION: batch dependent Store at 4˚C for three months and -20˚C, stable for up to one year. As with all antibodies STORAGE CONDITIONS: care should be taken to avoid repeated freeze thaw cycles. Antibodies should not be exposed to prolonged high temperatures. Additional Info OFFICIAL SYMBOL: IL17RB Interleukin-17 receptor B, IL-17 receptor B, IL-17RB, Cytokine receptor-like 4, IL-17 ALTERNATE NAMES: receptor homolog 1, IL-17Rh1, IL17Rh1, Interleukin-17B receptor, IL-17B receptor, IL17RB, CRL4, EVI27, IL17BR ACCESSION NO.: Q9NRM6 PROTEIN GI NO.: 21263748 GENE ID: 55540 USER NOTE: Optimal dilutions for each application to be determined by the researcher. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -
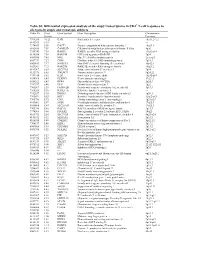
Table S1| Differential Expression Analysis of the Atopy Transcriptome
Table S1| Differential expression analysis of the atopy transcriptome in CD4+ T-cell responses to allergens in atopic and nonatopic subjects Probe ID S.test Gene Symbol Gene Description Chromosome Statistic Location 7994280 10.32 IL4R Interleukin 4 receptor 16p11.2-12.1 8143383 8.95 --- --- --- 7974689 8.50 DACT1 Dapper, antagonist of beta-catenin, homolog 1 14q23.1 8102415 7.59 CAMK2D Calcium/calmodulin-dependent protein kinase II delta 4q26 7950743 7.58 RAB30 RAB30, member RAS oncogene family 11q12-q14 8136580 7.54 RAB19B GTP-binding protein RAB19B 7q34 8043504 7.45 MAL Mal, T-cell differentiation protein 2cen-q13 8087739 7.27 CISH Cytokine inducible SH2-containing protein 3p21.3 8000413 7.17 NSMCE1 Non-SMC element 1 homolog (S. cerevisiae) 16p12.1 8021301 7.15 RAB27B RAB27B, member RAS oncogene family 18q21.2 8143367 6.83 SLC37A3 Solute carrier family 37 member 3 7q34 8152976 6.65 TMEM71 Transmembrane protein 71 8q24.22 7931914 6.56 IL2R Interleukin 2 receptor, alpha 10p15-p14 8014768 6.43 PLXDC1 Plexin domain containing 1 17q21.1 8056222 6.43 DPP4 Dipeptidyl-peptidase 4 (CD26) 2q24.3 7917697 6.40 GFI1 Growth factor independent 1 1p22 7903507 6.39 FAM102B Family with sequence similarity 102, member B 1p13.3 7968236 5.96 RASL11A RAS-like, family 11, member A --- 7912537 5.95 DHRS3 Dehydrogenase/reductase (SDR family) member 3 1p36.1 7963491 5.83 KRT1 Keratin 1 (epidermolytic hyperkeratosis) 12q12-q13 7903786 5.72 CSF1 Colony stimulating factor 1 (macrophage) 1p21-p13 8019061 5.67 SGSH N-sulfoglucosamine sulfohydrolase (sulfamidase) 17q25.3 -

The Role of Interleukin-1 Cytokine Family (IL-1Β, IL-37) and Interleukin-12 Cytokine
bioRxiv preprint doi: https://doi.org/10.1101/502609; this version posted December 22, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. 1 Title: 2 The Role of Interleukin-1 cytokine family (IL-1β, IL-37) and interleukin-12 cytokine 3 family (IL-12, IL-35) in eumycetoma infection pathogenesis. 4 5 6 Authors 7 Amir Abushouk1,2, Amre Nasr1,2,3, Emad Masuadi4, Gamal Allam5,6, Emmanuel E. Siddig7, 8 Ahmed H. Fahal7 9 10 1Department of Basic Medical Sciences, College of Medicine, King Saud Bin Abdul-Aziz 11 University for Health Sciences, Jeddah, Kingdom of Saudi Arabia. E. mail: shouka@ksau- 12 hs.edu.sa 13 14 2King Abdullah International Medical Research Centre, National Guard Health Affairs, 15 Kingdom of Saudi Arabia. 16 17 3Department of Microbiology, College of Sciences and Technology, Al-Neelain University, 18 P.O. Box 1027, Khartoum, Sudan. [email protected] 19 20 4Research Unit, Department of Medical Education, College of Medicine-Riyadh, King Saud 21 Bin Abdul-Aziz University for Health Sciences, Riyadh, Kingdom of Saudi Arabia. E. mail: 22 [email protected] 23 bioRxiv preprint doi: https://doi.org/10.1101/502609; this version posted December 22, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Table SI. Characteristics of 9 Patients Chosen from 60 Patients with Hepatocellular Carcinoma
Table SI. Characteristics of 9 patients chosen from 60 patients with hepatocellular carcinoma. Characteristic No. of patients (%) Age, years <50 3 (33.3) ≥50 6 (66.7) Sex Female 4 (44.4) Male 5 (55.6) Albumin, g/l ≥35 8 (88.9) <35 1 (11.1) AFP, ng/ml ≤20 2 (22.2) >20 7 (77.8) Tumor diameter, cm ≤5 3 (33.3) >5 6 (66.7) Portal vein invasion Without 3 (33.3) With 6 (66.7) BCLC stage A 2 (22.2) B/C 7 (77.8) AFP, α-fetoprotein; BCLC, Barcelona Clinic Liver Cancer. Table SII. List of the gene symbol and relative expression of six significant clusters. Gene symbol SPOT Profile Day 0 Day 3 Day 7 Day 14 Day 21 ALCAM ID_10 16 0 -356.5 378.5 0.1 307.8 ANGPT2 ID_13 16 0 -45.7 197.2 96 60.2 CD80 ID_24 16 0 -177 362 -29 187 CTNNB1 ID_29 16 0 -58 294 83 49 BMP4 ID_37 16 0 -107 189 0 53 CCL28 ID_49 16 0 -307 856 491 347 CCR1 ID_50 16 0 -189 391 236 112 CCR4 ID_53 16 0 -101 361 89 240 CCR7 ID_56 16 0 -98 361 103 66 CLC ID_70 16 0 -147 274 0 188 CRIM1 ID_74 16 0 -642 229 -48 -112 CXCL14 ID_83 16 0 -434 1,154.00 778 376 CXCL16 ID_84 16 0 -128 800 477 180 DKK3 ID_97 16 0 -199 610 107 3 FGFBP1 ID_129 16 0 -224 484 -155 37 FGF20 ID_147 16 0 -113 264 -32 -4 WFIKKN1 ID_163 16 0 -195 544 262 41 GFRA1 ID_175 16 0 -536 443 29 280 GFRA4 ID_178 16 0 -211 208 -47 116 TNFSF18 ID_180 16 0 -14 246 89 68 GPC5 ID_187 16 0 -180 1,345.00 298 509 GH1 ID_194 16 0 -596 206 10 -9 CCHCR1 ID_198 16 0 -108 322 50 238 NRG1 ID_200 16 0 -81 378.5 170.5 187 ICAM1 ID_207 16 0 -136 98 -17 63 IFNB1 ID_213 16 0 -87 233 143 29 IGFBP3 ID_218 16 0 -1,311.00 609 -368 -178 IGFBP4 ID_219 16 0 -

EBI3 Regulates the NK Cell Response to Mouse Cytomegalovirus Infection
EBI3 regulates the NK cell response to mouse cytomegalovirus infection Helle Jensena, Shih-Yu Chenb, Lasse Folkersenc, Garry P. Nolanb, and Lewis L. Laniera,d,1 aDepartment of Microbiology and Immunology, University of California, San Francisco, CA 94143; bDepartment of Microbiology and Immunology, Stanford University, Palo Alto, CA 94304; cDepartment of Systems Biology, Center for Biological Sequence Analysis, Technical University of Denmark, Lyngby DK-2800, Denmark; and dParker Institute for Cancer Immunotherapy, San Francisco, CA 94143 Contributed by Lewis L. Lanier, January 7, 2017 (sent for review November 1, 2016; reviewed by Michael A. Caligiuri and Daniel J. Cua) Natural killer (NK) cells are key mediators in the control of a role in shaping the subsequent adaptive immune responses. cytomegalovirus infection. Here, we show that Epstein–Barr virus- Crosstalk between NK cells and dendritic cells (DCs) early induced 3 (EBI3) is expressed by human NK cells after NKG2D or IL-12 during MCMV infection affects the outcome of the T-cell re- plus IL-18 stimulation and by mouse NK cells during mouse cytomeg- sponses. IL-10 secreted by various immune cells, including NK alovirus (MCMV) infection. The induction of EBI3 protein expression in cells, dampens the T-cell response by negatively affecting the mouse NK cells is a late activation event. Thus, early activation events maturation of DCs, and in the absence of IL-10 secretion of of NK cells, such as IFNγ production and CD69 expression, were not IFNγ and TNFα by NK cells enhances the maturation of DCs, −/− affected in EBI3-deficient (Ebi3 ) C57BL/6 (B6) mice during MCMV which boosts the T-cell response (11). -

Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association Between BMI and Adult-Onset Non- Atopic
Supplementary Material DNA Methylation in Inflammatory Pathways Modifies the Association between BMI and Adult-Onset Non- Atopic Asthma Ayoung Jeong 1,2, Medea Imboden 1,2, Akram Ghantous 3, Alexei Novoloaca 3, Anne-Elie Carsin 4,5,6, Manolis Kogevinas 4,5,6, Christian Schindler 1,2, Gianfranco Lovison 7, Zdenko Herceg 3, Cyrille Cuenin 3, Roel Vermeulen 8, Deborah Jarvis 9, André F. S. Amaral 9, Florian Kronenberg 10, Paolo Vineis 11,12 and Nicole Probst-Hensch 1,2,* 1 Swiss Tropical and Public Health Institute, 4051 Basel, Switzerland; [email protected] (A.J.); [email protected] (M.I.); [email protected] (C.S.) 2 Department of Public Health, University of Basel, 4001 Basel, Switzerland 3 International Agency for Research on Cancer, 69372 Lyon, France; [email protected] (A.G.); [email protected] (A.N.); [email protected] (Z.H.); [email protected] (C.C.) 4 ISGlobal, Barcelona Institute for Global Health, 08003 Barcelona, Spain; [email protected] (A.-E.C.); [email protected] (M.K.) 5 Universitat Pompeu Fabra (UPF), 08002 Barcelona, Spain 6 CIBER Epidemiología y Salud Pública (CIBERESP), 08005 Barcelona, Spain 7 Department of Economics, Business and Statistics, University of Palermo, 90128 Palermo, Italy; [email protected] 8 Environmental Epidemiology Division, Utrecht University, Institute for Risk Assessment Sciences, 3584CM Utrecht, Netherlands; [email protected] 9 Population Health and Occupational Disease, National Heart and Lung Institute, Imperial College, SW3 6LR London, UK; [email protected] (D.J.); [email protected] (A.F.S.A.) 10 Division of Genetic Epidemiology, Medical University of Innsbruck, 6020 Innsbruck, Austria; [email protected] 11 MRC-PHE Centre for Environment and Health, School of Public Health, Imperial College London, W2 1PG London, UK; [email protected] 12 Italian Institute for Genomic Medicine (IIGM), 10126 Turin, Italy * Correspondence: [email protected]; Tel.: +41-61-284-8378 Int. -

Single-Cell Analysis of Crohn's Disease Lesions Identifies
bioRxiv preprint doi: https://doi.org/10.1101/503102; this version posted December 20, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Single-cell analysis of Crohn’s disease lesions identifies a pathogenic cellular module associated with resistance to anti-TNF therapy JC Martin1,2,3, G Boschetti1,2,3, C Chang1,2,3, R Ungaro4, M Giri5, LS Chuang5, S Nayar5, A Greenstein6, M. Dubinsky7, L Walker1,2,5,8, A Leader1,2,3, JS Fine9, CE Whitehurst9, L Mbow9, S Kugathasan10, L.A. Denson11, J.Hyams12, JR Friedman13, P Desai13, HM Ko14, I Laface1,2,8, Guray Akturk1,2,8, EE Schadt15,16, S Gnjatic1,2,8, A Rahman1,2,5,8, , M Merad1,2,3,8,17,18*, JH Cho5,17,*, E Kenigsberg1,15,16,17* 1 Precision Immunology Institute, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. 2 Tisch Cancer Institute, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. 3 Department of Oncological Sciences, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. 4 The Dr. Henry D. Janowitz Division of Gastroenterology, Icahn School of Medicine at Mount Sinai, New York City, NY 10029, USA. 5 Charles Bronfman Institute for Personalized Medicine, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. 6 Department of Colorectal Surgery, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA 7 Department of Pediatrics, Susan and Leonard Feinstein IBD Clinical Center, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. -

Evolutionary Divergence and Functions of the Human Interleukin (IL) Gene Family Chad Brocker,1 David Thompson,2 Akiko Matsumoto,1 Daniel W
UPDATE ON GENE COMPLETIONS AND ANNOTATIONS Evolutionary divergence and functions of the human interleukin (IL) gene family Chad Brocker,1 David Thompson,2 Akiko Matsumoto,1 Daniel W. Nebert3* and Vasilis Vasiliou1 1Molecular Toxicology and Environmental Health Sciences Program, Department of Pharmaceutical Sciences, University of Colorado Denver, Aurora, CO 80045, USA 2Department of Clinical Pharmacy, University of Colorado Denver, Aurora, CO 80045, USA 3Department of Environmental Health and Center for Environmental Genetics (CEG), University of Cincinnati Medical Center, Cincinnati, OH 45267–0056, USA *Correspondence to: Tel: þ1 513 821 4664; Fax: þ1 513 558 0925; E-mail: [email protected]; [email protected] Date received (in revised form): 22nd September 2010 Abstract Cytokines play a very important role in nearly all aspects of inflammation and immunity. The term ‘interleukin’ (IL) has been used to describe a group of cytokines with complex immunomodulatory functions — including cell proliferation, maturation, migration and adhesion. These cytokines also play an important role in immune cell differentiation and activation. Determining the exact function of a particular cytokine is complicated by the influence of the producing cell type, the responding cell type and the phase of the immune response. ILs can also have pro- and anti-inflammatory effects, further complicating their characterisation. These molecules are under constant pressure to evolve due to continual competition between the host’s immune system and infecting organisms; as such, ILs have undergone significant evolution. This has resulted in little amino acid conservation between orthologous proteins, which further complicates the gene family organisation. Within the literature there are a number of overlapping nomenclature and classification systems derived from biological function, receptor-binding properties and originating cell type. -

Inflammatory Modulation of Hematopoietic Stem Cells by Magnetic Resonance Imaging
Electronic Supplementary Material (ESI) for RSC Advances. This journal is © The Royal Society of Chemistry 2014 Inflammatory modulation of hematopoietic stem cells by Magnetic Resonance Imaging (MRI)-detectable nanoparticles Sezin Aday1,2*, Jose Paiva1,2*, Susana Sousa2, Renata S.M. Gomes3, Susana Pedreiro4, Po-Wah So5, Carolyn Ann Carr6, Lowri Cochlin7, Ana Catarina Gomes2, Artur Paiva4, Lino Ferreira1,2 1CNC-Center for Neurosciences and Cell Biology, University of Coimbra, Coimbra, Portugal, 2Biocant, Biotechnology Innovation Center, Cantanhede, Portugal, 3King’s BHF Centre of Excellence, Cardiovascular Proteomics, King’s College London, London, UK, 4Centro de Histocompatibilidade do Centro, Coimbra, Portugal, 5Department of Neuroimaging, Institute of Psychiatry, King's College London, London, UK, 6Cardiac Metabolism Research Group, Department of Physiology, Anatomy & Genetics, University of Oxford, UK, 7PulseTeq Limited, Chobham, Surrey, UK. *These authors contributed equally to this work. #Correspondence to Lino Ferreira ([email protected]). Experimental Section Preparation and characterization of NP210-PFCE. PLGA (Resomers 502 H; 50:50 lactic acid: glycolic acid) (Boehringer Ingelheim) was covalently conjugated to fluoresceinamine (Sigma- Aldrich) according to a protocol reported elsewhere1. NPs were prepared by dissolving PLGA (100 mg) in a solution of propylene carbonate (5 mL, Sigma). PLGA solution was mixed with perfluoro- 15-crown-5-ether (PFCE) (178 mg) (Fluorochem, UK) dissolved in trifluoroethanol (1 mL, Sigma). This solution was then added to a PVA solution (10 mL, 1% w/v in water) dropwise and stirred for 3 h. The NPs were then transferred to a dialysis membrane and dialysed (MWCO of 50 kDa, Spectrum Labs) against distilled water before freeze-drying. Then, NPs were coated with protamine sulfate (PS). -

The Innate Cytokines IL-25, IL-33, and TSLP Cooperate in the Induction Of
The Innate Cytokines IL-25, IL-33, and TSLP Cooperate in the Induction of Type 2 Innate Lymphoid Cell Expansion and Mucous Metaplasia in Rhinovirus-Infected This information is current as Immature Mice of September 29, 2021. Mingyuan Han, Charu Rajput, Jun Y. Hong, Jing Lei, Joanna L. Hinde, Qian Wu, J. Kelley Bentley and Marc B. Hershenson J Immunol 2017; 199:1308-1318; Prepublished online 12 Downloaded from July 2017; doi: 10.4049/jimmunol.1700216 http://www.jimmunol.org/content/199/4/1308 http://www.jimmunol.org/ Supplementary http://www.jimmunol.org/content/suppl/2017/07/12/jimmunol.170021 Material 6.DCSupplemental References This article cites 64 articles, 15 of which you can access for free at: http://www.jimmunol.org/content/199/4/1308.full#ref-list-1 by guest on September 29, 2021 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2017 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. -

KRAS Mutations Are Negatively Correlated with Immunity in Colon Cancer
www.aging-us.com AGING 2021, Vol. 13, No. 1 Research Paper KRAS mutations are negatively correlated with immunity in colon cancer Xiaorui Fu1,2,*, Xinyi Wang1,2,*, Jinzhong Duanmu1, Taiyuan Li1, Qunguang Jiang1 1Department of Gastrointestinal Surgery, The First Affiliated Hospital of Nanchang University, Nanchang, Jiangxi, People's Republic of China 2Queen Mary College, Medical Department, Nanchang University, Nanchang, Jiangxi, People's Republic of China *Equal contribution Correspondence to: Qunguang Jiang; email: [email protected] Keywords: KRAS mutations, immunity, colon cancer, tumor-infiltrating immune cells, inflammation Received: March 27, 2020 Accepted: October 8, 2020 Published: November 26, 2020 Copyright: © 2020 Fu et al. This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT The heterogeneity of colon cancer tumors suggests that therapeutics targeting specific molecules may be effective in only a few patients. It is therefore necessary to explore gene mutations in colon cancer. In this study, we obtained colon cancer samples from The Cancer Genome Atlas, and the International Cancer Genome Consortium. We evaluated the landscape of somatic mutations in colon cancer and found that KRAS mutations, particularly rs121913529, were frequent and had prognostic value. Using ESTIMATE analysis, we observed that the KRAS-mutated group had higher tumor purity, lower immune score, and lower stromal score than the wild- type group. Through single-sample Gene Set Enrichment Analysis and Gene Set Enrichment Analysis, we found that KRAS mutations negatively correlated with enrichment levels of tumor infiltrating lymphocytes, inflammation, and cytolytic activities.