Development of a Comprehensive Genetic Tool for Identification Of
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Effect of Adding Garlic Powder (Allium Sativum) and Black Seed
Journal of Natural Sciences Research www.iiste.org ISSN 2224-3186 (Paper) ISSN 2225-0921 (Online) Vol.3, No.1, 2013 Effect of adding Garlic Powder ( Allium sativum ) and Black Seed ( Nigella sativa ) in Feed on Broiler Growth Performance and Intestinal Wall Structure Jamel M. Saeid , Arkan B. Mohamed * and Maad A. AL-Baddy Department of Animal Resources, College of Agriculture, University of Tikrit,Tikrit,Iraq * E-mail of the corresponding author: [email protected] The research is financed by Asian Development Bank. No. 2006-A171(Sponsoring information) Abstract This study was conducted to investigate the effect of garlic powder (GP) black seed (BS) and plant premix (GP and BS) in feed on broiler growth and intestinal wall structure. The result included 480 Hubbard broiler chicks( day- old) There were 4 treatment groups each consisting of 3 replicates .The four dietary treatments consisted of a control ( basal diet ) , basal diet + 0.5% GP , basal diet +0.5% BS and basal diet +0.5% plant premix (GP and BS) , to the starter and finisher diet. The experiment lasted 42 days. Body weight , body weight gain , feed intake and feed conversion ratio were determined weekly and intestinal characteristics were determined at the end of the study (42 day) .The addition of GP and BS plant premix (GP and BS) to the diet resulted in significantly higher body weight , body weight gain and feed intake as compared to that of control group. However , feed conversion ratio was not influenced by dietary treatment (p>0.05) . The villus height , crypt depth and crypt depth to villus height ratio was significantly higher in group garlic powder and plant premix (GP and BS) than other groups. -

Studies on the Dual Antioxidant and Antibacterial Properties of Parsley (Petroselinum Crispum) and Cilantro (Coriandrum Sativum) Extracts
Food Chemistry Food Chemistry 97 (2006) 505–515 www.elsevier.com/locate/foodchem Studies on the dual antioxidant and antibacterial properties of parsley (Petroselinum crispum) and cilantro (Coriandrum sativum) extracts Peter Y.Y. Wong, David D. Kitts * Department of Food, Nutrition and Health, The University of British Columbia, 6640 N.W. Marine Drive, Vancouver, BC, Canada V6T 1Z4 Received 23 August 2004; received in revised form 10 May 2005; accepted 10 May 2005 Abstract Antioxidant and antibacterial activities of freeze-dried and irradiated parsley (Petroselinum crispum) and cilantro (Coriandrum sativum) leaves and stems were determined on methanol and water extracts. The total phenolic content was quantified with the Folin–Ciocalteau reagent. Several mechanisms of potential antioxidant activity of all extracts, including determining relative free radical-scavenging and ferrous ion-chelating activities, as well as reducing power, were examined. Assessment of the total antioxi- dant activity of all extracts was done using an iron-induced linoleic acid oxidation model system. Antimicrobial activity towards Bacillus subtilis and Escherichia coli by different extracts was assessed by determining cell damage. Total phenolic content varied between parsley and cilantro, leaf and stem, as well as methanol and water extracts. Methanol-derived leaf extracts exhibited sig- nificantly (p < 0.05) greater radical-scavenging activity towards both lipid- and water-soluble radicals, which was attributed to the total phenolic content. Ferrous ion-chelating activity was significantly (p < 0.05) greater in the stem methanol extracts, and corre- sponded to antioxidant activity. Prooxidant activity was a feature of all aqueous extracts and corresponded to the reducing activity of both leaf and stem parts of parsley and cilantro. -

Evaluation of Cuminum Cyminum and Coriandrum Sativum on Profenofos Induced Nephrotoxicity in Swiss Albino Mice Arun Kumar*,Jiv Kant Singh**, Md
4771 Arun Kumar et al./ Elixir Appl. Botany 39 (2011) 4771-4773 Available online at www.elixirpublishers.com (Elixir International Journal) Applied Botany Elixir Appl. Botany 39 (2011) 4771-4773 Evaluation of Cuminum cyminum and Coriandrum sativum on Profenofos induced nephrotoxicity in Swiss albino mice Arun Kumar*,Jiv Kant Singh**, Md. Ali*, Ranjit Kumar*, Arvind Kumar**, A.Nath*, A.K.Roy***, S.P.Roy** and J.K.Singh* *Mahavir Cancer Institute & Research Center, Patna **Department of Biotechnology, T.M Bhagalpur University, Bhagalpur ***Department of Botany, T.M Bhagalpur University, Bhagalpur. ARTICLE INFO ABSTRACT Article history: Now a day’s various types of organophosphate compound are being utilised by farmers in Received: 21 July 2011; massive amount to increase the productivity of crops by destroying the insecticides. Received in revised form: Profenofos is one of the organophosphate which is widely used by the farmers. It is largely 21 September 2011; known to cause toxicity in various organs, such as the liver and brain. So, to evaluate the Accepted: 28 September 2011; effect of Profenofos, experiment was designed on Swiss albino model. So, the present study aims to observe the effect of Cuminum cyminum (Cumin) and Coriandrum sativum Keywords (Coriander) on kidney exposed with Profenofos on female swiss albino mice. The Profenofos, parameters utilized for the study were through biochemical analysis. The impact of herbal Coriander; plants Cumin and Coriander were observed on the Profenofos induced nephrotoxicity. The Cumin and Nephrotoxicity. study shows Cumin was more effective in normalizing the Uric acid level rather than Coriander, while the coriander is more effective in normalizing the creatinine level than Cumin. -

Cannabis – a Complex and Rapidly Evolving Landscape
CANNABIS – A COMPLEX AND RAPIDLY EVOLVING LANDSCAPE Abstract ABOUT THE AUTHOR The humble Cannabis sativa plant, cultivated for millennia for its psychoactive properties and more, is today considered one of the most controversial and complex plants in the world. Starting in the early to mid-20th century, much of its use became recreational, but by the early 1970s discoveries began to emerge around its potential medical efficacies. This article will discuss current knowledge of how cannabis engages with the brain and the endocannabinoid system (ECS) and provide an overview of the new market landscapes brought about by changes in governing laws Dr. Georgiana Willwerth-Pascutiu and regulations, which are affecting usage by our current and potential [email protected] customers. It will also explore the additional hazards, concerns, and Georgiana Willwerth-Pascutiu is Vice President, Global Medical Director considerations of cannabis use in countries where it remains illegal. at RGA. She is board certified in Insurance Medicine by the American Introduction Academy of Insurance Medicine (AAIM) and specialized in internal medicine, Naturally occurring psychoactive substances have been part of human life nephrology and ultrasonography. for millennia. One of the most frequently utilized plant sources of these Dr. Willwerth-Pascutiu is also a past substances, Cannabis sativa, is also the best-known worldwide. For the president of the Canadian Life Insurance Medical Officers Association (CLIMOA) past half-century, scientific and medical interest in its many compounds, and currently chairs its scientific known as cannabinoids, has been increasing. Today, the two best-known, committee. She is a frequent presenter and has contributed several articles to delta-9 tetrahydrocannabinol (THC), its psychoactive chemical, and insurance industry publications. -

Special Report on ASX-Listed Cannabis and Hemp Stocks
Special Report on ASX-listed Cannabis and Hemp stocks An exciting new sector 24 March 2020 From humble beginnings in Canada around ten years ago the cannabis and hemp industries have blossomed into a major force to be reckoned with by investors the world over. Australia is no exception, with many cannabis and hemp companies having gone live on ASX over the last five years. However, many investors are unfamiliar with the dynamics of this exciting new sector. Pitt Street Research now seeks to close that information gap with our Special Report on Cannabis and Hemp, released 24 March 2020. Welcome to the cannabis and hemp revolution Cannabis and hemp have fuelled a major investment boom since 2014 largely because of the known therapeutic benefits of medicinal cannabis. Governments around the world have responded to the scientific evidence and made it easier for patients to access cannabis-based medicine. Concurrently, voters in many countries have become more favourably disposed towards the legalisation of recreational cannabis. These two trends have fuelled a boom in cannabis, while hemp, from a different plant, had also benefited as investors have moved to use this plant for a variety of purposes, most notably in food. It’s fair to say that cannabis and hemp have quickly become respectable industries worthy of investor attention. Many have come to the view that cannabis and hemp are agents of serious economic change, with potential to seriously disrupt Subscribe to our research HERE sectors as diverse as drinks, building materials and, of course, medicine. Analyst: Stuart Roberts Why should the Canadians have all the fun? Tel: +61 (0)447 247 909 Canada was the origin of the current cannabis and hemp boom because the regulatory framework changed in that [email protected] country around 2013 in a way that allowed entrepreneurs to flourish while the public equity markets allowed large amounts of capital to be raised. -

A Review on Cannabis Sativa: Its Compounds and Their Effects
Int. J. Pharm. Sci. Rev. Res., 53(2), November - December 2018; Article No. 12, Pages: 59-63 ISSN 0976 – 044X Review Article A Review on Cannabis sativa: Its Compounds and Their Effects Ranju Rajput, *Dr. Krishan Kumar Department of Food and Biotechnology, Jayoti Vidyapeeth Women’s University, Jaipur, Rajasthan, India. *Corresponding author’s E-mail: [email protected] Received: 01-11-2018; Revised: 25-11-2018; Accepted: 10-12-2018. ABSTRACT Our society often considered the use of cannabis is an under-reported activity. Cannabis is used to relieve neuropathic and chronic pain. Cannabis, produced from the Cannabis sativa plant, have been used in three forms: herbal cannabis, the dried leaves and flowering tops The resin of the cannabis is the pressed secretions of the plant, known as ‘hashish’ or ‘charash. Cannabis sativa is an herbaceous species originated from Central Asia. It has been used in medicine and as a source of textile fiber since ancient times. The cannabis sativa is a fast growing plant attracted the people’s interest because of its multi-purpose applications. It is a rich source of photochemical, cellulose and woody fibers. The more interest is also due to its metabolites which show potent bioactivities on human health. In this review, the phytochemicals is discussed by putting a special emphasis on molecules including cannabinoids, terpenes and phenolic compounds. Cannabinoids are represented as the most studied group of compounds, because of their wide range of pharmaceutical effects in humans, including psychotropic activities. This article aims to update the current knowledge and evidence of using cannabis and its derivatives with a view to the sociolegal context and perspectives for future research. -

Forensic Palynology: Why Do It and How It Works
Forensic Science International 163 (2006) 163–172 www.elsevier.com/locate/forsciint Editorial Forensic palynology: Why do it and how it works Abstract Forensic palynology has been a law enforcement tool for over 50 years. Forensic palynology is the application of pollen and spores in solving legal issues, either civil or criminal. Pollen and spores can be obtained from an extremely wide range of items, including bodies. Pollen and spores provide clues as to the source of the items and the characteristics of the environments from which the material on them is sourced. Their usefulness lies in a combination of their abundance, dispersal mechanisms, resistance to mechanical and chemical destruction, microscopic size, and morphology. Their often complex morphology allows identification to an individual parent plant taxon that can be related to a specific ecological habitat or a specific scene. Pollen and spore assemblages characterise different environments and scenes and can easily be picked up and transported away from scenes of interest without providing any visual clue to a suspect as to what has occurred. With so many publications and high-profile cases involving forensic palynology and environmental analysis now receiving publicity, the future of this branch of forensic science is assured. Furthermore, with the development of multi-disciplinary approaches to environmental analyses of crime scenes, far more detailed information is now available to law enforcement agencies, enabling them to determine with greater accuracy what may have happened during the commission of criminal activities. # 2006 Elsevier Ireland Ltd. All rights reserved. Keywords: Forensic science; Forensic palynology; Pollen; Spores; Environmental analysis; Trace evidence 1. -

Familial Mediterranean Fever
:: Familial Mediterranean fever – This document is a translation of the French recommendations drafted by Prof Gilles Grateau, Dr Véronique Hentgen, Dr Katia Stankovic Stojanovic and Dr Gilles Bagou reviewed and published by Orphanet in 2010. – Some of the procedures mentioned, particularly drug treatments, may not be validated in the country where you practice. Synonyms: Periodic disease, FMF Definition: Familial Mediterranean fever (FMF) is an auto-inflammatory disease of genetic origin, affecting Mediterranean populations and characterised by recurrent attacks of fever accompanied by polyserositis that causes the symptoms. Colchicine is the basic reference treatment and is designed to tackle inflammatory attacks and prevent amyloidosis, the most severe complication of FMF. Further information: See the Orphanet abstract http://www.orpha.net/data/patho/Pro/en/Emergency_FamilialMediterraneanFever-enPro920.pdf ©Orphanet UK 1/5 Pre-hospital emergency care recommendations Call for a patient suffering from Familial mediterranean fever Synonyms periodic disease FMF Mechanisms auto-inflammatory disease that affects Mediterranean populations in particular, due to mutation of the MEFV gene that codes pyrin or marenostrin, this being the underlying cause of congenital immune dysfunction; repeated inflammatory attacks can lead to amyloidosis, particularly renal Specific risks in emergency situations acute inflammatory attack, particularly abdominal (pseudo-surgical), but also thoracic, articular (knees) or testicular fever as an expression -

Past, Present, and Future of Medical Cannabis
PAST, PRESENT AND FUTURE OF MEDICAL CANNABIS Asare B. Christian, MD, MPH Associate Outpatient Medical Director, Good Shepherd Rehabilitation Network Clinical Faculty, Penn Medicine Outline • Medical History of Cannabis • Pharmacology of Cannabinoids • Endocannabinoid System Physiology • Safety and Adverse Effects of Cannabis • Impairments from Cannabis • Future of Cannabis in Medicine HISTORICAL USE OF CANNABIS IN MEDICINE Historical Use of Cannabis in Medicine • 2700 BC Emperor Shen-Nung • Analgesia, rheumatism, beriberi, malaria, gout and poor memory • 1839 William O’ Shaughnessy • Introduced medical cannabis to England • 1854 Cannabis enters Dispensatory of US • Sir William Osler on migraine…“cannabis Indica is probably the most satisfactory remedy.” • Empirical Medicine of the 19th Century • Combined morphine, cannabis and capsicum • Provided-phyto-opiod, Phytocannabinoids and phytovanilloid in one prep • Better outpatient pain relief than is currently available in st 21 centuryLi HL: An archaeological and historical account of cannabis in China. Econ Bot (1974) 28:437- 448. O'Shaughnessy WB: On the preparations of the Indian hemp, or gunjah (Cannabis indica): Their effects on the animal system in health, and their utility in the treatment of tetanus and other convulsive diseases. Trans Med Phys Soc Bengal (1838-1840) Historical Use of Cannabis in Medicine • 19th and 20th Century • US Pharmacopoeia 1850-1942 • Restrictions of sale and use 1937 • Boggs 1951 and Narcotic Control Act of 1956 – legal penalties • 1996 - California permits cannabis use for medicine • Compassionate Use Act • 2017 - 28 states as well as Washington, DC, Guam and Puerto Rico • 21 states decriminalized • 8 states allow recreational use (AL, CA, CO, MN, MA, NV, OR, WA, DC) Bridgeman, Mary Barna, and Daniel T. -
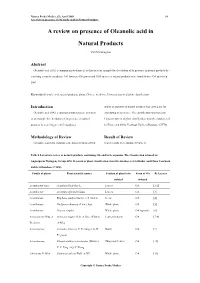
A Review on Presence of Oleanolic Acid in Natural Products
Natura Proda Medica, (2), April 2009 64 A review on presence of Oleanolic acid in Natural Products A review on presence of Oleanolic acid in Natural Products YEUNG Ming Fai Abstract Oleanolic acid (OA), a common phytochemical, is chosen as an example for elucidation of its presence in natural products by searching scientific databases. 146 families, 698 genera and 1620 species of natural products were found to have OA up to Sep 2007. Keywords Oleanolic acid, natural products, plants, Chinese medicine, Linnaeus system of plant classification Introduction and/or its saponins in natural products was carried out for Oleanolic acid (OA), a common phytochemical, is chosen elucidating its pressence. The classification was based on as an example for elucidation of its presence in natural Linnaeus system of plant classification from the databases of products by searching scientific databases. SciFinder and China Yearbook Full-text Database (CJFD). Methodology of Review Result of Review Literature search for isolation and characterization of OA Search results were tabulated (Table 1). Table 1 Literature review of natural products containing OA and/or its saponins. The classification is based on Angiosperm Phylogeny Group APG II system of plant classification from the databases of SciFinder and China Yearbook Full-text Database (CJFD). Family of plants Plant scientific names Position of plant to be Form of OA References isolated isolated Acanthaceae Juss. Acanthus illicifolius L. Leaves OA [1-2] Acanthaceae Avicennia officinalis Linn. Leaves OA [3] Acanthaceae Blepharis sindica Stocks ex T. Anders Seeds OA [4] Acanthaceae Dicliptera chinensis (Linn.) Juss. Whole plant OA [5] Acanthaceae Justicia simplex Whole plant OA saponins [6] Actinidiaceae Gilg. -

On the Origin of Hops: Genetic Variability, Phylogenetic Relationships, and Ecological Plasticity of Humulus (Cannabaceae)
ON THE ORIGIN OF HOPS: GENETIC VARIABILITY, PHYLOGENETIC RELATIONSHIPS, AND ECOLOGICAL PLASTICITY OF HUMULUS (CANNABACEAE) A DISSERTATION SUBMITTED TO THE GRADUATE DIVISION OF THE UNIVERSITY OF HAWAI‘I AT MĀNOA IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY IN BOTANY MAY 2014 By Jeffrey R. Boutain DISSERTATION COMMITTEE: Will C. McClatchey, Chairperson Mark D. Merlin Sterling C. Keeley Clifford W. Morden Stacy Jørgensen Copyright © 2014 by Jeffrey R. Boutain ii This dissertation is dedicated to my family tree. iii ACKNOWLEDGEMENTS There are a number of individuals to whom I am indebted in many customs. First and foremost, I thank my committee members for their contribution, patience, persistence, and motivation that helped me complete this dissertation. Specifically, thank you Dr. Will McClatchey for the opportunity to study in a botany program with you as my advisor and especially the encouragement to surf plant genomes. Also with great gratitude, thank you Dr. Sterling Keeley for the opportunity to work on much of this dissertation in your molecular phylogenetics and systematics lab. In addition, thank you Dr. Mark Merlin for numerous brainstorming sessions as well as your guidance and expert perspective on the Cannabaceae. Also, thank you Dr. Cliff Morden for the opportunity to work in your lab where the beginnings of this molecular research took place. Thank you Dr. Jianchu Xu for welcoming me into your lab group at the Kunming Institute of Botany, Chinese Academy of Sciences (CAS) and the opportunity to study the Yunnan hop. In many ways, major contributions towards the completion of this dissertation have come from my family, and I thank you for your unconditional encouragement, love, and support. -

A Belated Green Revolution for Cannabis: Virtual Genetic Resources to Fast-Track Cultivar Development
REVIEW published: 29 July 2016 doi: 10.3389/fpls.2016.01113 A Belated Green Revolution for Cannabis: Virtual Genetic Resources to Fast-Track Cultivar Development Matthew T. Welling 1, Tim Shapter 1, 2, Terry J. Rose 1, Lei Liu 1, Rhia Stanger 1 and Graham J. King 1* 1 Southern Cross Plant Science, Southern Cross University, Lismore, NSW, Australia, 2 Ecofibre Industries Operations Pty Ltd, Maleny, QLD, Australia Cannabis is a predominantly diecious phenotypically diverse domesticated genus with few if any extant natural populations. International narcotics conventions and associated legislation have constrained the establishment, characterization, and use of Cannabis genetic resource collections. This has resulted in the underutilization of genepool variability in cultivar development and has limited the inclusion of secondary genepools associated with genetic improvement strategies of the Green Revolution. The structured screening of ex situ germplasm and the exploitation of locally-adapted intraspecific traits is expected to facilitate the genetic improvement Edited by: Jaime Prohens, of Cannabis. However, limited attempts have been made to establish the full extent of Polytechnic University of Valencia, genetic resources available for pre-breeding. We present a thorough critical review of Spain Cannabis ex situ genetic resources, and discuss recommendations for conservation, Reviewed by: pre-breeding characterization, and genetic analysis that will underpin future cultivar Ryan C. Lynch, University of Colorado Boulder, USA development. We consider East Asian germplasm to be a priority for conservation Giuseppe Mandolino, based on the prolonged historical cultivation of Cannabis in this region over a range Council for Agricultural Research and Economics, Italy of latitudes, along with the apparent high levels of genetic diversity and relatively *Correspondence: low representation in published genetic resource collections.